Abstract
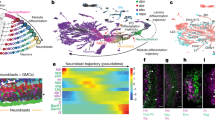
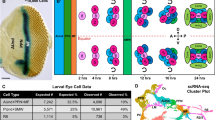
Elegant tools are available for the genetic analysis of neural stem cell lineages in Drosophila, but a methodology for purifying stem cells and their differentiated progeny for transcriptome analysis is currently missing. Previous attempts to overcome this problem either involved using RNA isolated from whole larval brain tissue or co-transcriptional in vivo mRNA tagging. As both methods have limited cell type specificity, we developed a protocol for the isolation of Drosophila neural stem cells (neuroblasts, NBs) and their differentiated sibling cells by FACS. We dissected larval brains from fly strains expressing GFP under the control of a NB lineage–specific GAL4 line. Upon dissociation, we made use of differences in GFP intensity and cell size to separate NBs and neurons. The resulting cell populations are over 98% pure and can readily be used for live imaging or gene expression analysis. Our method is optimized for neural stem cells, but it can also be applied to other Drosophila cell types. Primary cell suspensions and sorted cell populations can be obtained within 1 d; material for deep-sequencing library preparation can be obtained within 4 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Chia, W., Somers, W.G. & Wang, H. Drosophila neuroblast asymmetric divisions: cell cycle regulators, asymmetric protein localization, and tumorigenesis. J. Cell Biol. 180, 267–272 (2008).
Doe, C.Q. Neural stem cells: balancing self-renewal with differentiation. Development 135, 1575–1587 (2008).
Homem, C.C. & Knoblich, J.A. Drosophila neuroblasts: a model for stem cell biology. Development 139, 4297–4310 (2012).
Bello, B.C., Izergina, N., Caussinus, E. & Reichert, H. Amplification of neural stem cell proliferation by intermediate progenitor cells in Drosophila brain development. Neural Dev. 3, 5 (2008).
Boone, J.Q. & Doe, C.Q. Identification of Drosophila type II neuroblast lineages containing transit amplifying ganglion mother cells. Dev. Neurobiol. 68, 1185–1195 (2008).
Bowman, S.K. et al. The tumor suppressors Brat and Numb regulate transit-amplifying neuroblast lineages in Drosophila. Dev. Cell 14, 535–546 (2008).
Ito, K. & Hotta, Y. Proliferation pattern of postembryonic neuroblasts in the brain of Drosophila melanogaster. Dev. Biol. 149, 134–148 (1992).
Truman, J.W. & Bate, M. Spatial and temporal patterns of neurogenesis in the central nervous system of Drosophila melanogaster. Dev. Biol. 125, 145–157 (1988).
Carney, T.D. et al. Functional genomics identifies neural stem cell sub-type expression profiles and genes regulating neuroblast homeostasis. Dev. Biol. 361, 137–146 (2012).
Miller, M.R., Robinson, K.J., Cleary, M.D. & Doe, C.Q. TU-tagging: cell type-specific RNA isolation from intact complex tissues. Nat. Methods 6 (6): 439–41 (2009).
Wang, X., Starz-Gaiano, M., Bridges, T. & Montell,, D. Purification of specific cell populations from Drosophila tissues by magnetic bead sorting, for use in gene expression profiling. Protoc. Exchange http://www.nature.com/doifinder/10.1038/nprot.2008.28 (2008).
Cumberledge, S. & Krasnow, M.A. Preparation and analysis of pure cell populations from Drosophila. Methods Cell Biol. 44, 143–159 (1994).
Kai, T., Williams, D. & Spradling, A.C. The expression profile of purified Drosophila germline stem cells. Dev. Biol. 283, 486–502 (2005).
Tirouvanziam, R., Davidson, C.J., Lipsick, J.S. & Herzenberg, L.A. Fluorescence-activated cell sorting (FACS) of Drosophila hemocytes reveals important functional similarities to mammalian leukocytes. Proc. Natl. Acad. Sci. USA 101, 2912–2917 (2004).
de la Cruz, A.F. & Edgar, B.A. Flow cytometric analysis of Drosophila cells. Methods Mol. Biol. 420, 373–389 (2008).
