Abstract
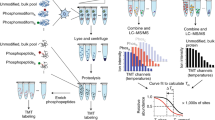
We outline NMR protocols for site-specific mapping and time-resolved monitoring of protein phosphorylation reactions using purified kinases and mammalian cell extracts. These approaches are particularly amenable to intrinsically disordered proteins and unfolded, regulatory protein domains. We present examples for the 15N isotope-labeled N-terminal transactivation domain of human p53, which is either sequentially reacted with recombinant enzymes or directly added to mammalian cell extracts and phosphorylated by endogenous kinases. Phosphorylation reactions with purified enzymes are set up in minutes, whereas NMR samples in cell extracts are prepared within 1 h. Time-resolved NMR measurements are performed over minutes to hours depending on the activities of the probed kinases. Phosphorylation is quantitatively monitored with consecutive 2D 1H-15N band-selective optimized-flip-angle short-transient (SOFAST)-heteronuclear multiple-quantum (HMQC) NMR experiments, which provide atomic-resolution insights into the phosphorylation levels of individual substrate residues and time-dependent changes thereof, thereby offering unique advantages over western blotting and mass spectrometry.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Change history
28 October 2015
In the version of this article initially published, Equation 11 was incorrect. This sign within the brackets was 'plus' and it should have been 'minus'. The error has been corrected in the HTML and PDF versions of the article.
References
Deribe, Y.L., Pawson, T. & Dikic, I. Post-translational modifications in signal integration. Nat. Struct. Mol. Biol. 17, 666–672 (2010).
Walsh, C.T., Garneau-Tsodikova, S. & Gatto, G.J. Jr. Protein posttranslational modifications: the chemistry of proteome diversifications. Angew. Chem. Int. Ed. Engl. 44, 7342–7372 (2005).
Khoury, G.A., Baliban, R.C. & Floudas, C.A. Proteome-wide post-translational modification statistics: frequency analysis and curation of the SwissProt database. Sci. Rep. http://dx.doi.org/10.1038/srep00090 (13 September 2011).
Hunter, T. Tyrosine-phosphorylation: thirty years and counting. Curr. Opin. Cell Biol. 21, 140–146 (2009).
Kee, J.M. & Muir, T.W. Chasing phosphohistidine, an elusive sibling in the phosphoamino acid family. ACS Chem. Biol. 7, 44–51 (2012).
Manning, G., Whyte, D.B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
McConnell, J.L. & Wadzinski, B.E. Targeting protein serine/threonine phosphatases for drug development. Mol. Pharmacol. 75, 1249–1261 (2009).
Eglen, R.M. & Reisine, T. The current status of drug discovery against the human kinome. Assay Drug Dev. Technol. 7, 22–43 (2009).
Schwartz, P.A. & Murray, B.W. Protein kinase biochemistry and drug discovery. Bioorg. Chem. 39, 192–210 (2011).
Cohen, P. The regulation of protein function by multisite phosphorylation–a 25-year update. Trends Biochem. Sci. 25, 596–601 (2000).
Yang, X.J. Multisite protein modification and intramolecular signaling. Oncogene 24, 1653–1662 (2005).
Gnad, F., Gunawardena, J. & Mann, M. PHOSIDA 2011: the posttranslational modification database. Nucleic Acids Res. 39, D253–260 (2011).
Schweiger, R. & Linial, M. Cooperativity within proximal phosphorylation sites is revealed from large-scale proteomics data. Biol. Direct 5, 6 (2010).
Thomson, M. & Gunawardena, J. Unlimited multistability in multisite phosphorylation systems. Nature 460, 274–277 (2009).
Salazar, C. & Hofer, T. Multisite protein phosphorylation–from molecular mechanisms to kinetic models. FEBS J. 276, 3177–3198 (2009).
Liu, X., Bardwell, L. & Nie, Q. A combination of multisite phosphorylation and substrate sequestration produces switchlike responses. Biophys. J. 98, 1396–1407 (2010).
Wang, L., Nie, Q. & Enciso, G. Non-essential sites improve phosphorylation switch. Biophys. J. 99, L41–L43 (2010).
Selenko, P. et al. In situ observation of protein phosphorylation by high-resolution NMR spectroscopy. Nat. Struct. Mol. Biol. 15, 321–329 (2008).
Liokatis, S., Dose, A., Schwarzer, D. & Selenko, P. Simultaneous detection of protein phosphorylation and acetylation by high-resolution NMR spectroscopy. J. Am. Chem. Soc. 132, 14704–14705 (2010).
Theillet, F.X. et al. Site-specific mapping and time-resolved monitoring of lysine methylation by high-resolution NMR spectroscopy. J. Am. Chem. Soc. 134, 7616–7619 (2012).
