Abstract
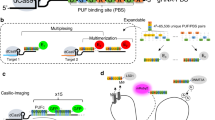
Traditional methods for the generation of DNA damage are not well suited for the observation of spatiotemporal aspects of damaged chromosomal loci. We describe a protocol for the derivation of a cellular system to induce and to visualize chromosome damage at specific sites of the mammalian genome in living cells. The system is based on the stable integration of endonuclease I-SceI recognition sites flanked by bacterial LacO/TetO operator arrays, coupled with retroviral-mediated integration of their fluorescent repressors (LacR/TetR) to visualize the LacO/TetO sites. Expression of the I-SceI endonuclease induces double-strand breaks (DSBs) specifically at the sites of integration, and it permits the dynamics of damaged chromatin to be followed by time-lapse microscopy. Sequential LacO-I-SceI/TetO-I-SceI integrations in multiple chromosomes permit the generation of a system to visualize the formation of chromosome translocations in living cells. This protocol requires intermediate cell culture and molecular biology skills, and it is adaptable to the efficient derivation of any integrated clonal reporter system of interest in ∼3–5 months.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ciccia, A. & Elledge, S.J. The DNA damage response: making it safe to play with knives. Mol. Cell. 40, 179–204 (2010).
Misteli, T. & Soutoglou, E. The emerging role of nuclear architecture in DNA repair and genome maintenance. Nat. Rev. Mol. Cell Biol. 10, 243–254 (2009).
Soria, G., Polo, S.E. & Almouzni, G. Prime, repair, restore: the active role of chromatin in the DNA damage response. Mol. Cell 46, 722–734 (2012).
Dion, V. & Gasser, S.M. Chromatin movement in the maintenance of genome stability. Cell 152, 1355–1364 (2013).
Jakob, B., Splinter, J., Durante, M. & Taucher-Scholz, G. Live cell microscopy analysis of radiation-induced DNA double-strand break motion. Proc. Natl. Acad. Sci. USA 106, 3172–3177 (2009).
Aten, J.A. et al. Dynamics of DNA double-strand breaks revealed by clustering of damaged chromosome domains. Science 303, 92–95 (2004).
Krawczyk, P.M. et al. Chromatin mobility is increased at sites of DNA double-strand breaks. J. Cell Sci. 125, 2127–2133 (2012).
Agarwal, S. et al. ATP-dependent and independent functions of Rad54 in genome maintenance. J. Cell Biol. 192, 735–750 (2011).
Mehta, I.S., Kulashreshtha, M., Chakraborty, S., Kolthur-Seetharam, U. & Rao, B.J. Chromosome territories reposition during DNA damage-repair response. Genome Biol. 14, R135 (2013).
Zink, D., Bornfleth, H., Visser, A., Cremer, C. & Cremer, T. Organization of early and late replicating DNA in human chromosome territories. Exp. Cell Res. 247, 176–188 (1999).
Zink, D. et al. Structure and dynamics of human interphase chromosome territories in vivo. Hum. Genet. 102, 241–251 (1998).
Gotoh, E. Visualizing the dynamics of chromosome structure formation coupled with DNA replication. Chromosoma 116, 453–462 (2007).
Kruhlak, M.J. et al. Changes in chromatin structure and mobility in living cells at sites of DNA double-strand breaks. J. Cell Biol. 172, 823–834 (2006).
Michaelis, C., Ciosk, R. & Nasmyth, K. Cohesins: chromosomal proteins that prevent premature separation of sister chromatids. Cell 91, 35–45 (1997).
Robinett, C.C. et al. In vivo localization of DNA sequences and visualization of large-scale chromatin organization using lac operator/repressor recognition. J. Cell Biol. 135, 1685–1700 (1996).
Roukos, V. et al. Spatial dynamics of chromosome translocations in living cells. Science 341, 660–664 (2013).
Soutoglou, E. et al. Positional stability of single double-strand breaks in mammalian cells. Nat. Cell Biol. 9, 675–682 (2007).
