Abstract
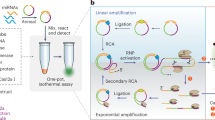
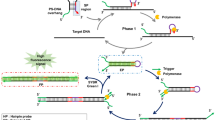
This protocol describes an isothermal amplification approach for ultrasensitive detection of specific microRNAs (miRNAs). It achieves this level of sensitivity through quadratic amplification of the target oligonucleotide by using a Bst DNA polymerase–induced strand-displacement reaction and a lambda exonuclease–aided recycling reaction. First, the target miRNA binds to a specifically designed molecular beacon, causing it to become a fluorescence emitter. A primer then binds to the activated beacon, and Bst polymerase initiates the synthesis of a double-stranded DNA segment templated on the molecular beacon. This causes the concomitant release of the target miRNA from the beacon—the first round of 'recycling'. Second, the duplex beacon thus produced is a suitable substrate for a nicking enzyme present in solution. After the duplex beacon is nicked, the lambda exonuclease digests the beacon and releases the DNA single strand just synthesized, which is complementary to the molecular beacon, inducing the second round of recycling. The miRNA detection limit of this protocol is 10 fmol at 37 °C and 1 amol at 4 °C. This approach also affords high selectivity when applied to miRNA extracted from MCF-7 and PC3 cell lines and even from breast cancer tissue samples. Upon isolation of miRNA, the detection process can be completed in ∼2 h.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
He, L. & Hannon, G.J. MicroRNAs: small RNAs with a big role in gene regulation. Nat. Rev. Genet. 5, 522–531 (2004).
Válóczi, A. et al. Sensitive and specific detection of microRNAs by northern blot analysis using LNA-modified oligonucleotide probes. Nucleic Acids Res. 32, e175 (2004).
Lee, J., Cho, H. & Jung, Y. Fabrication of a structure-specific RNA binder for array detection of label-free microRNA. Angew. Chem. Int. Ed. 49, 8662–8665 (2010).
Lee, I. et al. Discriminating single-base difference miRNA expressions using microarray Probe Design Guru (ProDeG). Nucleic Acids Res. 36, e27 (2008).
Chen, C. et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 33, e179 (2005).
Li, J. et al. Real-time polymerase chain reaction microRNA detection based on enzymatic stem-loop probes ligation. Anal. Chem. 81, 5446–5451 (2009).
Jia, H., Li, Z., Liu, C. & Cheng, Y. Ultrasensitive detection of microRNAs by exponential isothermal amplification. Angew. Chem. Int. Ed. 49, 5498–5501 (2010).
Song, S. et al. Gold-nanoparticle-based multicolor nanobeacons for sequence-specific DNA analysis. Angew. Chem. Int. Ed. 48, 1–5 (2009).
Wang, K.M. et al. Molecular engineering of DNA: molecular beacons. Angew. Chem. Int. Ed. 948, 856–870 (2009).
Wei, F. et al. Electrochemical detection of low-copy-number salivary RNA based on specific signal amplification with a hairpin probe. Nucleic Acids Res. 36, e65 (2008).
Zhang, X., Wang, Z., Xing, H., Xiang, Y. & Lu, Y. Catalytic and molecular beacons for amplied detection of metal ions and organic molecules with high sensitivity. Anal. Chem. 82, 5005–5011 (2010).
Li, J., Chu, Y., Lee, B.Y.H. & Xie, X.S. Enzymatic signal amplification of molecular beacons for sensitive DNA detection. Nucleic Acids Res. 36, e36 (2008).
Kiesling, T. et al. Sequence-specific detection of DNA using nicking endonuclease signal amplification (NESA). Nucleic Acids Res. 35, e117 (2007).
Van Ness, J., Van Ness, L.K. & Galas, D.J. Isothermal reactions for the amplification of oligonucleotides. Proc. Natl. Acad. Sci. USA 100, 4504–4509 (2003).
Hosoda, K. et al. A novel sequence-specific RNA quantification method using nicking endonuclease, dual-labeled fluorescent DNA probe, and conformation-interchangeable oligo-DNA. RNA 14, 584–592 (2008).
