Abstract
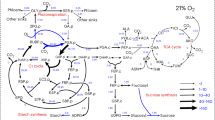
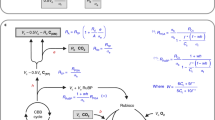
Flux analysis has been carried out in plants for decades, but technical innovations are now enabling it to be carried out in photosynthetic tissues in a more precise fashion with respect to the number of metabolites measured. Here we describe a protocol, using gas chromatography (GC)- and liquid chromatography (LC)-mass spectrometry (MS), to resolve intracellular fluxes of the central carbon metabolism in illuminated intact Arabidopsis thaliana rosettes using the time course of the unlabeled fractions in 40 major constituents of the metabolome after switching to 13CO2. We additionally simplify modeling assumptions, specifically to cope with the presence of multiple cellular compartments. We summarize all steps in this 8–10-week-long process, including setting up the chamber; harvesting; liquid extraction and subsequent handling of sample plant material to chemical derivatization procedures such as silylation and methoxymation (necessary for gas chromatography only); choosing instrumentation settings and evaluating the resultant chromatogram in terms of both unlabeled and labeled peaks. Furthermore, we describe how quantitative insights can be gained by estimating both benchmark and previously unknown fluxes from collected data sets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ong, S.-E. & Mann, M. A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC). Nat. Protoc. 1, 2650–2660 (2006).
Trapnell, C. et al. Differential gene and transcript expression analysis of Rna-Seq experiments with Tophat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Lisec, J., Schauer, N., Kopka, J., Willmitzer, L. & Fernie, A.R. Gas chromatography–mass spectrometry-based metabolite profiling in plants. Nat. Protoc. 1, 387–396 (2006).
Tohge, T. & Fernie, A.R. Combining genetic diversity, informatics and metabolomics to facilitate annotation of plant gene function. Nat. Protoc. 5, 1210–1227 (2010).
Kim, H.K., Choi, Y.H. & Verpoorte, R. NMR-based metabolomic analysis of plants. Nat. Protoc. 5, 536–549 (2010).
Chan, E.C.Y., Pasikanti, K.K. & Nicholson, J.K. Global urinary metabolic profiling procedures using gas chromatography-mass spectrometry. Nat. Protoc. 6, 1483–1499 (2011).
Sellick, C.A., Hansen, R., Stephens, G.M., Goodacre, R. & Dickson, A.J. Metabolite extraction from suspension-cultured mammalian cells for global metabolite profiling. Nat. Protoc. 6, 1241–1249 (2011).
Nielsen, J. It is all about metabolic fluxes. J. Bacteriol. 185, 7031–7035 (2003).
Fernie, A.R., Geigenberger, P. & Stitt, M. Flux an important, but neglected, component of functional genomics. Curr. Opin. Plant Biol. 8, 174–182 (2005).
Poolman, M.G., Miguet, L., Sweetlove, L.J. & Fell, D.A.A Genome-scale metabolic model of Arabidopsis and some of its properties. Plant Physiol. 151, 1570–1581 (2009).
Jensen, P.A. & Papin, J.A. Functional integration of a metabolic network model and expression data without arbitrary thresholding. Bioinformatics 27, 541–547 (2011).
Blazier, A.S. & Papin, J.A. Integration of expression data in genome-scale metabolic network reconstructions. Front. Physiol. 3, 299 (2012).
Toepfer, N. et al. Integration of genome-scale modeling and transcript profiling reveals metabolic pathways underlying light and temperature acclimation in Arabidopsis. Plant Cell 25, 1197–1211 (2013).
Yuan, J., Fowler, W.U., Kimball, E., Lu, W. & Rabinowitz, J.D. Kinetic flux profiling of nitrogen assimilation in Escherichia coli. Nat. Chem. Biol. 2, 529–530 (2006).
Yuan, J., Bennett, B.D. & Rabinowitz, J.D. Kinetic flux profiling for quantitation of cellular metabolic fluxes. Nat. Protoc. 3, 1328–1340 (2008).
Oliver, S.G. Introduction to functional analysis of the yeast genome. Yeast Gene Anal. 26, 1–13 (1998).
Fiehn, O. et al. Metabolite profiling for plant functional genomics. Nat. Biotechnol. 18, 1157–1161 (2000).
