Abstract
Prions are proteinaceous infectious agents responsible for the transmission of prion diseases. The lack of a procedure for cultivating prions in the laboratory has been a major limitation to the study of the unorthodox nature of this infectious agent and the molecular mechanism by which the normal prion protein (PrPC) is converted into the abnormal isoform (PrPSc). Protein misfolding cyclic amplification (PMCA), described in detail in this protocol, is a simple, fast and efficient methodology to mimic prion replication in the test tube. PMCA involves incubating materials containing minute amounts of infectious prions with an excess of PrPC and boosting the conversion by cycles of sonication to fragment the converting units, thereby leading to accelerated prion replication. PMCA is able to detect the equivalent of a single molecule of infectious PrPSc and propagate prions that maintain high infectivity, strain properties and species specificity. A single PMCA assay takes little more than 3 d to replicate a large amount of prions, which could take years in an in vivo situation. Since its invention 10 years ago, PMCA has helped to answer fundamental questions about this intriguing infectious agent and has been broadly applied in research areas that include the food industry, blood bank safety and human and veterinary disease diagnosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Prusiner, S.B. Prions. Proc. Natl. Acad. Sci. USA 95, 13363–13383 (1998).
Collinge, J. Prion diseases of humans and animals: their causes and molecular basis. Annu. Rev. Neurosci. 24, 519–550 (2001).
Collee, J.G. & Bradley, R. BSE: a decade on—Part I. Lancet 349, 636–641 (1997).
Collee, J.G. & Bradley, R. BSE: a decade on—Part 2. Lancet 349, 715–721 (1997).
Llewelyn, C.A. et al. Possible transmission of variant Creutzfeldt-Jakob disease by blood transfusion. Lancet 363, 417–421 (2004).
Peden, A.H., Head, M.W., Ritchie, D.L., Bell, J.E. & Ironside, J.W. Preclinical vCJD after blood transfusion in a PRNP codon 129 heterozygous patient. Lancet 364, 527–529 (2004).
Wroe, S.J. et al. Clinical presentation and pre-mortem diagnosis of variant Creutzfeldt-Jakob disease associated with blood transfusion: a case report. Lancet 368, 2061–2067 (2006).
Soto, C. Prion hypothesis: the end of the controversy? Trends Biochem. Sci. 36, 151–158 (2011).
Soto, C. & Satani, N. The intricate mechanisms of neurodegeneration in prion diseases. Trends Mol. Med. 36, 151–158 (2011).
Saborio, G.P., Permanne, B. & Soto, C. Sensitive detection of pathological prion protein by cyclic amplification of protein misfolding. Nature 411, 810–813 (2001).
Soto, C., Saborio, G.P. & Anderes, L. Cyclic amplification of protein misfolding: application to prion-related disorders and beyond. Trends Neurosci. 25, 390–394 (2002).
Caughey, B., Kocisko, D.A., Raymond, G.J. & Lansbury, P.T. Jr. Aggregates of scrapie-associated prion protein induce the cell-free conversion of protease-sensitive prion protein to the protease-resistant state. Chem. Biol. 2, 807–817 (1995).
Ghetti, B. et al. Prion protein amyloidosis. Brain Pathol. 6, 127–145 (1996).
Jarrett, J.T. & Lansbury, P.T. Jr. Seeding 'one-dimensional crystallization' of amyloid: a pathogenic mechanism in Alzheimer's disease and scrapie? Cell 73, 1055–1058 (1993).
Soto, C., Estrada, L. & Castilla, J. Amyloids, prions and the inherent infectious nature of misfolded protein aggregates. Trends Biochem. Sci. 31, 150–155 (2006).
Castilla, J., Saá, P., Hetz, C. & Soto, C. In vitro generation of infectious scrapie prions. Cell 121, 195–206 (2005).
Saa, P., Castilla, J. & Soto, C. Ultra-efficient replication of infectious prions by automated protein misfolding cyclic amplification. J. Biol. Chem. 281, 35245–35252 (2006).
Gonzalez-Montalban, N. et al. Highly efficient protein misfolding cyclic amplification. PLoS Pathog. 7, e1001277 (2011).
Fernandez-Borges, N. & Castilla, J. PMCA. A decade of in vitro prion replication. Curr. Chem. Biol. 4, 200–207 (2010).
Pritzkow, S. et al. Quantitative detection and biological propagation of scrapie seeding activity in vitro facilitate use of prions as model pathogens for disinfection. PLoS ONE 6, e20384 (2011).
Castilla, J. et al. Cell-free propagation of prion strains. EMBO J. 27, 2557–2566 (2008).
Green, K.M. et al. Accelerated high fidelity prion amplification within and across prion species barriers. PLoS Pathog. 4, e1000139 (2008).