Zuo, X., Xia, F., Xiao, Y. & Plaxco, K.W. Sensitive and selective amplified fluorescence DNA detection based on exonuclease III-aided target recycling. J. Am. Chem. Soc. 132, 1816–1818 (2010).
Freeman, R., Liu, X. & Willner, I. Amplified multiplexed analysis of DNA by the exonuclease III-catalyzed regeneration of the target DNA in the presence of functionalized semiconductor quantum dots. Nano Lett. 11, 4456–4461 (2011).
Zuo, X., et al. Two-step, PCR-free telomerase detection using exonuclease III–aided target recycling. Chembiochem. 12, 2745–2747 (2011).
Connolly, A.R. & Trau, M. Isothermal detection of DNA by beacon-assisted detection amplification. Angew. Chem. Int. Ed. 49, 2720–2723 (2010).
Guo, Q. et al. Sensitive fluorescence detection of nucleic acids based on isothermal circular strand-displacement polymerization reaction. Nucleic Acids Res. 37, e20 (2009).
Connolly, A.R. & Trau, M. Rapid DNA detection by beacon-assisted detection amplification. Nat. Protoc. 6, 772–778 (2011).
Fang, S., Lee, H.J., Wark, A.W. & Corn, R.M. Attomole microarray detection of microRNAs by nanoparticle-amplified SPR imaging measurements of Surface polyadenylation reactions. J. Am. Chem. Soc. 128, 14044–14046 (2006).
Li, J., Schachermeyer, S., Wang, Y., Yin, Y. & Zhong, W. Detection of microRNA by fluorescence amplification based on cation-exchange in nanocrystals. Anal. Chem. 81, 9723–9729 (2009).
Zhou, Y. et al. A dumbbell probe-mediated rolling-circle amplification strategy for highly sensitive microRNA detection. Nucleic Acids Res. 38, e156 (2010).
Cheng, Y. et al. Highly sensitive determination of microRNA using target-primed and branched rolling-circle amplification. Angew. Chem. Int. Ed. 121, 3318–3322 (2009).
Duan, R. et al. Lab in a tube: ultrasensitive detection of microRNAs at the single-cell level and in breast cancer patients using quadratic isothermal amplification. J. Am. Chem. Soc. 135, 4604–4607 (2013).
Acknowledgements
This research was supported by initiatory financial support from HUST, the National Basic Research Program of China (2013CB933000), the 100 Talents Program from the CAS, the Shanghai Pujiang Project (grant no. 13PJ1410700 to X. Z.), the 1,000 Young Talents program (to F. X.) and the National Natural Science Foundation (21375042).
Author information
Authors and Affiliations
Contributions
R.D., F.X., X.Z., L.J., S.W., Z.C., C.F., X.Q. and D.C. conceived the projects, designed and conducted experiments, analyzed the data and wrote the manuscript. R.D., X.Z. and S.W. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Duan, R., Zuo, X., Wang, S. et al. Quadratic isothermal amplification for the detection of microRNA. Nat Protoc 9, 597–607 (2014). https://doi.org/10.1038/nprot.2014.036
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.036
This article is cited by
-
Isothermal exponential amplification reactions triggered by circular templates (cEXPAR) targeting miRNA
Molecular Biology Reports (2023)
-
Concentration-Dependent Study of Nucleic Acid Blockers Used for Sequence-Specificity Enhancement in Nucleic Acids Detection
Arabian Journal for Science and Engineering (2023)
-
Label-free detection of microRNA: two-stage signal enhancement with hairpin assisted cascade isothermal amplification and light-up DNA-silver nanoclusters
Microchimica Acta (2020)
-
Ultrasensitive detection of cancer biomarker microRNA by amplification of fluorescence of lanthanide nanoprobes
Nano Research (2018)
-
Multiplex quantitative analysis of microRNA expression via exponential isothermal amplification and conformation-sensitive DNA separation
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.