Roessner, U. et al. Metabolic profiling allows comprehensive phenotyping of genetically or environmentally modified plant systems. Plant Cell 13, 11–29 (2001).
Fernie, A.R. & Stitt, M. On the discordance of metabolomics with proteomics and transcriptomics: coping with increasing complexity in logic, chemistry, and network interactions. Plant Physiol. 158, 1139–1145 (2012).
Raamsdonk, L.M. et al. A functional genomics strategy that uses metabolome data to reveal the phenotype of silent mutations. Nat. Biotechnol. 19, 45–50 (2001).
Hirai, M.Y. et al. Integration of transcriptomics and metabolomics for understanding of global responses to nutritional stresses in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 101, 10205–10210 (2004).
Sreekumar, A. et al. Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature 457, 910–914 (2009).
De Vos, R.C.H. et al. Untargeted large-scale plant metabolomics using liquid chromatography coupled to mass spectrometry. Nat. Protoc. 2, 778–791 (2007).
Kruger, N.J., Troncoso-Ponce, M.A. & Ratcliffe, R.G. H NMR metabolite fingerprinting and metabolomic analysis of perchloric acid extracts from plant tissues. Nat. Protoc. 3, 1001–1012 (2008).
Patti, G.J., Tautenhahn, R. & Siuzdak, G. Meta-analysis of untargeted metabolomic data from multiple profiling experiments. Nat. Protoc. 7, 508–516 (2012).
Smith, C.A., Want, E.J., O'Maille, G., Abagyan, R. & Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Luedemann, A., Strassburg, K., Erban, A. & Kopka, J. Tagfinder for the quantitative analysis of gas chromatography–mass spectrometry (GC-MS)-based metabolite profiling experiments. Bioinformatics 24, 732–737 (2008).
Lommen, A. MetAlign: interface-driven, versatile metabolomics tool for hyphenated full-scan mass spectrometry data preprocessing. Anal. Chem. 81, 3079–3086 (2009).
Tohge, T. & Fernie, A.R. Web-based resources for mass-spectrometry-based metabolomics: a user's guide. Phytochemistry 70, 450–456 (2009).
Jenkins, H. et al. A proposed framework for the description of plant metabolomics experiments and their results. Nat. Biotechnol. 22, 1601–1606 (2004).
Fiehn, O. et al. Establishing reporting standards for metabolomic and metabonomic studies: a call for participation. OMICS 10, 158–163 (2006).
Sumner, L.W. et al. Proposed minimum reporting standards for chemical analysis. Metabolomics 3, 211–221 (2007).
Fiehn, O. et al. The Metabolomics Standards Initiative (MSI). Metabolomics 3, 175–178 (2007).
Fiehn, O. et al. Quality control for plant metabolomics: reporting MSI-compliant studies. Plant J. 53, 691–704 (2008).
Taylor, C.F. et al. Promoting coherent minimum reporting guidelines for biological and biomedical investigations: the MIBBI project. Nat. Biotechnol. 26, 889–896 (2008).
Fernie, A.R. et al. Recommendations for reporting metabolite data. Plant Cell 23, 2477–2482 (2011).
Matsuda, F. et al. AtMetExpress development: a phytochemical atlas of Arabidopsis development. Plant Physiol. 152, 566–578 (2010).
Kueger, S., Steinhauser, D., Willmitzer, L. & Giavalisco, P. High-resolution plant metabolomics: from mass spectral features to metabolites and from whole-cell analysis to subcellular metabolite distributions. Plant J. 70, 39–50 (2012).
Fernie, A.R. & Morgan, J.A. Analysis of metabolic flux using dynamic labelling and metabolic modelling. Plant Cell Environ. 36, 1738–1750 (2013).
Sweetlove, L.J., Last, R.L. & Fernie, A.R. Predictive metabolic engineering: a goal for systems biology. Plant Physiol. 132, 420–425 (2003).
Schwender, J., Goffman, F., Ohlrogge, J.B. & Shachar-Hill, Y. RuBisCo without the Calvin cycle improves the carbon efficiency of developing green seeds. Nature 432, 779–782 (2004).
Sulpice, R. et al. Starch as a major integrator in the regulation of plant growth. Proc. Natl. Acad. Sci. USA 106, 10348–10353 (2009).
