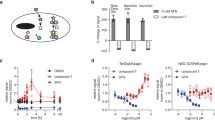
Abstract
Coimmunoprecipitation (co-IP) analysis is a useful method for studying protein-protein interactions. It currently involves electrophoresis and western blotting, which are not optimized for detecting weak and transient interactions. In this protocol we describe an advanced version of co-IP analysis that uses real-time, single-molecule fluorescence imaging as its detection scheme. Bait proteins are pulled down onto the imaging plane of a total internal reflection (TIR) microscope. With unpurified cells or tissue extracts kept in reaction chambers, we observe single protein-protein interactions between the surface-immobilized bait and the fluorescent protein–labeled prey proteins in real time. Such direct recording provides an improvement of five orders of magnitude in the time resolution of co-IP analysis. With the single-molecule sensitivity and millisecond time resolution, which distinguish our method from other methods for measuring weak protein-protein interactions, it is possible to quantify the interaction kinetics and active fraction of native, unlabeled bait proteins. Real-time single-molecule co-IP analysis, which takes ∼4 h to complete from lysate preparation to kinetic analysis, provides a general avenue for revealing the rich kinetic picture of target protein-protein interactions, and it can be used, for example, to investigate the molecular lesions that drive individual cancers at the level of protein-protein interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Sambrook, J. & Russell, D.W. Molecular Cloning: a Laboratory Manual 3rd edn. (Cold Spring Harbor Laboratory Press, 2001).
Lee, H.W. et al. Real-time single-molecule co-immunoprecipitation analyses reveal cancer-specific Ras signalling dynamics. Nat. Commun. 4, 1505 (2013).
Jain, A. et al. Probing cellular protein complexes using single-molecule pull-down. Nature 473, 484–488 (2011).
Yeom, K.H. et al. Single-molecule approach to immunoprecipitated protein complexes: insights into miRNA uridylation. EMBO Rep. 12, 690–696 (2011).
Morelle, W. et al. Mass spectrometric approach for screening modifications of total serum N-glycome in human diseases: application to cirrhosis. Glycobiology 16, 281–293 (2006).
Lemmon, M.A. & Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 141, 1117–1134 (2010).
Inoue, T., Heo, W.D., Grimley, J.S., Wandless, T.J. & Meyer, T. An inducible translocation strategy to rapidly activate and inhibit small GTPase signaling pathways. Nat. Methods 2, 415–418 (2005).
Lee, K.H., Lee, S., Lee, W.Y., Yang, H.W. & Heo, W.D. Visualizing dynamic interaction between calmodulin and calmodulin-related kinases via a monitoring method in live mammalian cells. Proc. Natl. Acad. Sci. USA 107, 3412–3417 (2010).
Soderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006).
Nilsson, I. et al. VEGF receptor 2/-3 heterodimers detected in situ by proximity ligation on angiogenic sprouts. EMBO J. 29, 1377–1388 (2010).
Weinberg, R.A. The Biology of Cancer (Garland Science, 2007).
Wyer, J.R. et al. T cell receptor and coreceptor CD8αα bind peptide-MHC independently and with distinct kinetics. Immunity 10, 219–225 (1999).
Garrus, J.E. et al. Tsg101 and the vacuolar protein sorting pathway are essential for HIV-1 budding. Cell 107, 55–65 (2001).
Burman, J.D. et al. Interaction of human complement with Sbi, a staphylococcal immunoglobulin-binding protein: indications of a novel mechanism of complement evasion by Staphylococcus aureus. J. Biol. Chem. 283, 17579–17593 (2008).
Homola, J., Yee, S.S. & Gauglitz, G. Surface plasmon resonance sensors: review. Sens. Actuators B Chem. 54, 3–15 (1999).
Joo, C., Balci, H., Ishitsuka, Y., Buranachai, C. & Ha, T. Advances in single-molecule fluorescence methods for molecular biology. Annu. Rev. Biochem. 77, 51–76 (2008).