Berger, C. et al. FACS purification and transcriptome analysis of Drosophila neural stem cells reveals a role for Klumpfuss in self-renewal. Cell Rep. 2, 407–418 (2012).
Goulas, S., Conder, R. & Knoblich, J.A. The par complex and integrins direct asymmetric cell division in adult intestinal stem cells. Cell Stem Cell 11, 529–540 (2012).
McGuire, S.E., Le, P.T., Osborn, A.J., Matsumoto, K. & Davis, R.L. Spatiotemporal rescue of memory dysfunction in Drosophila. Science 302, 1765–1768 (2003).
McGuire, S.E., Mao, Z. & Davis, R.L. Spatiotemporal gene expression targeting with the TARGET and gene-switch systems in Drosophila. Sci. STKE 2004, pl6 (2004).
Fristrom, J.W. & Mitchell, H.K. The preparative isolation of imaginal discs from larvae of Drosophila melanogaster. J. Cell Biol. 27, 445–448 (1965).
Zweidler, A. & Cohen, L.H. Large-scale isolation and fractionation of organs of Drosophila melanogaster larvae. J. Cell Biol. 51, 240–248 (1971).
Zhu, S. et al. Gradients of the Drosophila chinmo BTB-zinc finger protein govern neuronal temporal identity. Cell 127, 409–422 (2006).
Neumuller, R.A. et al. Genome-wide analysis of self-renewal in Drosophila neural stem cells by transgenic RNAi. Cell Stem Cell 8, 580–593 (2011).
Zhu, S., Barshow, S., Wildonger, J., Jan, L.Y. & Jan, Y.N. Ets transcription factor pointed promotes the generation of intermediate neural progenitors in Drosophila larval brains. Proc. Natl. Acad. Sci. USA 108, 20615–20620 (2011).
Barolo, S., Carver, L.A. & Posakony, J.W. GFP and -galactosidase transformation vectors for promoter/enhancer analysis in Drosophila. Biotechniques 29, 726, 728, 730, 732 (2000).
Ceron, J., Tejedor, F.J. & Moya, F. A primary cell culture of Drosophila postembryonic larval neuroblasts to study cell cycle and asymmetric division. Eur. J. Cell Biol. 85, 567–575 (2006).
Winnebeck, E.C., Millar, C.D. & Warman, G.R. Why does insect RNA look degraded? J. Insect Sci. 10, 159 (2010).
Acknowledgements
We thank G. Stengl, T. Lendl and N. Corsini for FACS, P. Pasierbek for imaging support, T.R. Burkard for bioinformatics analyses, and all members of the Knoblich group for discussion and suggestions. We are grateful to the Campus Science Support Facilities Next Generation Sequencing Unit for performing library preparation and next-generation sequencing. C.B. is supported by an EMBO Long Term Fellowship. Work in J.A.K.'s laboratory is supported by the Austrian Academy of Sciences, the Austrian Science Fund, the EU FP7 network EuroSystems and an advanced grant from the European Research Council.
Author information
Authors and Affiliations
Contributions
C.B., H.H. and J.A.K. designed the study. H.H., C.B. and R.C. conducted the experiments. G.S. assisted in the development of the FACS protocol. H.H., C.B. and J.A.K. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Harzer, H., Berger, C., Conder, R. et al. FACS purification of Drosophila larval neuroblasts for next-generation sequencing. Nat Protoc 8, 1088–1099 (2013). https://doi.org/10.1038/nprot.2013.062
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.062
This article is cited by
-
Continuous collection and simultaneous detection of picoliter volume of nucleic acid samples using a mille-feuille probe
Analytical and Bioanalytical Chemistry (2017)
-
Single neuron transcriptomics identify SRSF/SR protein B52 as a regulator of axon growth and Choline acetyltransferase splicing
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.