Sulpice, R. et al. Impact of the carbon and nitrogen supply on relationships and connectivity between metabolism and biomass in a broad panel of Arabidopsis accessions. Plant Physiol. 162, 347–363 (2013).
Stitt, M. Metabolic regulation of photosynthesis. In Advances in Photosynthesis; Photosynthesis and the Environment. Vol. 5 (ed. Baker, N.R.) 151–190, Springer (1996).
Arrivault, S. et al. Use of reverse-phase liquid chromatography, linked to tandem mass spectrometry, to profile the Calvin cycle and other metabolic intermediates in Arabidopsis rosettes at different carbon dioxide concentrations. Plant J. 59, 824–839 (2009).
Fernie, A.R., Roessner, U. & Geigenberger, P. The sucrose analog palatinose leads to a stimulation of sucrose degradation and starch synthesis when supplied to discs of growing potato tubers. Plant Physiol. 125, 1967–1977 (2001).
Sweetlove, L.J., Burrell, M.M. & Aprees, T. Starch metabolism in tubers of transgenic potato (Solanum tuberosum) with increased ADPglucose pyrophosphorylase. Biochem. J. 320, 493–498 (1996).
Chen, X. & Shachar-Hill, Y. Insights into metabolic efficiency from flux analysis. J. Exp. Bot. 63, 2343–2351 (2012).
Fernie, A.R. Grand challenges in plant systems biology: closing the circle(s). Front. Plant Sci. 3, 35 (2012).
Stitt, M. Systems-integration of plant metabolism: means, motive and opportunity. Curr. Opin. Plant Biol. 16, 381–388 (2013).
Calvin, M. Path of carbon in photosynthesis. Science 135, 879 (1962).
Kornberg, H.L. Metabolism of C2 compounds in microorganisms.1. incorporation of [2-C-14]acetate by Pseudomonas fluorescens, and by a corynebacterium, grown on ammonium acetate. Biochem. J. 68, 535–542 (1958).
Kornberg, H. & Beevers, H. A mechanism of conversion of fat to carbohydrate in castor beans. Nature 180, 35–36 (1957).
Wheeler, G.L., Jones, M.A. & Smirnoff, N. The biosynthetic pathway of vitamin C in higher plants. Nature 393, 365–369 (1998).
Bao, X.M., Katz, S., Pollard, M. & Ohlrogge, J. Carbocyclic fatty acids in plants: biochemical and molecular genetic characterization of cyclopropane fatty acid synthesis of Sterculia foetida. Proc. Natl. Acad. Sci. USA 99, 7172–7177 (2002).
Gage, D.A. et al. A new route for synthesis of dimethylsulphoniopropionate in marine algae. Nature 387, 891–894 (1997).
Tieman, D. et al. Tomato aromatic amino acid decarboxylases participate in synthesis of the flavor volatiles 2-phenylethanol and 2-phenylacetaldehyde. Proc. Natl. Acad. Sci. USA 103, 8287–8292 (2006).
Dal Cin, V. et al. Identification of genes in the phenylalanine metabolic pathway by ectopic expression of a myb transcription factor in tomato fruit. Plant Cell 23, 2738–2753 (2011).
Araújo, W.L. et al. Identification of the 2-hydroxyglutarate and isovaleryl-CoA dehydrogenases as alternative electron donors linking lysine catabolism to the electron transport chain of Arabidopsis mitochondria. Plant Cell 22, 1549–1563 (2010).
Blackman, F.F. Analytic Studies in Plant Respiration (Cambridge University Press, 1954).
Lipps, S.H. & Beevers, H. Compartmentation of organic acids in corn roots I. differential labeling of 2 malate pools. Plant Physiol. 41, 709–712 (1966).
Gibbs, M. & Beevers, H. Glucose dissimilation in the higher plant. Effect of age of tissue. Plant Physiol. 30, 343–347 (1955).
Beevers, H. & Gibbs, M. The direct oxidation pathway in plant respiration. Plant Physiol. 29, 322–324 (1954).
Ap Rees, T. & Royston, B.J. Control of respiration in disks of carrot storage tissue. Phytochemistry 10, 1199–1206 (1971).
Ap Rees, T., Fuller, W.A. & Wright, B.W. Pathways of carbohydrate oxidation during thermogenesis by the spadix of Arum maculatum. Biochim. Biophys. Acta 437, 22–35 (1976).
