Abstract
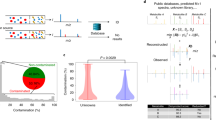
Data processing for 1D NMR spectra is a key bottleneck for metabolomic and other complex-mixture studies, particularly where quantitative data on individual metabolites are required. We present a protocol for automated metabolite deconvolution and quantification from complex NMR spectra by using the Bayesian automated metabolite analyzer for NMR (BATMAN) R package. BATMAN models resonances on the basis of a user-controllable set of templates, each of which specifies the chemical shifts, J-couplings and relative peak intensities for a single metabolite. Peaks are allowed to shift position slightly between spectra, and peak widths are allowed to vary by user-specified amounts. NMR signals not captured by the templates are modeled non-parametrically by using wavelets. The protocol covers setting up user template libraries, optimizing algorithmic input parameters, improving prior information on peak positions, quality control and evaluation of outputs. The outputs include relative concentration estimates for named metabolites together with associated Bayesian uncertainty estimates, as well as the fit of the remainder of the spectrum using wavelets. Graphical diagnostics allow the user to examine the quality of the fit for multiple spectra simultaneously. This approach offers a workflow to analyze large numbers of spectra and is expected to be useful in a wide range of metabolomics studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Sreekumar, A. et al. Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature 457, 910–914 (2009).
Holmes, E. et al. Human metabolic phenotype diversity and its association with diet and blood pressure. Nature 453, 396–400 (2008).
Weckwerth, W. et al. Differential metabolic networks unravel the effects of silent plant phenotypes. Proc. Natl. Acad. Sci. USA 101, 7809–7814 (2004).
Lindon, J.C. et al. The Consortium for Metabonomic Toxicology (COMET): aims, activities and achievements. Pharmacogenomics 6, 691–699 (2005).
Nicholson, J.K. & Wilson, I.D. High-resolution proton magnetic-resonance spectroscopy of biological-fluids. Prog. Nucl. Magn. Reson. Spectrosc. 21, 449–501 (1989).
Fan, T.W.M. Metabolite profiling by one- and two-dimensional NMR analysis of complex mixtures. Prog. Nucl. Magn. Reson. Spectrosc. 28, 161–219 (1996).
Zhang, S. et al. Advances in NMR-based biofluid analysis and metabolite profiling. Analyst 135, 1490–1498 (2010).
Beckonert, O. et al. Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nat. Protoc. 2, 2692–2703 (2007).
Ebbels, T.M.D. & Cavill, R. Bioinformatic methods in NMR-based metabolic profiling. Prog. Nucl. Magn. Reson. Spectrosc. 55, 361–374 (2009).
Ebbels, T.M.D. & De Iorio, M. Statistical data analysis in metabolomics. in Handbook of Statistical Systems Biology (eds. Girolami, M.A., Balding, D. & Stumpf, M.) (Thompson Digital, 2011).
Wishart, D.S. et al. HMDB: the Human Metabolome Database. Nucleic Acids Res. 35 (Database issue), D521–D526 (2007).
Ulrich, E.L. et al. BioMagResBank. Nucleic Acids Res. 36 (Database issue), D402–D408 (2008).
Holmes, E. et al. Automatic data reduction and pattern recognition methods for analysis of 1H nuclear magnetic resonance spectra of human urine from normal and pathological states. Anal. Biochem. 220, 284 (1994).
Spraul, M. et al. Automatic reduction of NMR spectroscopic data for statistical and pattern recognition classification of samples. J. Pharm. Biomed. Anal. 12, 1215–1225 (1994).
Cloarec, O. et al. Evaluation of the orthogonal projection on latent structure model limitations caused by chemical shift variability and improved visualization of biomarker changes in 1H NMR spectroscopic metabonomic studies. Anal. Chem. 77, 517 (2005).
Csenki, L. et al. Proof of principle of a generalized fuzzy Hough transform approach to peak alignment of one-dimensional 1H NMR data. Anal. Bioanal. Chem. 389, 875–885 (2007).