Belmont, A.S., Li, G., Sudlow, G. & Robinett, C. Visualization of large-scale chromatin structure and dynamics using the lac operator/lac repressor reporter system. Methods Cell Biol. 58, 203–222 (1999).
Gonzalez-Serricchio, A.S. & Sternberg, P.W. Visualization of C. elegans transgenic arrays by GFP. BMC Genet. 7, 36 (2006).
Lassadi, I. & Bystricky, K. Tracking of single and multiple genomic loci in living yeast cells. Methods Mol. Biol. 745, 499–522 (2011).
Lisby, M., Mortensen, U.H. & Rothstein, R. Colocalization of multiple DNA double-strand breaks at a single Rad52 repair centre. Nat. Cell Biol. 5, 572–577 (2003).
Meister, P., Gehlen, L.R., Varela, E., Kalck, V. & Gasser, S.M. Visualizing yeast chromosomes and nuclear architecture. Methods Enzymol. 470, 535–567 (2010).
Nagai, S. et al. Functional targeting of DNA damage to a nuclear pore-associated SUMO-dependent ubiquitin ligase. Science 322, 597–602 (2008).
Rohner, S., Gasser, S.M. & Meister, P. Modules for cloning-free chromatin tagging in Saccharomyces cerevisiae. Yeast 25, 235–239 (2008).
Strukov, Y.G. & Belmont, A.S. Development of mammalian cell lines with lac operator-tagged chromosomes. Cold Spring Harb. Protoc. 2008 pdb prot4903 (2008).
Vazquez, J., Belmont, A.S. & Sedat, J.W. The dynamics of homologous chromosome pairing during male Drosophila meiosis. Curr. Biol. 12, 1473–1483 (2002).
Dion, V., Kalck, V., Horigome, C., Towbin, B.D. & Gasser, S.M. Increased mobility of double-strand breaks requires Mec1, Rad9 and the homologous recombination machinery. Nat. Cell Biol. 14, 502–509 (2012).
Mine-Hattab, J. & Rothstein, R. Increased chromosome mobility facilitates homology search during recombination. Nat. Cell Biol. 14, 510–517 (2012).
Chubb, J.R., Boyle, S., Perry, P. & Bickmore, W.A. Chromatin motion is constrained by association with nuclear compartments in human cells. Curr. Biol. 12, 439–445 (2002).
Thomson, I., Gilchrist, S., Bickmore, W.A. & Chubb, J.R. The radial positioning of chromatin is not inherited through mitosis but is established de novo in early G1. Curr. Biol. 14, 166–172 (2004).
Vazquez, J., Belmont, A.S. & Sedat, J.W. Multiple regimes of constrained chromosome motion are regulated in the interphase Drosophila nucleus. Curr. Biol. 11, 1227–1239 (2001).
Chuang, C.H. et al. Long-range directional movement of an interphase chromosome site. Curr. Biol. 16, 825–831 (2006).
Janicki, S.M. et al. From silencing to gene expression: real-time analysis in single cells. Cell 116, 683–698 (2004).
Shanbhag, N.M., Rafalska-Metcalf, I.U., Balane-Bolivar, C., Janicki, S.M. & Greenberg, R.A. ATM-dependent chromatin changes silence transcription in cis to DNA double-strand breaks. Cell 141, 970–981 (2010).
Tsukamoto, T. et al. Visualization of gene activity in living cells. Nat. Cell Biol. 2, 871–878 (2000).
Soutoglou, E. & Misteli, T. Activation of the cellular DNA damage response in the absence of DNA lesions. Science 320, 1507–1510 (2008).
Bonilla, C.Y., Melo, J.A. & Toczyski, D.P. Colocalization of sensors is sufficient to activate the DNA damage checkpoint in the absence of damage. Mol. Cell 30, 267–276 (2008).
Bergmann, J.H. et al. Epigenetic engineering shows H3K4me2 is required for HJURP targeting and CENP-A assembly on a synthetic human kinetochore. EMBO J. 30, 328–340 (2011).