Kim, E. et al. A single-molecule dissection of ligand binding to a protein with intrinsic dynamics. Nat. Chem. Biol. 9, 313–318 (2013).
Roy, R., Hohng, S. & Ha, T. A practical guide to single-molecule FRET. Nat. Methods 5, 507–516 (2008).
Jain, A., Liu, R., Xiang, Y.K. & Ha, T. Single-molecule pull-down for studying protein interactions. Nat. Protoc. 7, 445–452 (2012).
Hancock, J.F. Ras proteins: different signals from different locations. Nat. Rev. Mol. Cell Biol. 4, 373–384 (2003).
Larson, D.R., Zenklusen, D., Wu, B., Chao, J.A. & Singer, R.H. Real-time observation of transcription initiation and elongation on an endogenous yeast gene. Science 332, 475–478 (2011).
Adams, S.R. et al. New biarsenical ligands and tetracysteine motifs for protein labeling in vitro and in vivo: synthesis and biological applications. J. Am. Chem. Soc. 124, 6063–6076 (2002).
Juillerat, A. et al. Directed evolution of O6-alkylguanine-DNA alkyltransferase for efficient labeling of fusion proteins with small molecules in vivo. Chem. Biol. 10, 313–317 (2003).
Shi, X. et al. Quantitative fluorescence labeling of aldehyde-tagged proteins for single-molecule imaging. Nat. Methods 9, 499–503 (2012).
Xie, S.N. Single-molecule approach to enzymology. Single Mol. 2, 229–236 (2001).
Selvin, P.R. & Ha, T. Single-Molecule Techniques: a Laboratory Manual (Cold Spring Harbor Laboratory Press, 2008).
Yanagida, T. & Ishii, Y. Single Molecule Dynamics in Life Science (Wiley, 2009).
Sako, Y. & Ueda, M. Cell Signaling Reactions: Single-molecular Kinetic Analysis (Springer, 2011).
Dickson, R.M., Cubitt, A.B., Tsien, R.Y. & Moerner, W.E. On/off blinking and switching behaviour of single molecules of green fluorescent protein. Nature 388, 355–358 (1997).
Ha, T. & Tinnefeld, P. Photophysics of fluorescent probes for single-molecule biophysics and super-resolution imaging. Annu. Rev. Phys. Chem. 63, 595–617 (2012).
Schubbert, S., Shannon, K. & Bollag, G. Hyperactive Ras in developmental disorders and cancer. Nat. Rev. Cancer 7, 295–308 (2007).
Sydor, J.R., Engelhard, M., Wittinghofer, A., Goody, R.S. & Herrmann, C. Transient kinetic studies on the interaction of Ras and the Ras-binding domain of c-Raf-1 reveal rapid equilibration of the complex. Biochemistry 37, 14292–14299 (1998).
Gorman, C., Skinner, R.H., Skelly, J.V., Neidle, S. & Lowe, P.N. Equilibrium and kinetic measurements reveal rapidly reversible binding of Ras to Raf. J. Biol. Chem. 271, 6713–6719 (1996).
Nassar, N. et al. The 2.2 Å crystal structure of the Ras-binding domain of the serine/threonine kinase c-Raf1 in complex with Rap1A and a GTP analogue. Nature 375, 554–560 (1995).
Chuang, E. et al. Critical binding and regulatory interactions between Ras and Raf occur through a small, stable -termina N-terminal domain of Raf and specific Ras effector residues. Mol. Cell. Biol. 14, 5318–5325 (1994).
Der, C.J., Finkel, T. & Cooper, G.M. Biological and biochemical properties of human rasH genes mutated at codon 61. Cell 44, 167–176 (1986).
Mather, J.P. & Barnes, D.W. Animal Cell Culture Methods (Academic Press, 1998).
