Abstract
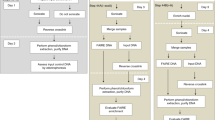
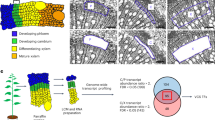
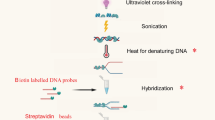
Woody cells and tissues are recalcitrant to standard chromatin immunoprecipitation (ChIP) procedures. However, we recently successfully implemented ChIP in wood-forming tissue of the model woody plant Populus trichocarpa. Here we provide the detailed ChIP protocol optimized for wood-forming tissue that we used in those studies. By using stem-differentiating xylem (SDX; a wood-forming tissue), we identified all steps that were ineffective in standard ChIP protocols and systematically modified them to develop and optimize a robust ChIP protocol. The protocol includes tissue collection, cross-linking, nuclear isolation, chromatin extraction, DNA fragmentation, immunoprecipitation, DNA purification and sequence analysis. The protocol takes 2.5 d to complete and allows a robust 8–10-fold enrichment of transcription factor (TF)–bound genomic fragments (∼150 ng/g of SDX) over nonspecific DNAs. The enriched DNAs are of high quality and can be used for subsequent PCR and DNA-seq analyses. We used this protocol to identify genome-wide specific TF-DNA interactions during wood formation and histone modifications associated with regulation of wood formation. Our protocol, which may be suitable for many tissue types, is so far the only working ChIP system for wood-forming tissue.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Sarkanen, K.V. Renewable resources for the production of fuels and chemicals. Science 191, 773–776 (1976).
Chiang, V.L. From rags to riches. Nat. Biotechnol. 20, 557–558 (2002).
Ragauskas, A.J. et al. The path forward for biofuels and biomaterials. Science 311, 484–489 (2006).
Hinchee, M. et al. Short-rotation woody crops for bioenergy and biofuels applications. In Vitro Cell Dev. Biol. Plant 45, 619–629 (2009).
Evert, R.F. Esau's Plant Anatomy: Meristems, Cells, and Tissues of the Plant Body: Their Structure, Function, and Development 3rd edn. (John Wiley & Sons, 2006).
Li, Q. et al. Splice variant of the SND1 transcription factor is a dominant negative of SND1 members and their regulation in Populus trichocarpa. Proc. Natl. Acad. Sci. USA 109, 14699–14704 (2012).
Lin, Y.-C. et al. SND1 transcription factor–directed quantitative functional hierarchical genetic regulatory network in wood formation in Populus trichocarpa. Plant Cell 25, 4324–4341 (2013).
Solomon, M.J., Larsen, P.L. & Varshavsky, A. Mapping protein–DNA interactions in vivo with formaldehyde: evidence that histone H4 is retained on a highly transcribed gene. Cell 53, 937–947 (1988).
Li, W. et al. DNA methylation and histone modifications regulate de novo shoot regeneration in Arabidopsis by modulating WUSCHEL expression and auxin signaling. PLoS Genet. 7, e1002243 (2011).
Kaufmann, K. et al. Chromatin immunoprecipitation (ChIP) of plant transcription factors followed by sequencing (ChIP-SEQ) or hybridization to whole genome arrays (ChIP-CHIP). Nat. Protoc. 5, 457–472 (2010).
Saleh, A., Alvarez-Venegas, R. & Avramova, Z. An efficient chromatin immunoprecipitation (ChIP) protocol for studying histone modifications in Arabidopsis plants. Nat. Protoc. 3, 1018–1025 (2008).
Bowler, C. et al. Chromatin techniques for plant cells. Plant J. 39, 776–789 (2004).
Zhu, J.Y., Sun, Y. & Wang, Z.Y. Genome-wide identification of transcription factor-binding sites in plants using chromatin immunoprecipitation followed by microarray (ChIP-Chip) or sequencing (ChIP-seq). Plant Signalling Networks: Methods and Protocols (eds. Wang, Z. & Yang, Z.) 173–188 (Humana Press, 2012).
Freudenberg, K. Biosynthesis and constitution of lignin. Nature 183, 1152–1155 (1959).
Lockhart, J. Breaking Down the complex regulatory web underlying lignin biosynthesis. Plant Cell 25, 4282–4282 (2013).
Kim, T.H. & Ren, B. Genome-wide analysis of protein-DNA interactions. Annu. Rev. Genomics Hum. Genet. 7, 81–102 (2006).
Farnham, P.J. Insights from genomic profiling of transcription factors. Nat. Rev. Genet. 10, 605–616 (2009).
Zhou, V.W., Goren, A. & Bernstein, B.E. Charting histone modifications and the functional organization of mammalian genomes. Nat. Rev. Genet. 12, 7–18 (2011).
Margueron, R., Trojer, P. & Reinberg, D. The key to development: interpreting the histone code? Curr. Opin. Genet. Dev. 15, 163–176 (2005).
Pfluger, J. & Wagner, D. Histone modifications and dynamic regulation of genome accessibility in plants. Curr. Opin. Plant Biol. 10, 645–652 (2007).
Nystedt, B. et al. The Norway spruce genome sequence and conifer genome evolution. Nature 497, 579–584 (2013).
Neale, D.B. et al. Decoding the massive genome of loblolly pine using haploid DNA and novel assembly strategies. Genome Biol. 15, R59 (2014).
Amborella Genome Project. The Amborella genome and the evolution of flowering plants. Science 342, 1241089 (2013).
Tuskan, G.A. et al. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313, 1596–1604 (2006).
Myburg, A. et al. The Eucalyptus grandis Genome Project: Genome and transcriptome resources for comparative analysis of woody plant biology. BMC Proc. 5, 20 (2011).