Stitt, M., Wirtz, W. & Heldt, H.W. Metabolite levels during induction in the chloroplast and extra-chloroplast compartments of spinach protoplasts. Biochim. Biophys. Acta 593, 85–102 (1980).
Stitt, M., Gerhardt, R., Kurzel, B. & Heldt, H.W. A role for fructose 2,6-bisphosphate in the regulation of sucrose synthesis in spinach leaves. Plant Physiol. 72, 1139–1141 (1983).
Fricke, W., Hinde, P.S., Leigh, R.A. & Tomos, A.D. Vacuolar solutes in the upper epidermis of barley leaves - intercellular differences follow patterns. Planta 196, 40–49 (1995).
Koroleva, O.A., Farrar, J.F., Tomos, A.D. & Pollock, C.J. Carbohydrates in individual cells of epidermis, mesophyll, and bundle sheath in barley leaves with changed export or photosynthetic rate. Plant Physiol. 118, 1525–1532 (1998).
Wiechert, W. C-13 metabolic flux analysis. Metab. Eng. 3, 195–206 (2001).
Noeh, K. & Wiechert, W. Experimental design principles for isotopically instationary 13C labeling experiments. Biotechnol. Bioeng. 94, 234–251 (2006).
Szecowka, M. et al. Metabolic fluxes in an illuminated Arabidopsis rosette. Plant Cell 25, 694–714 (2013).
Ratcliffe, R.G. & Shachar-Hill, Y. Measuring multiple fluxes through plant metabolic networks. Plant J. 45, 490–511 (2006).
Schmidt, K., Nielsen, J. & Villadsen, J. Quantitative anaylsis of metabolic fluxes in Escherichia coli, using two-dimensional NMR spectroscopy and complete isotopomer models. J. Biotechnol. 71, 175–190 (1999).
Ruehl, M. et al. Collisional fragmentation of central carbon metabolites in LC-MS/MS increases precision of 13C metabolic flux analysis. Biotechnol. Bioeng. 109, 763–771 (2012).
Schwender, J., Shachar-Hill, Y. & Ohlrogge, J.B. Mitochondrial metabolism in developing embryos of Brassica napus. J. Biol. Chem. 281, 34040–34047 (2006).
Alonso, A.P., Goffman, F.D., Ohlrogge, J.B. & Shachar-Hill, Y. Carbon conversion efficiency and central metabolic fluxes in developing sunflower (Helianthus annuus L.) embryos. Plant J. 52, 296–308 (2007).
Masakapalli, S.K. et al. Subcellular flux analysis of central metabolism in a heterotrophic Arabidopsis cell suspension using steady-state stable isotope labeling. Plant Physiol. 152, 602–619 (2010).
Williams, T.C.R. et al. A genome-scale metabolic model accurately predicts fluxes in central carbon metabolism under stress conditions. Plant Physiol. 154, 311–323 (2010).
Rontein, D., Dieuaide-Noubhani, M., Dufourc, E.J., Raymond, P. & Rolin, D. The metabolic architecture of plant cells: stability of central metabolism and flexibility of anabolic pathways during the growth cycle of tomato cells. J. Biol. Chem. 277, 43948–43960 (2002).
Kruger, N.J., Huddlestone, J.E., Le Lay, P., Brown, N.D. & Ratcliffe, R.G. Network flux analysis: impact Of 13C substrates on metabolism in Arabidopsis thaliana cell suspension cultures. Phytochemistry 68, 2176–2188 (2007).
Sriram, G. et al. Quantification of compartmented metabolic fluxes in developing soybean embryos by employing biosynthetic Ally directed fractional 13C labeling, [13C, 1H] two-dimensional nuclear magnetic resonance, and comprehensive isotopomer balancing. Plant Physiol. 136, 3043–3057 (2004).
Allen, D.K., Ohlrogge, J.B. & Shachar-Hill, Y. The role of light in soybean seed filling metabolism. Plant J. 58, 220–234 (2009).
Alonso, A.P., Val, D.L. & Shachar-Hill, Y. Central metabolic fluxes in the endosperm of developing maize seeds and their implication for metabolic engineering. Metab. Eng. 13, 96–107 (2011).