Luijsterburg, M.S. et al. A new non-catalytic role for ubiquitin ligase RNF8 in unfolding higher-order chromatin structure. EMBO J. 31, 2511–2527 (2012).
Tumbar, T. & Belmont, A.S. Interphase movements of a DNA chromosome region modulated by VP16 transcriptional activator. Nat. Cell Biol. 3, 134–139 (2001).
Verschure, P.J. et al. In vivo HP1 targeting causes large-scale chromatin condensation and enhanced histone lysine methylation. Mol. Cell Biol. 25, 4552–4564 (2005).
Kaiser, T.E., Intine, R.V. & Dundr, M. De novo formation of a subnuclear body. Science 322, 1713–1717 (2008).
Shevtsov, S.P. & Dundr, M. Nucleation of nuclear bodies by RNA. Nat. Cell Biol. 13, 167–173 (2011).
Zolghadr, K. et al. A fluorescent two-hybrid assay for direct visualization of protein interactions in living cells. Mol. Cell Proteomics 7, 2279–2287 (2008).
Zolghadr, K., Rothbauer, U. & Leonhardt, H. The fluorescent two-hybrid (F2H) assay for direct analysis of protein-protein interactions in living cells. Methods Mol. Biol. 812, 275–282 (2012).
Aymard, F. et al. Transcriptionally active chromatin recruits homologous recombination at DNA double-strand breaks. Nat. Struct. Mol. Biol. 21, 366–374 (2014).
Joung, J.K. & Sander, J.D. TALENs: a widely applicable technology for targeted genome editing. Nat. Rev. Mol. Cell Biol. 14, 49–55 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Khanna, N., Bian, Q., Plutz, M. & Belmont, A.S. BAC manipulations for making BAC transgene arrays. Methods Mol. Biol. 1042, 197–210 (2013).
Stap, J. et al. Induction of linear tracks of DNA double-strand breaks by alpha-particle irradiation of cells. Nat. Methods 5, 261–266 (2008).
Zotter, A. et al. Recruitment of the nucleotide excision repair endonuclease XPG to sites of UV-induced dna damage depends on functional TFIIH. Mol. Cell Biol. 26, 8868–8879 (2006).
Bekker-Jensen, S. et al. Spatial organization of the mammalian genome surveillance machinery in response to DNA strand breaks. J. Cell Biol. 173, 195–206 (2006).
Kim, J.S., Krasieva, T.B., LaMorte, V., Taylor, A.M. & Yokomori, K. Specific recruitment of human cohesin to laser-induced DNA damage. J. Biol. Chem. 277, 45149–45153 (2002).
Mari, P.O. et al. Dynamic assembly of end-joining complexes requires interaction between Ku70/80 and XRCC4. Proc. Natl. Acad. Sci. USA 103, 18597–18602 (2006).
Roukos, V. et al. Dynamic recruitment of licensing factor Cdt1 to sites of DNA damage. J. Cell Sci. 124, 422–434 (2011).
Haaf, T., Golub, E.I., Reddy, G., Radding, C.M. & Ward, D.C. Nuclear foci of mammalian Rad51 recombination protein in somatic cells after DNA damage and its localization in synaptonemal complexes. Proc. Natl. Acad. Sci. USA 92, 2298–2302 (1995).
Stathopoulou, A. et al. Cdt1 is differentially targeted for degradation by anticancer chemotherapeutic drugs. PLoS ONE 7, e34621 (2012).
Nagy, Z. & Soutoglou, E. DNA repair: easy to visualize, difficult to elucidate. Trends Cell Biol. 19, 617–629 (2009).
Nelms, B.E., Maser, R.S., MacKay, J.F., Lagally, M.G. & Petrini, J.H. In situ visualization of DNA double-strand break repair in human fibroblasts. Science 280, 590–592 (1998).
Polo, S.E. & Jackson, S.P. Dynamics of DNA damage response proteins at DNA breaks: a focus on protein modifications. Genes Dev. 25, 409–433 (2011).