Liokatis, S. et al. Phosphorylation of histone H3 Ser10 establishes a hierarchy for subsequent intramolecular modification events. Nat. Struct. Mol. Biol. 19, 819–823 (2012).
Theillet, F.X. et al. Cell signaling, post-translational protein modifications and NMR spectroscopy. J. Biomol. NMR 54, 217–236 (2012).
Sattler, M., Schleucher, J. & Griesinger, C. Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog. Nucl. Mag. Res. Sp. 34, 93–158 (1999).
Brutscher, B. & Lescop, E. Fast protein backbone NMR resonance assignment using the BATCH strategy. Methods Mol. Biol. 831, 407–428 (2012).
Schanda, P., Kupce, E. & Brutscher, B. SOFAST-HMQC experiments for recording two-dimensional heteronuclear correlation spectra of proteins within a few seconds. J. Biomol. NMR 33, 199–211 (2005).
Ishida, A., Kameshita, I., Sueyoshi, N., Taniguchi, T. & Shigeri, Y. Recent advances in technologies for analyzing protein kinases. J. Pharmacol. Sci. 103, 5–11 (2007).
Kubota, K. et al. Sensitive multiplexed analysis of kinase activities and activity-based kinase identification. Nat. Biotechnol. 27, 933–940 (2009).
Arsenault, R., Griebel, P. & Napper, S. Peptide arrays for kinome analysis: new opportunities and remaining challenges. Proteomics 11, 4595–4609 (2011).
Gonzalez-Vera, J.A. Probing the kinome in real time with fluorescent peptides. Chem. Soc. Rev. 41, 1652–1664 (2012).
Kettenbach, A.N. et al. Rapid determination of multiple linear kinase substrate motifs by mass spectrometry. Chem. Biol. 19, 608–618 (2012).
Mollica, J.P., Oakhill, J.S., Lamb, G.D. & Murphy, R.M. Are genuine changes in protein expression being overlooked? Reassessing western blotting. Anal. Biochem. 386, 270–275 (2009).
Prabakaran, S. et al. Comparative analysis of Erk phosphorylation suggests a mixed strategy for measuring phospho-form distributions. Mol. Syst. Biol. 7, 482 (2011).
Goto, H. & Inagaki, M. Production of a site- and phosphorylation state-specific antibody. Nat. Protoc. 2, 2574–2581 (2007).
Archuleta, A.J., Stutzke, C.A., Nixon, K.M. & Browning, M.D. Optimized protocol to make phospho-specific antibodies that work. Methods Mol. Biol. 717, 69–88 (2011).
Egelhofer, T.A. et al. An assessment of histone-modification antibody quality. Nat. Struct. Mol. Biol. 18, 91–93 (2011).
Fuchs, S.M., Krajewski, K., Baker, R.W., Miller, V.L. & Strahl, B.D. Influence of combinatorial histone modifications on antibody and effector protein recognition. Curr. Biol. 21, 53–58 (2011).
Bock, I. et al. Detailed specificity analysis of antibodies binding to modified histone tails with peptide arrays. Epigenetics 6, 256–263 (2011).
Witze, E.S., Old, W.M., Resing, K.A. & Ahn, N.G. Mapping protein post-translational modifications with mass spectrometry. Nat. Methods 4, 798–806 (2007).
Kosako, H. & Nagano, K. Quantitative phosphoproteomics strategies for understanding protein kinase-mediated signal transduction pathways. Expert Rev. Proteomics 8, 81–94 (2011).
Nilsson, C.L. Advances in quantitative phosphoproteomics. Anal. Chem. 84, 735–746 (2011).
Stasyk, T. & Huber, L.A. Mapping in vivo signal transduction defects by phosphoproteomics. Trends Mol. Med. 18, 43–51 (2012).
Boehm, M.E., Seidler, J., Hahn, B. & Lehmann, W.D. Site-specific degree of phosphorylation in proteins measured by liquid chromatography-electrospray mass spectrometry. Proteomics 12, 2167–2178 (2012).
Gafken, P.R. & Lampe, P.D. Methodologies for characterizing phosphoproteins by mass spectrometry. Cell Commun. Adhes. 13, 249–262 (2006).
Britton, L.M., Gonzales-Cope, M., Zee, B.M. & Garcia, B.A. Breaking the histone code with quantitative mass spectrometry. Expert Rev. Proteomics 8, 631–643 (2011).
Diedrich, J.K. & Julian, R.R. Facile identification of phosphorylation sites in peptides by radical directed dissociation. Anal. Chem. 83, 6818–6826 (2011).