Geigenberger, P., Stitt, M. & Fernie, A.R. Metabolic control analysis and regulation of the conversion of sucrose to starch in growing potato tubers. Plant Cell Environ. 27, 655–673 (2004).
Roessner-Tunali, U. et al. Kinetics of labelling of organic and amino acids in potato tubers by gas chromatography–mass spectrometry following incubation in 13C labelled isotopes. Plant J. 39, 668–679 (2004).
Roscher, A., Kruger, N.J. & Ratcliffe, R.G. Strategies for metabolic flux analysis in plants using isotope labelling. J. Biotechnol. 77, 81–102 (2000).
Hasunuma, T. et al. Metabolic turnover analysis by a combination of in vivo C-13-labelling from 13CO2 and metabolic profiling with CE-MS/MS reveals rate-limiting steps of the C3 photosynthetic pathway in Nicotiana tabacum leaves. J. Exp. Bot. 61, 1041–1051 (2010).
Noeh, K. et al. Metabolic flux analysis at ultrashort time scale: isotopically non-stationary 13C labeling experiments. J. Biotechnol. 129, 249–267 (2007).
Noack, S., Noeh, K., Moch, M., Oldiges, M. & Wiechert, W. Stationary versus non-stationary 13C-MFA: a comparison using a consistent dataset. J. Biotechnol. 154, 179–190 (2011).
Shastri, A.A. & Morgan, J.A. A transient isotopic labeling methodology for 13C metabolic flux analysis of photo auto trophic microorganisms. Phytochemistry 68, 2302–2312 (2007).
Wiechert, W. & Noeh, K. Isotopically non-stationary metabolic flux analysis: complex yet highly informative. Curr. Opin. Biotechnol. 24, 979–986 (2013).
Antoniewicz, M.R. et al. Metabolic flux analysis in a nonstationary system: fed-batch fermentation of a high yielding strain of E. coli producing 1,3-propanediol. Metab. Eng. 9, 277–292 (2007).
Hörl, M., Schnidder, J., Sauer, U. & Zamboni, N. Non-stationary 13C-metabolic flux ratio analysis. Biotechnol. Bioeng. 12, 3164–3177 (2013).
Antoniewicz, M.R., Kelleher, J.K. & Stephanopoulos, G. Elementary metabolite units (EMU): a novel framework for modeling isotopic distributions. Metab. Eng. 9, 68–86 (2007).
Klie, S. et al. Analysis of the compartmentalized metabolome—a validation of the non-aqueous fractionation technique. Front. Plant Sci. 2, 55 (2011).
Pal, S.K. et al. Diurnal changes of polysome loading track sucrose content in the rosette of wild-type Arabidopsis and the starchless pgm mutant. Plant Physiol. 162, 1246–1265 (2013).
Boyes, D.C. et al. Growth stage-based phenotypic analysis of Arabidopsis: a model for high throughput functional genomics in plants. Plant Cell 13, 1499–1510 (2001).
Badger, M.R., Sharkey, T.D. & Von Caemmerer, S. The relationship between steady-state gas exchange of bean leaves and the levels of carbon-reduction cycle intermediates. Planta 160, 305–313 (1984).
Ruuska, S.A. et al. The interplay between limiting processes in C3 photosynthesis studied by rapid-response gas exchange using transgenic tobacco impaired in photosynthesis. Austral. J. Plant Physiol. 25, 859–887 (1998).
Lunn, J.E. et al. Sugar-induced increases in trehalose 6-phosphate are correlated with redox activation of ADPglucose pyrophosphorylase and higher rates of starch synthesis in Arabidopsis thaliana. Biochem. J. 397, 139–148 (2006).
Cohen, S.D. & Hindmarsh, A.C. CVODE, a stiff/nonstiff ODE solver in C. Computers Phys. 10, 138–143 (1996).
Rowan,, T. Functional stability analysis of numerical algorithms. Ph.D. Thesis, Department of Computer Sciences, University Of Texas at Austin (1990).
Tohge, T., Mettler, T., Arrivault, S., Carroll, A.J., Stitt, M. & Fernie, A.R. From models to crop species: caveats and solutions for translational metabolomics. Front. Plant Sci. 2, 61 (2011).
Gerhardt, R. & Heldt, H.W. Measurement of subcellular metabolite levels in leaves by fractionation of freeze-stopped material in nonaqueous media. Plant Physiol. 75, 542–547 (1984).