Müller, C.W. Transcription factors: global and detailed views. Curr. Opin. Struct. Biol. 11, 26–32 (2001).
Chen, K. & Rajewsky, N. The evolution of gene regulation by transcription factors and microRNAs. Nat. Rev. Genet. 8, 93–103 (2007).
Hobert, O. Gene regulation by transcription factors and microRNAs. Science 319, 1785–1786 (2008).
Lu, S. et al. Ptr-miR397a is a negative regulator of laccase genes affecting lignin content in Populus trichocarpa. Proc. Natl. Acad. Sci. USA 110, 10848–10853 (2013).
Courtois-Moreau, C.L. et al. A unique program for cell death in xylem fibers of Populus stem. Plant J. 58, 260–274 (2009).
Hu, W. et al. Repression of lignin biosynthesis promotes cellulose accumulation and growth in transgenic trees. Nat. Biotechnol. 17, 808–812 (1999).
Li, L. et al. Combinatorial modification of multiple lignin traits in trees through multigene cotransformation. Proc. Natl. Acad. Sci. USA 100, 4939–4944 (2003).
Lu, S. et al. Novel and mechanical stress-responsive microRNAs in Populus trichocarpa that are absent from Arabidopsis. Plant Cell 17, 2186–2203 (2005).
Shi, R. et al. Towards a systems approach for lignin biosynthesis in Populus trichocarpa: transcript abundance and specificity of the monolignol biosynthetic genes. Plant Cell Physiol. 51, 144–163 (2010).
Mochida, K. et al. TreeTFDB: An integrative database of the transcription factors from six economically important tree crops for functional predictions and comparative and functional genomics. DNA Res. 20, 151–162 (2013).
Jin, J., Zhang, H., Kong, L., Gao, G. & Luo, J. PlantTFDB 3.0: a portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res. 42, D1182–D1187 (2014).
Chen, H. et al. Membrane protein complexes catalyze both 4- and 3-hydroxylation of cinnamic acid derivatives in monolignol biosynthesis. Proc. Natl. Acad. Sci. USA 108, 21253–21258 (2011).
Chen, H. et al. Monolignol pathway 4-coumaric acid:coenzyme A ligases in Populus trichocarpa: novel specificity, metabolic regulation, and simulation of coenzyme A ligation fluxes. Plant Physiol. 161, 1501–1516 (2013).
Acknowledgements
This work was supported by the Office of Science (Biological and Environmental Research), US Department of Energy Grant DE-SC000691 (to V.L.C.). We also thank the support of the North Carolina State University Jordan Family Distinguished Professor Endowment.
Author information
Authors and Affiliations
Contributions
W.L. and Y.-C.L. designed and performed experiments, analyzed data and wrote the paper; Q.L. designed the experiments and antibodies, and edited the manuscript; H.C. validated the specificity of antibodies; R.S., C.-Y.L. and L.C. performed the imaging experiments; R.R.S. and G.-Z.Q. analyzed the data and edited the manuscript; V.L.C. supervised and designed the experiments, analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Optimization of the cross-linking time.
5 g freshly collected SDX were merged into Crosslinking Buffer, and crosslinking performed by vacuum (5 min)/release/mix at room temperature, three times (15 min), six times (30 min) or eight times (40 min). A positive direct target PtrMYB021 of PtrSND1-B16,7 was used. PtrACTIN was used as a negative control. The binding of PtrSND1-B1 to the promoter of PtrMYB021 or PtrACTIN was determined by ChIP using 5 μl of anti-PtrSND1-B1 antibodies followed by detection using ChIP-PCR. Vacuum (5 min)/release/mix at room temperature, six times (30 min) was determined to be optimal. Input, Mock and Anti-B1 are PCR reactions using the chromatin preparations before immunoprecipitation, immunoprecipitated with preimmune serum and immunoprecipitated with anti-PtrSND1-B1 antibody, respectively. Three independent biological replicates of ChIP assays were performed, and the results of one biological replicate are shown. The primers used are described in ref. 7.
Supplementary Figure 2 Microscopic image of isolated nuclei.
Examples of P. trichocarpa SDX nuclei stained with DAPI before adding the nuclei lysis buffer. The scale bar is equivalent to 100 μm.
Supplementary information
Supplementary Figure 1
Optimization of the cross-linking time. (PDF 123 kb)
Supplementary Figure 2
Microscopic image of isolated nuclei. (PDF 116 kb)
Rights and permissions
About this article
Cite this article
Li, W., Lin, YC., Li, Q. et al. A robust chromatin immunoprecipitation protocol for studying transcription factor–DNA interactions and histone modifications in wood-forming tissue. Nat Protoc 9, 2180–2193 (2014). https://doi.org/10.1038/nprot.2014.146
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.146
This article is cited by
-
Cell-type-specific PtrWOX4a and PtrVCS2 form a regulatory nexus with a histone modification system for stem cambium development in Populus trichocarpa
Nature Plants (2023)
-
Protoplast isolation and transcriptome analysis of developing xylem in Pinus massoniana (Pinaceae)
Molecular Biology Reports (2022)
-
High-quality genome assembly of 'Cuiguan' pear (Pyrus pyrifolia) as a reference genome for identifying regulatory genes and epigenetic modifications responsible for bud dormancy
Horticulture Research (2021)
-
ChIP-seq and RNA-seq for complex and low-abundance tree buds reveal chromatin and expression co-dynamics during sweet cherry bud dormancy
Tree Genetics & Genomes (2020)
-
ChIP-Seq reveals that QsMYB1 directly targets genes involved in lignin and suberin biosynthesis pathways in cork oak (Quercus suber)
BMC Plant Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.