Meyerett, C. et al. In vitro strain adaptation of CWD prions by serial protein misfolding cyclic amplification. Virology 382, 267–276 (2008).
Deleault, N.R., Harris, B.T., Rees, J.R. & Supattapone, S. Formation of native prions from minimal components in vitro. Proc. Natl. Acad. Sci. USA 104, 9741–9746 (2007).
Wang, F., Wang, X., Yuan, C.-G. & Ma, J. Generating a prion with bacterially expressed recombinant prion protein. Science 327, 1132–1135 (2010).
Abid, K., Morales, R. & Soto, C. Cellular factors implicated in prion replication. FEBS Lett. 584, 2409–2414 (2010).
Supattapone, S. Biochemistry. What makes a prion infectious? Science 327, 1091–1092 (2010).
Deleault, N.R., Kascsak, R., Geoghegan, J.C. & Supattapone, S. Species-dependent differences in cofactor utilization for formation of the protease-resistant prion protein in vitro. Biochemistry 49, 3928–3934 (2010).
Castilla, J. et al. Crossing the species barrier by PrP(Sc) replication in vitro generates unique infectious prions. Cell 134, 757–768 (2008).
Barria, M.A., Mukherjee, A., Gonzalez-Romero, D., Morales, R. & Soto, C. De novo generation of infectious prions in vitro produces a new disease phenotype. PLoS Pathog. 5, e1000421 (2009).
Soto, C. Diagnosing prion diseases: needs, challenges and hopes. Nat. Rev. Microbiol. 2, 809–819 (2004).
Castilla, J., Saa, P. & Soto, C. Detection of prions in blood. Nat. Med. 11, 982–985 (2005).
Gonzalez-Romero, D., Barria, M.A., Leon, P., Morales, R. & Soto, C. Detection of infectious prions in urine. FEBS Lett. 582, 3161–3166 (2008).
Murayama, Y. et al. Urinary excretion and blood level of prions in scrapie-infected hamsters. J. Gen. Virol. 88, 2890–2898 (2007).
Tattum, M.H., Jones, S., Pal, S., Collinge, J. & Jackson, G.S. Discrimination between prion-infected and normal blood samples by protein misfolding cyclic amplification. Transfusion 50, 996–1002 (2010).
Saa, P., Castilla, J. & Soto, C. Presymptomatic detection of prions in blood. Science 313, 92–94 (2006).
Chen, B., Morales, R., Barria, M.A. & Soto, C. Estimating prion concentration in fluids and tissues by quantitative PMCA. Nat. Methods 7, 519–520 (2010).
Barret, A. et al. Evaluation of quinacrine treatment for prion diseases. J. Virol. 77, 8462–8469 (2003).
Morales, R. et al. Reduction of prion infectivity in packed red blood cells. Biochem. Biophys. Res. Commun. 377, 373–378 (2008).
Haley, N.J., Seelig, D.M., Zabel, M.D., Telling, G.C. & Hoover, E.A. Detection of CWD prions in urine and saliva of deer by transgenic mouse bioassay. PLoS ONE 4, e4848 (2009).
Haley, N.J., Mathiason, C.K., Zabel, M.D., Telling, G.C. & Hoover, E.A. Detection of sub-clinical CWD infection in conventional test-negative deer long after oral exposure to urine and feces from CWD+ deer. PLoS ONE 4, e7990 (2009).
Nichols, T.A. et al. Detection of protease-resistant cervid prion protein in water from a CWD-endemic area. Prion 3, 171–183 (2009).
Nagaoka, K. et al. Sensitive detection of scrapie prion protein in soil. Biochem. Biophys. Res. Commun. 397, 626–630 (2010).
Bessen, R.A. & Marsh, R.F. Biochemical and physical properties of the prion protein from two strains of the transmissible mink encephalopathy agent. J. Virol. 66, 2096–2101 (1992).
Murayama, Y. et al. Efficient in vitro amplification of a mouse-adapted scrapie prion protein. Neurosci. Lett. 413, 270–273 (2006).
Barria, M.A., Telling, G.C., Gambetti, P., Mastrianni, J.A. & Soto, C. Generation of a new form of human PrPSc in vitro by interspecies transmission from cervid prions. J. Biol. Chem. 286, 7490–7495 (2011).
Kurt, T.D. et al. Efficient in vitro amplification of chronic wasting disease PrPRES. J. Virol. 81, 9605–9608 (2007).
Cosseddu, G.M. et al. Ultraefficient PrPSc amplification highlights potentialities and pitfalls of the PMCA technology. PLoS Pathog. 7, e1002370 (2011).