Bollag, D.M., Rozycki, M.D. & Edelstein,, S.J. Protein Methods 2nd edn. (Wiley-Liss, 1996).
Acknowledgements
This work was supported by the National Creative Research Initiative Program (Center for Single-Molecule Systems Biology to T.-Y.Y.) and the Basic Science Research Program (2011-0012385 to K.K.) through the National Research Foundation of Korea (NRF) funded by the Korean government.
Author information
Authors and Affiliations
Contributions
T.-Y.Y. designed the experiments. H.-W.L., J.Y. and B.C. performed single-molecule experiments. H.-W.L., J.Y.R. and K.K. wrote the analysis programs. H.-W.L., J.Y.R., J.Y., K.K. and T.-Y.Y. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Quantitative immunoblot of endogenous and exogenous cRaf in HeLa cells.
(a) Quantitative immunoblot image of endogenous and exogenous cRaf in HeLa cells. We transfected an EGFP-cRaf plasmid to HeLa cells and performed serial dilutions of EGFP fluorescence in lysates from 200 pM to 2,000 pM for quantitative immunoblotting. To measure endogenous level of cRaf, we also loaded 1 and 2 mg ml−1 of HeLa cell lysates. All samples were run on SDS-PAGE gel, followed by immunoblotting against full length cRaf. Each band signal intensity was quantified using ImageJ. (b) We plotted the concentration [pM] of EGFP-cRaf versus band intensity from 200 pM to 2,000 pM, which showed a linear relationship. To measure the endogenous cRaf level, each band intensity from endogenous cRaf was quantified using ImageJ and converted to concentration with the calibration curve. The endogenous cRaf concentration was 71 pM in 1 mg ml−1 and 166 pM in 2 mg ml−1 of cell extracts. Thus, the expression level of endogenous cRaf was ∼77 pM per mg ml−1 in HeLa cell. In the case of exogenous cRaf, typically 105 HeLa cells yielded a few tens of nM of EGFP fluorescence level per mg ml−1 of cell extract. Thus, in the case of cRaf, we could obtain exogenously expressed preys at a concentration higher than that of the endogenous protein, by at least two orders of magnitude.
Supplementary Figure 2 An alternative model to explain the kinetic rates of kbind with two-exponential fitting.
We used two models to analyze the distribution of τon (kbind) in Figure 2g; one with a single exponential (green line) model and the other with a double exponential (red line) model. The single exponential model gives ~0.4 s−1 as the kinetic value of kbind and the double exponential model gives two kinetic values, ∼0.3 s−1 and 3.8 s−1. The two-exponential fitting identifies a small component of high kbind (3.8 s−1), but the main component still gives a kbind of 0.3 s−1, which is very close to 0.4 s−1 obtained by single exponential fitting.
Supplementary Figure 3 Simulation result to calculate the practical limit of kdiss.
To examine the practical limit of kdiss, we sampled 3,000 random values from the single exponential distribution having a fixed kdiss_theoretical. Then we plotted the distribution of the resulting population and fitted it to a single exponential model to infer kdiss. We repeated this simulation 100 times under equal conditions to get values of kdiss and calculated the relative error as the following: Relative error (%) = kdiss_theoretical − kdiss /kdiss_theoretical × 100 . Then we calculated the average value and s.d. (error bars, n = 100) of the Relative error at each kdiss. Increasing kdiss−1 by increments of 50 ms, we collected a dataset of kdiss−1 from 50 ms to 1 s. Our simulation result showed that the relative error of kdiss−1 was suppressed to less than 5% above 150 ms of kdiss−1, which corresponds to 6.7 s−1 of kdiss. Thus if kdiss is lower than 6.7 s−1, we are able to measure kdiss with less than a 5% error.
Supplementary Figure 4 Calibration curve to determine the fluorescence level in cell lysates described in Step 7.