Taus, T. et al. Universal and confident phosphorylation site localization using phosphoRS. J. Proteome Res. 10, 5354–5362 (2011).
Courcelles, M., Bridon, G., Lemieux, S. & Thibault, P. Occurrence and detection of phosphopeptide isomers in large-scale phosphoproteomics experiments. J. Proteome Res. 11, 3753–3765 (2012).
Young, N.L., Plazas-Mayorca, M.D. & Garcia, B.A. Systems-wide proteomic characterization of combinatorial post-translational modification patterns. Expert Rev. Proteomics 7, 79–92 (2010).
Sidoli, S., Cheng, L. & Jensen, O.N. Proteomics in chromatin biology and epigenetics: Elucidation of post-translational modifications of histone proteins by mass spectrometry. J. Proteomics 75, 3419–3433 (2012).
Kettenbach, A.N., Rush, J. & Gerber, S.A. Absolute quantification of protein and post-translational modification abundance with stable isotope-labeled synthetic peptides. Nat. Protoc. 6, 175–186 (2011).
Singh, S.A. et al. FLEXIQinase, a mass spectrometry–based assay, to unveil multikinase mechanisms. Nat. Methods 9, 504–508 (2012).
Cordier, F. et al. Ordered phosphorylation events in two independent cascades of the PTEN C-tail revealed by NMR. J. Am. Chem. Soc. 134, 20533–20543 (2012).
Landrieu, I. et al. Molecular implication of PP2A and Pin1 in the Alzheimer's disease specific hyperphosphorylation of Tau. PLoS ONE 6, e21521 (2011).
Gallego, M. & Virshup, D.M. Protein serine/threonine phosphatases: life, death, and sleeping. Curr. Opin. Cell Biol. 17, 197–202 (2005).
Shi, Y. Serine/threonine phosphatases: mechanism through structure. Cell 139, 468–484 (2009).
Virshup, D.M. & Shenolikar, S. From promiscuity to precision: protein phosphatases get a makeover. Mol. Cell 33, 537–545 (2009).
Purvis, J.E. & Lahav, G. Encoding and decoding cellular information through signaling dynamics. Cell 152, 945–956 (2013).
Deshmukh, L., Meller, N., Alder, N., Byzova, T. & Vinogradova, O. Tyrosine phosphorylation as a conformational switch: a case study of integrin beta3 cytoplasmic tail. J. Biol. Chem. 286, 40943–40953 (2011).
Smet-Nocca, C., Launay, H., Wieruszeski, J.M., Lippens, G. & Landrieu, I. Unraveling a phosphorylation event in a folded protein by NMR spectroscopy: phosphorylation of the Pin1 WW domain by PKA. J. Biomol. NMR 55, 323–337 (2013).
West, X.Z. et al. Integrin β3 crosstalk with VEGFR accommodating tyrosine phosphorylation as a regulatory switch. PLoS ONE 7, e31071 (2013).
Zhang, Y. et al. Structure, phosphorylation and U2AF65 binding of the N-terminal domain of splicing factor 1 during 3'-splice site recognition. Nucleic Acids Res. 41, 1343–1354 (2013).
Iakoucheva, L.M. et al. The importance of intrinsic disorder for protein phosphorylation. Nucleic Acids Res. 32, 1037–1049 (2004).
Gu, B. & Zhu, W.G. Surf the post-translational modification network of p53 regulation. Int. J. Biol. Sci. 8, 672–684 (2012).
Landrieu, I. et al. NMR analysis of a Tau phosphorylation pattern. J. Am. Chem. Soc. 128, 3575–3583 (2006).
Bienkiewicz, E.A. & Lumb, K.J. Random-coil chemical shifts of phosphorylated amino acids. J. Biomol. NMR 15, 203–206 (1999).
Ulrich, E.L. et al. BioMagResBank. Nucleic Acids Res. 36, D402–D408 (2008).
Guerry, P. & Herrmann, T. Advances in automated NMR protein structure determination. Q. Rev. Biophys. 44, 257–309 (2011).
Hiller, S., Wasmer, C., Wider, G. & Wuthrich, K. Sequence-specific resonance assignment of soluble nonglobular proteins by 7D APSY-NMR spectroscopy. J. Am. Chem. Soc. 129, 10823–10828 (2007).
Zawadzka-Kazimierczuk, A., Kozminski, W., Sanderova, H. & Krasny, L. High dimensional and high resolution pulse sequences for backbone resonance assignment of intrinsically disordered proteins. J. Biomol. NMR 52, 329–337 (2012).
Bagai, I., Ragsdale, S.W. & Zuiderweg, E.R. Pseudo-4D triple-resonance experiments to resolve HN overlap in the backbone assignment of unfolded proteins. J. Biomol. NMR 49, 69–74 (2011).