Alm, E. et al. A solution to the 1D NMR alignment problem using an extended generalized fuzzy Hough transform and mode support. Anal. Bioanal. Chem. 395, 213–223 (2009).
Astle, W. et al. A bayesian model of NMR spectra for the deconvolution and quantification of metabolites in complex biological mixtures. J. Am. Stat. Assoc. 107, 1259–1271 (2012).
Hao, J. et al. BATMAN—an R package for the automated quantification of metabolites from nuclear magnetic resonance spectra using a Bayesian model. Bioinformatics 28, 2088–2090 (2012).
Liebeke, M. et al. Combining spectral ordering with peak fitting for one-dimensional NMR quantitative metabolomics. Anal. Chem. 85, 4605–4612 (2013).
Reily, M.D. et al. DFTMP, an NMR reagent for assessing the near-neutral pH of biological samples. J. Am. Chem. Soc. 128, 12360–12361 (2006).
Behrends, V. et al. Metabolite profiling to characterize disease-related bacteria: gluconate excretion by Pseudomonas aeruginosa mutants and clinical isolates from cystic fibrosis patients. J. Biol. Chem. 288, 15098–15109 (2013).
Zheng, C. et al. Identification and quantification of metabolites in (1)H NMR spectra by Bayesian model selection. Bioinformatics 27, 1637–1644 (2011).
Weljie, A.M. et al. Targeted profiling: quantitative analysis of 1H NMR metabolomics data. Anal. Chem. 78, 4430–4442 (2006).
Tredwell, G.D. et al. Between-person comparison of metabolite fitting for NMR-based quantitative metabolomics. Anal. Chem. 83, 8683–8687 (2011).
Sokolenko, S. et al. Understanding the variability of compound quantification from targeted profiling metabolomics of 1D-1H-NMR spectra in synthetic mixtures and urine with additional insights on choice of pulse sequences and robotic sampling. Metabolomics 9, 887–903 (2013).
Slupsky, C.M. et al. Investigations of the effects of gender, diurnal variation, and age in human urinary metabolomic profiles. Anal. Chem. 79, 6995–7004 (2007).
Viant, M.R. et al. International NMR-based environmental metabolomics intercomparison exercise. Environ. Sci. Technol. 43, 219–225 (2009).
Acknowledgements
T.M.D.E., M.D.I., W.A. and J.H. acknowledge support from the Biotechnology and Biological Sciences Research Council (BBSRC), UK, grant BB/E20372/1. J.G.B., T.M.D.E., J.H. and M.L. acknowledge support from the Natural Environment Research Council (NERC), UK, grant NE/H009973/1. T.M.D.E. and J.H. acknowledge support from the European Commission COSMOS project, contract EC312941. We thank the eMICE consortium for use of the synthetic mixture spectra.
Author information
Authors and Affiliations
Contributions
T.M.D.E. and M.D.I. conceived the project and supervised the development of the Bayesian model. W.A. developed the Bayesian model and J.H. implemented the corresponding R package. J.G.B. and M.L. provided theoretical and practical input on NMR spectroscopy, including software testing and ideas for new features. T.M.D.E. and J.H. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Data
Comparison of BATMAN and Chenomx NMR Suite results. (PDF 805 kb)
Rights and permissions
About this article
Cite this article
Hao, J., Liebeke, M., Astle, W. et al. Bayesian deconvolution and quantification of metabolites in complex 1D NMR spectra using BATMAN. Nat Protoc 9, 1416–1427 (2014). https://doi.org/10.1038/nprot.2014.090
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.090
This article is cited by
-
Problems, principles and progress in computational annotation of NMR metabolomics data
Metabolomics (2022)
-
Metabolic alterations in meningioma reflect the clinical course
BMC Cancer (2021)
-
Metabolomic signatures in elite cyclists: differential characterization of a seeming normal endocrine status regarding three serum hormones
Metabolomics (2021)
-
Targeting redox metabolism: the perfect storm induced by acrylamide poisoning in the brain
Scientific Reports (2020)
-
A comparison of high-throughput plasma NMR protocols for comparative untargeted metabolomics
Metabolomics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.