Abstract
Gene knockout experiments are often essential in functional genomics and metabolic engineering studies. However, repeated multiple gene knockout experiments are laborious, time consuming and sometimes impossible to perform for those genes that are essential for cell function. Small regulatory RNAs (sRNAs) are short noncoding RNAs in prokaryotes that can finely control the expression of target genes in trans at the post-transcriptional level. Here we describe the protocol for synthetic sRNA-based gene expression control, including sRNA design principles. Customized synthetic sRNAs consist of a scaffold and a target-binding sequence, and they can be created by simply replacing an existing target-binding sequence with one that is complementary to the target mRNA to be repressed, while retaining the scaffold. Our plasmid-based synthetic sRNA system does not require chromosomal modifications, and it enables one to perform high-throughput studies on the effects of knockdowns on host cell physiology, and it further allows the simultaneous screening of target genes in different Escherichia coli strains for applications in metabolic engineering and synthetic biology. Once an sRNA scaffold-harboring plasmid is constructed, customized synthetic sRNAs can be made within 3–4 d; after this time, the synthetic sRNAs can be applied to the desired experiments.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Warner, J.R. et al. Rapid profiling of a microbial genome using mixtures of barcoded oligonucleotides. Nat. Biotechnol. 28, 856–862 (2010).
Wang, H.H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Sandoval, N.R. et al. Strategy for directing combinatorial genome engineering in Escherichia coli. Proc. Natl. Acad. Sci. USA 109, 10540–10545 (2012).
Lee, S.Y., Lee, K.M., Chan, H.N. & Steinbüchel, A. Comparison of recombinant Escherichia coli strains for synthesis and accumulation of poly-(3-hydroxybutyric acid) and morphological changes. Biotechnol. Bioeng. 44, 1337–1347 (1994).
Desai, S.K. & Gallivan, J.P. Genetic screens and selections for small molecules based on a synthetic riboswitch that activates protein translation. J. Am. Chem. Soc. 126, 13247–13254 (2004).
Nomura, Y. & Yokobayashi, Y. Reengineering a natural riboswitch by dual genetic selection. J. Am. Chem. Soc. 129, 13814–13815 (2007).
Jin, Y. & Huang, J.D. Engineering a portable riboswitch-LacP hybrid device for two-way gene regulation. Nucleic Acids Res. 39, e131 (2011).
Sun, X., Zhulin, I. & Wartell, R.M. Predicted structure and phyletic distribution of the RNA-binding protein Hfq. Nucleic Acids Res. 30, 3663–3671 (2002).
Hershberg, R., Altuvia, S. & Margalit, H. A survey of small RNA-encoding genes in Escherichia coli. Nucleic Acids Res. 31, 1813 (2003).
Win, M.N. & Smolke, C.D. Higher-order cellular information processing with synthetic RNA devices. Science 322, 456–460 (2008).
Man, S. et al. Artificial trans-encoded small non-coding RNAs specifically silence the selected gene expression in bacteria. Nucleic Acids Res. 39, e50 (2011).
Urban, J.H. & Vogel, J. Translational control and target recognition by Escherichia coli small RNAs in vivo. Nucleic Acids Res. 35, 1018–1037 (2007).
Na, D. et al. Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nat. Biotechnol. 31, 170–174 (2013).
Gottesman, S. The small RNA regulators of Escherichia coli: roles and mechanisms. Annu. Rev. Microbiol. 58, 303–328 (2004).
Repoila, F. & Darfeuille, F. Small regulatory non-coding RNAs in bacteria: physiology and mechanistic aspects. Biol. Cell 101, 117–131 (2009).
Storz, G., Opdyke, J. & Zhang, A. Controlling mRNA stability and translation with small, noncoding RNAs. Curr. Opin. Microbiol. 7, 140–144 (2004).