Bermel, W. et al. Speeding up sequence specific assignment of IDPs. J. Biomol. NMR 53, 293–301 (2012).
Novacek, J. et al. 5D 13C-detected experiments for backbone assignment of unstructured proteins with a very low signal dispersion. J. Biomol. NMR 50, 1–11 (2011).
Theillet, F.X., Binolfi, A., Liokatis, S., Verzini, S. & Selenko, P. Paramagnetic relaxation enhancement to improve sensitivity of fast NMR methods: application to intrinsically disordered proteins. J. Biomol. NMR 51, 487–495 (2011).
Bai, Y., Milne, J.S., Mayne, L. & Englander, S.W. Primary structure effects on peptide group hydrogen exchange. Proteins 17, 75–86 (1993).
Murray-Zmijewski, F., Slee, E.A. & Lu, X. A complex barcode underlies the heterogeneous response of p53 to stress. Nat. Rev. Mol. Cell Biol. 9, 702–712 (2008).
Vousden, K.H. & Ryan, K.M. p53 and metabolism. Nat. Rev. Cancer 9, 691–700 (2009).
Menendez, D., Inga, A. & Resnick, M.A. The expanding universe of p53 targets. Nat. Rev. Cancer 9, 724–737 (2009).
Kruse, J.P. & Gu, W. Modes of p53 regulation. Cell 137, 609–622 (2009).
Wong, T.S. et al. Physical and functional interactions between human mitochondrial single-stranded DNA-binding protein and tumour suppressor p53. Nucleic Acids Res. 37, 568–581 (2009).
Shieh, S.Y., Ikeda, M., Taya, Y. & Prives, C. DNA damage-induced phosphorylation of p53 alleviates inhibition by MDM2. Cell 91, 325–334 (1997).
Lees-Miller, S.P., Sakaguchi, K., Ullrich, S.J., Appella, E. & Anderson, C.W. Human DNA-activated protein kinase phosphorylates serines 15 and 37 in the amino-terminal transactivation domain of human p53. Mol. Cell Biol. 12, 5041–5049 (1992).
Venerando, A. et al. Isoform specific phosphorylation of p53 by protein kinase CK1. Cell Mol. Life Sci. 67, 1105–1118 (2010).
Saito, S. et al. Phosphorylation site interdependence of human p53 post-translational modifications in response to stress. J. Biol. Chem. 278, 37536–37544 (2003).
Putz, M.V., Lacrama, A.M. & Ostafe, V. Full analytic progress curves of enzymic reactions in vitro. Int. J. Mol. Sci. 7, 469–484 (2006).
Acknowledgements
We thank P. Schmieder and M. Beerbaum for excellent maintenance of NMR infrastructure. We also thank F. Cordier for many insightful discussions, G. Lippens for expert advice and all members of the Selenko laboratory for carefully reading the manuscript and providing helpful comments. F.-X.T. was supported by a grant from the Association pour la Recherche sur le Cancer (ARC). P.S. acknowledges support by an Emmy Noether research grant (SE1-1/1794) by the Deutsche Forschungsgemeinschaft (DFG).
Author information
Authors and Affiliations
Contributions
F.-X.T., H.M.R., S.L., A.B., R.T., M.S. and P.S. devised and executed the experiments. Specifically, F.-X.T., H.M.R. and M.S. prepared cell extracts and measured in-extract phosphorylation reactions; F.-X.T., S.L., A.B. and R.T. performed in vitro phosphorylation measurements; and F.-X.T., H.M.R. and M.S. performed western blotting. F.-X.T., H.M.R. and P.S. prepared figures and wrote the manuscript. All authors carefully read the manuscript and approved of the conclusions drawn therein.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Theillet, FX., Rose, H., Liokatis, S. et al. Site-specific NMR mapping and time-resolved monitoring of serine and threonine phosphorylation in reconstituted kinase reactions and mammalian cell extracts. Nat Protoc 8, 1416–1432 (2013). https://doi.org/10.1038/nprot.2013.083
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.083
This article is cited by
-
An intrinsic temporal order of c-JUN N-terminal phosphorylation regulates its activity by orchestrating co-factor recruitment
Nature Communications (2022)
-
Deciphering protein post-translational modifications using chemical biology tools
Nature Reviews Chemistry (2020)
-
Time-resolved NMR monitoring of tRNA maturation
Nature Communications (2019)
-
Random coil shifts of posttranslationally modified amino acids
Journal of Biomolecular NMR (2019)
-
Farseer-NMR: automatic treatment, analysis and plotting of large, multi-variable NMR data
Journal of Biomolecular NMR (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.