Berkovich, E., Monnat, R.J. Jr. & Kastan, M.B. Assessment of protein dynamics and DNA repair following generation of DNA double-strand breaks at defined genomic sites. Nat. Protoc. 3, 915–922 (2008).
Iacovoni, J.S. et al. High-resolution profiling of gammaH2AX around DNA double strand breaks in the mammalian genome. EMBO J. 29, 1446–1457 (2010).
Sadler, J.R., Sasmor, H. & Betz, J.L. A perfectly symmetric lac operator binds the lac repressor very tightly. Proc. Natl. Acad. Sci. USA 80, 6785–6789 (1983).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Miyanari, Y., Ziegler-Birling, C. & Torres-Padilla, M.E. Live visualization of chromatin dynamics with fluorescent TALEs. Nat. Struct. Mol. Biol. 20, 1321–1324 (2013).
Papapetrou, E.P. & Sadelain, M. Generation of transgene-free human induced pluripotent stem cells with an excisable single polycistronic vector. Nat. Protoc. 6, 1251–1273 (2011).
Jacome, A. & Fernandez-Capetillo, O. Lac operator repeats generate a trac0eable fragile site in mammalian cells. EMBO Rep. 12, 1032–1038 (2011).
Rouet, P., Smih, F. & Jasin, M. Expression of a site-specific endonuclease stimulates homologous recombination in mammalian cells. Proc. Natl. Acad. Sci. USA 91, 6064–6068 (1994).
Masui, O. et al. Live-cell chromosome dynamics and outcome of X chromosome pairing events during ES cell differentiation. Cell 145, 447–458 (2011).
Acknowledgements
Work in the Misteli laboratory is supported by the Intramural Research Program of the US National Institutes of Health (NIH), National Cancer Insitute (NCI), Center for Cancer Research. We thank E. Soutoglou and C. Schmidt for the optimization of specific steps of this protocol, and we thank T. Karpova (NCI Fluorescence Imaging Microscopy Facility) for help with microscopy.
Author information
Authors and Affiliations
Contributions
V.R. developed the protocol. V.R. and R.C.B. wrote the manuscript. T.M. supervised the study and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Optimal size and density of clones.
Representative images of cell colonies that are: (a) smaller than optimal, (b) optimal or (c) colonies plated at a density higher than the desired.
Supplementary Figure 2 Retroviral construct maps and optimal plating density for Plat-GP cells.
A. Retroviral bicistronic vectors designed to express a fusion of Tet repressor (TetR) tagged with the fluorescent protein mCherry along with the puromycin selection marker (a) or the Lac repressor (LacR) fused to GFP along with a neomycin resistance gene (b) via an internal ribosome entry site (IRES) B. Plating density of Platinum-GP cells. Representative images of cells plated at confluencies that are: (a) lower than optimal, (b) optimal for transfection and (c) overconfluent.
Supplementary Figure 3 Cells with high expression of the fluorescent repressors.
High expression of the fluorescent repressors prevents the visualization of LacO/TetO arrays (arrows). Nuclei stained with Hoechst. Scale bar is 10 μm.
Supplementary information
Supplementary Figure 1
Optimal size and density of clones. (PDF 993 kb)
Supplementary Figure 2
Retroviral construct maps and optimal plating density for Plat-GP cells. (PDF 1256 kb)
Supplementary Figure 3
Cells with high expression of the fluorescent repressors. (PDF 1575 kb)
Supplementary Table 1
A list of the plasmids used in this protocol. (PDF 75 kb)
Rights and permissions
About this article
Cite this article
Roukos, V., Burgess, R. & Misteli, T. Generation of cell-based systems to visualize chromosome damage and translocations in living cells. Nat Protoc 9, 2476–2492 (2014). https://doi.org/10.1038/nprot.2014.167
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.167
This article is cited by
-
Control of DNA replication in vitro using a reversible replication barrier
Nature Protocols (2024)
-
A fluorogenic array for temporally unlimited single-molecule tracking
Nature Chemical Biology (2019)
-
Deep Imaging: the next frontier in microscopy
Histochemistry and Cell Biology (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.