Moll, I., Afonyushkin, T., Vytvytska, O., Kaberdin, V.R. & Bläsi, U. Coincident Hfq binding and RNase E cleavage sites on mRNA and small regulatory RNAs. RNA 9, 1308–1314 (2003).
Aiba, H. Mechanism of RNA silencing by Hfq-binding small RNAs. Curr. Opin. Microbiol. 10, 134–139 (2007).
Delebecque, C.J., Lindner, A.B., Silver, P.A. & Aldaye, F.A. Organization of intracellular reactions with rationally designed RNA assemblies. Science 333, 470–474 (2011).
Kang, Z., Wang, X., Li, Y., Wang, Q. & Qi, Q. Small RNA RyhB as a potential tool used for metabolic engineering in Escherichia coli. Biotechnol. Lett. 34, 527–531 (2012).
Sharma, V., Yamamura, A. & Yokobayashi, Y. Engineering artificial small RNAs for conditional gene silencing in Escherichia coli. ACS Synth. Biol. 1, 6–13 (2012).
Liang, J.C., Bloom, R.J. & Smolke, C.D. Engineering biological systems with synthetic RNA molecules. Mol. Cell 43, 915–926 (2011).
Duckert, P., Brunak, S. & Blom, N. Prediction of proprotein convertase cleavage sites. Protein Eng. Des. Sel. 17, 107–112 (2004).
McDaniel, R. & Weiss, R. Advances in synthetic biology: on the path from prototypes to applications. Curr. Opin. Biotechnol. 16, 476–483 (2005).
Landrain, T.E., Carrera, J., Kirov, B., Rodrigo, G. & Jaramillo, A. Modular model-based design for heterologous bioproduction in bacteria. Curr. Opin. Biotechnol. 20, 272–279 (2009).
McDaniel, R., Endy, D. & Weiss, R. Foundations for engineering biology. Nature 438, 449–453 (2005).
Na, D., Lee, S. & Lee, D. Mathematical modeling of translation initiation for the estimation of its efficiency to computationally design mRNA sequences with a desired expression level in prokaryotes. BMC Syst. Biol. 4, 71 (2010).
Jana, N.K., Roy, S., Bhattacharyya, B. & Mandal, N.C. Amino acid changes in the repressor of bacteriophage lambda due to temperature-sensitive mutations in its cI gene and the structure of a highly temperature-sensitive mutant repressor. Protein Eng. 12, 225–233 (1999).
Zhang, R., Ou, H.Y. & Zhang, C.T. DEG: a database of essential genes. Nucleic Acids Res. 32, D271–272 (2004).
Markham, N. & Zuker, M. UNAFold: software for nucleic acid folding and hybridization. Methods Mol. Biol. 453, 3–31 (2008).
Nolan, T., Hands, E.R. & Bustin, A.S. Quantification of mRNA using real-time RT-PCR. Nat. Protoc. 1, 1559–1582 (2006).
Mahmood, T. & Yang, P.-C. Western blot: technique, theory, and trouble shooting. N. Am. J. Med. Sci. 4, 429–434 (2012).
Sambrook, J. & Russell, D. Molecular Cloning: a Laboratory Manual (Cold Spring Harbor Laboratory Press, 2001).
Schägger, H. Tricine–SDS-PAGE. Nat. Protoc. 1, 16–22 (2006).
Acknowledgements
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries (NRF-2012-C1AAA001-2012M1A2A2026556) and the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) of the Ministry of Science, ICT and Future Planning through the National Research Foundation of Korea. This protocol paper describes the detailed strategies and step-by-step procedures for the work reported in our previous paper.