Stitt, M., Lilley, R.M., Gerhardt, R. & Heldt, H.W. Metabolite levels in specific cells and subcellular compartments of plant leaves. Methods Enzymol. 174, 518–552 (1989).
Riens, B., Lohaus, G., Heineke, D. & Heldt, H.W. Amino acid and sucrose content determined in the cytosolic, chloroplastic, and vacuolar compartments and in the phloem sap of spinach leaves. Plant Physiol. 97, 227–233 (1991).
Lawson, C.L. & Hanson, R.J. Solving least-squares problems. in Classics Appl. Math. (Siam, 1995).
Schuster, R. & Schuster, S. Refined algorithm and computer-program for calculating all nonnegative fluxes admissible in steady-states of biochemical reaction systems with or without some flux rates fixed. Comp. Appl. Biosci. 9, 79–85 (1993).
Wiechert, W., Mollney, M., Isermann, N., Wurzel, W. & De Graaf, A.A. Bidirectional reaction steps in metabolic networks: III. Explicit solution and analysis of isotopomer labeling systems. Biotechnol. Bioeng. 66, 69–85 (1999).
Acknowledgements
We thank J. Lunn, R. Feil and J. Huege for useful discussions and H. Ishihara and C. Abel for help with the video. We also thank B.O. Hansen for help in formatting.
Author information
Authors and Affiliations
Contributions
R.H., M. Szecowka, S.A., T.T., A.N.-N., M. Stitt, Z.N. and A.R.F. contributed sections to the manuscript. M. Stitt, Z.N. and A.R.F. collated and standardized the text. All authors read and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Histograms of Monte Carlo simulation.
Frequencies of selected parameters φ of the Monte-Carlo simulation including 500 repetitions. The parameters φ describe to steady-state net flux distribution according to Equation (7) (also see Supplementary Note 1). The selected parameters correspond to (A) φ(1) amount of photorespiration, (B) φ(2) amount of starch synthesis, (C) φ(3) amount of sucrose synthesis and (D) φ(4) amount of trehalse synthesis. The corresponding modes M1 - M4 are listed in Supplementary Data 2. The arrow indicates the optimal value and the dashed lines border the 95% confindence intervals.
Supplementary information
Supplementary Figure 1
Histograms of Monte Carlo simulation. (PDF 66 kb)
Supplementary Table 1
Metabolic contents used for flux estimation. (PDF 68 kb)
Supplementary Table 2
Parameters of the input models. (PDF 67 kb)
Supplementary Table 3
Estimates of fluxes. (PDF 68 kb)
Supplementary Table 4
Estimates of inactive fractions. (PDF 52 kb)
Supplementary Data 1
Amounts for phosphorylated metabolites measured by LC-MS/MS (in pmol gFW-1). (XLSX 18 kb)
Supplementary Data 2
The pathway model and its modes. (PDF 107 kb)
Supplementary Equations
System of ordinary differential Equations. (PDF 74 kb)
Supplementary Note 1
Implementation of the model. (PDF 119 kb)
Supplementary Note 2
Decomposition. (PDF 66 kb)
Supplementary Video 1
Quenching of Arabidopsis. (MP4 18131 kb)
Rights and permissions
About this article
Cite this article
Heise, R., Arrivault, S., Szecowka, M. et al. Flux profiling of photosynthetic carbon metabolism in intact plants. Nat Protoc 9, 1803–1824 (2014). https://doi.org/10.1038/nprot.2014.115
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.115
This article is cited by
-
Novel guard cell sink characteristics revealed by a multi-species/cell-types meta-analysis of 13C-labelling experiments
Theoretical and Experimental Plant Physiology (2024)
-
Recent advances in proteomics and metabolomics in plants
Molecular Horticulture (2022)
-
Design and fabrication of an improved dynamic flow cuvette for 13CO2 labeling in Arabidopsis plants
Plant Methods (2022)
-
An integrated isotopic labeling and freeze sampling apparatus (ILSA) to support sampling leaf metabolomics at a centi-second scale
Plant Methods (2022)
-
Stochastic simulation of enzymatic kinetics for 13C isotope labeling at the single-cell scale
Reaction Kinetics, Mechanisms and Catalysis (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.