Shown are 11 different concentrations of recombinant EGFP or mCherry used to measure fluorescence intensity. We plot the fluorescence intensity versus protein concentration and fit the data to a linear function to obtain the calibration curve. This calibration curve converts the fluorescence intensity observed to the concentration of EGFP or mCherry-labelled proteins in cell lysates. (a) Exemplary calibration curve of recombinant EGFP. (b) Exemplary calibration curve of recombinant mCherry.
Supplementary Figure 5 Calibration curve to determine the total number of bait proteins immobilized on the surface.
We count the numbers of immobilized single mCherry-bait proteins while varying the mCherry-bait concentration and fit the data to a linear function. Error bars denote standard deviation (n=10).
Supplementary Figure 6 Exemplary inappropriate traces for analysis.
(a) An exemplary trace generating false positive result. This trace shows fluctuations of small peaks occurring between 5-10 s and 30-35 s. When the threshold passes through this fluctuation, many short τoff and τon values are generated. These short τoff and τon values give a false-positive population of high kdiss and kbind value, leading to an overestimation of the kinetic values. (b) Exemplary trace with low TIR excitation intensity. Beam pattern causes inhomogeneity of the TIR excitation field, which cannot illuminate the entire field of view homogenously. Thus some molecules are excited by lower TIR excitation intensity, generating small peaks, as shown at 15 and 48 s. If the signal trace of these small peaks does not exceed the threshold, the Kinetics Analyzer program will miss these peaks and will generate longer τoff, which corresponds to small kbind. Even if the signal trace of a small peak exceeds the threshold, the trace lasts for a short time, which gives short τoff value and it result in an overestimation of kdiss. (c) Exemplary trace including a junk fluorescence molecule. Junk fluorescence molecules generally have a long lifetime, as shown between 10-30 s. This junk fluorescence molecule seems to be an autofluorescent contaminants in the PEG- cushion layer. When the trace of this junk fluorescence molecule is included in the analysis, many short τoff and τon values are generated, as shown. These values will generate a false-positive population of high kdiss and kbind values and cause the kinetic values to be overestimated.
Supplementary Figure 7 Single-molecule kinetic rates of native KRas pulled down from MDA-MB-231 cells.
Using the Kinetics Analyzer program, we analysed the kinetic rates of endogenous KRas obtained from the MDA-MB-231 cells used in the experiment depicted in Figure 2f. (a) Distribution of kbind for the experiment in Figure 2f. The kinetic rate was obtained through single exponential fitting (solid line), which gave a value of 0.215 s−1. (b) Distribution of kdiss for the experiment in Figure 2f. From the single exponential fit (solid line), we obtained a value of 2.51 s−1 for kdiss. Error bars denote standard errors.
Supplementary information
Supplementary Discussion
Supplementary Discussion (PDF 616 kb)
Supplementary Figure 1
Quantitative immunoblot of endogenous and exogenous cRaf in HeLa cells. (PDF 223 kb)
Supplementary Figure 2
An alternative model to explain the kinetic rates of kbind with two-exponential fitting. (PDF 37 kb)
Supplementary Figure 3
Simulation result to calculate the practical limit of kdiss. (PDF 80 kb)
Supplementary Figure 4
Calibration curve to determine the fluorescence level in cell lysates described in Step 7. (PDF 142 kb)
Supplementary Figure 5
Calibration curve to determine the total number of bait proteins immobilized on the surface. (PDF 128 kb)
Supplementary Figure 6
Exemplary inappropriate traces for analysis. (PDF 137 kb)
Supplementary Figure 7
Single-molecule kinetic rates of native KRas pulled down from MDA-MB-231 cells. (PDF 44 kb)
Rights and permissions
About this article
Cite this article
Lee, HW., Ryu, J., Yoo, J. et al. Real-time single-molecule coimmunoprecipitation of weak protein-protein interactions. Nat Protoc 8, 2045–2060 (2013). https://doi.org/10.1038/nprot.2013.116
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.116
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.