Author information
Authors and Affiliations
Contributions
D.N. and S.Y.L. conceived the project. S.M.Y. and D.N. performed all experiments and developed the experimental protocol. S.Y.L. supervised the project. All authors wrote the paper together and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figure 1
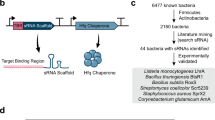
Scaffold selection for the construction of effective synthetic sRNAs. (a) Most sRNAs possess a consensus sequence that is able to recruit the Hfq protein. The left figure illustrates the process of analyzing sequence features of discovered sRNAs in E. coli and the selection of three candidate scaffolds from MicC, MicF and SgrS sRNAs. The respective scaffolds were fused with a sequence complementary to the TIR of DsRed2 mRNA in order to build anti-DsRed2 synthetic sRNAs with different scaffolds. The graph on the right shows that the three scaffolds are highly efficient (>90%) in repressing DsRed2 translation and that the MicC scaffold showed the highest repression efficiency. Consequently, the MicC scaffold was selected to be the synthetic sRNA scaffold. “C”, no synthetic sRNA; “-”, a scaffold without the DsRed2 targeting sequence; “+”, a scaffold with the DsRed2 targeting sequence. (b) Identification of effective mRNA binding regions and sRNA lengths are identified. The graph on the right shows repressed DsRed2 expression by synthetic sRNAs targeting different regions in the mRNA, as indicated in the left figure. “A” denotes the same anti-DsRed2 synthetic sRNA with a MicC scaffold used in (a). The TIR, denoted by a green bar, was estimated using a previously published algorithm27. (Reprinted from ref. 13; please see details there.) (PDF 316 kb)
Supplementary Figure 2
Testing the specificities of synthetic sRNA by using the web-based tools TargetRNA2 and RNAPredator. (a) TargetRNA2. Input information in the indicated boxes: select the target genome to search to E. coli (named ‘Replicon’), input the whole sequence of the synthetic sRNA (named ‘sRNA sequence’) – in this example anti-lacZ, enter ‘24’ in ‘Target parameters’, and then ‘Submit’. (b) Predicted target mRNAs for the anti-lacZ synthetic sRNA by TargetRNA2. (c) Predicted target mRNAs for the anti-lacZ synthetic sRNA by RNAPredator. (PDF 278 kb)
Supplementary Figure 3
Binding free energy–dependent repression capabilities of different anti-lacZ synthetic sRNA. (a) Primers for the anti-lacZ synthetic sRNAs with different binding free energies were designed. (b) The tunability of repression with respect to binding energy was investigated for lacZα (MITRegistry, BBa_I732006) and its corresponding synthetic sRNAs. The LacZ expression levels were measured according to the standard β-galactosidase assay in E. coli DH5α strain by α-complementation33. (Reprinted from ref. 13; please see details there.) (PDF 455 kb)
Supplementary Figure 4
Vector map of the pWAS plasmid. (a) This figure illustrates the brief structure of the pWAS plasmid and indicates promoters, coding sequences, synthetic sRNAs, etc. (b) The vector sequence is available in the Supplementary Data file, where the locations of relevant genetic components in the plasmid are listed. (PDF 136 kb)
Supplementary Discussion
Computational design of synthetic sRNAs and high-throughput library construction. (PDF 154 kb)
Supplementary Data
Sequence of the pWAS vector. (TXT 85 kb)
Rights and permissions
About this article
Cite this article
Yoo, S., Na, D. & Lee, S. Design and use of synthetic regulatory small RNAs to control gene expression in Escherichia coli. Nat Protoc 8, 1694–1707 (2013). https://doi.org/10.1038/nprot.2013.105
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.105
This article is cited by
-
Systems metabolic engineering of Escherichia coli for hyper-production of 5‑aminolevulinic acid
Biotechnology for Biofuels and Bioproducts (2023)
-
A programmable encapsulation system improves delivery of therapeutic bacteria in mice
Nature Biotechnology (2022)
-
Synthetic fused sRNA for the simultaneous repression of multiple genes
Applied Microbiology and Biotechnology (2022)
-
Post-transcriptional control of bacterial nitrogen metabolism by regulatory noncoding RNAs
World Journal of Microbiology and Biotechnology (2022)
-
Recent Research Advances in Small Regulatory RNAs in Streptococcus
Current Microbiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.