Thorne, L. & Terry, L.A. In vitro amplification of PrPSc derived from the brain and blood of sheep infected with scrapie. J. Gen. Virol. 89, 3177–3184 (2008).
Acknowledgements
We thank the many former laboratory members whose input has been important for reaching the optimal PMCA procedure, especially G. Saborio, L. Anderes, C. Adessi, K. Maundrell, J. Castilla, P. Saa, K. Abid, M. Barria, D. Gonzalez-Romero and B. Chen. We also thank M.-J. Liberona for critical reading of the manuscript. This work was partially supported by US National Institutes of Health grants R01 NS049173, P01 AI077774 and P01 AG014359 to C.S.
Author information
Authors and Affiliations
Contributions
R.M. designed the experiments, carried out the work, analyzed the results and wrote part of the manuscript. C.D.-A. performed various PMCA assays, wrote part of the manuscript and drafted the figures. M.V.C. performed various PMCA optimizations and wrote part of the manuscript. R.D.-E. performed experiments with recombinant prion protein and wrote part of the manuscript, and C.S. is the principal investigator on the project and was responsible for coordinating research activity, funding and producing the final version of the article.
Corresponding author
Ethics declarations
Competing interests
Dr Soto is inventor on several patents related to the PMCA technology and is currently Founder, Chief Scientific Officer and Vice -President of Amprion Inc., a biotech company focusing on the commercial exploitation of PMCA for prion diagnosis.
Supplementary information
Supplementary Fig. 1
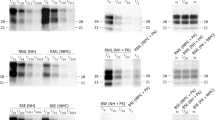
Schematic representation of a PMCA procedure using purified PrPC and PrPSc components. Purified PrPC and PrPSc submitted to PMCA in the sole presence of conversion buffer will not show any PrP27-30 signal as shown in the Western blot of the left panel. However, complementing the PMCA reaction with tissue homogenate from different sources or purified molecules will result in a typical PrPSc amplification (middle and right Western blots). This system could be applied to investigate in more detail the role of different molecules in the misfolding and aggregation processes of mammalian prions. Samples shown in the Western blots were all PK treated. (PDF 654 kb)
Supplementary Fig. 2
Perfused brains for PMCA substrate. A proper preparation of homogenates from non-infected animals is one of the most important steps in the PMCA procedure. We have observed that blood components can dramatically decrease the PrPSc amplification in a PMCA reaction. For that reason, brain perfusion is critical when preparing a good quality brain homogenate. Properly perfused brains (A) are easily differentiated from those that are not well perfused (B) by their appearance and color. (PDF 3423 kb)
Supplementary Fig. 3
Quality of the sonicator horn. Sonicator platforms from automated sonicators adapted for PMCA release white sediments that decrease the sonication efficiency inside the tube. The release of these sediments is especially abundant in new sonicators. We strongly recommend periodically cleaning the sonicator horn, especially when a new PMCA assay starts. A: Sonicator horn with typical white sediment. As observed, it is impossible to see the sonication platform. B: Sonicator horn filled with clean water. Note the rusted-like appearance in the sonicator platform typically observed when sonicator ages. (PDF 5005 kb)
Supplementary Fig. 4
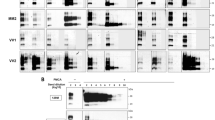
Quantification of PrPSc by qPMCA. A: A PrPSc standard for qPMCA is prepared by partially purifying PrPSc from the brain of infected animals by a series of sarkosyl precipitation steps. The concentration of PrPSc in the standard inoculum is estimated by comparison with known amounts of recombinant PrP (rPrP), produced as described in Supplementary Method 1. For better comparison of the signal, partially purified PrPSc can be deglycosylated by PNGase as described in Box 1 and treated with PK. The asterisk indicates a band of PrPSc incompleted digested by PK. B: In order to assess the extent of amplification in the standard sample, aliquots of PrPSc ranging from 10-8 to 10-21 g of partially purified PrPSc are subjected to sPMCA using standard conditions. All samples shown in B were PK digested with the sole exception of NBH which corresponds to the PrPC signal from a normal (non-infected) brain homogenate which is used as a control of electrophoretical mobility. (PDF 2120 kb)
Supplementary Methods
PMCA using purified components. (PDF 232 kb)
Rights and permissions
About this article
Cite this article
Morales, R., Duran-Aniotz, C., Diaz-Espinoza, R. et al. Protein misfolding cyclic amplification of infectious prions. Nat Protoc 7, 1397–1409 (2012). https://doi.org/10.1038/nprot.2012.067
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.067
This article is cited by
-
Nasal bots carry relevant titers of CWD prions in naturally infected white-tailed deer
EMBO Reports (2024)
-
Ticks harbor and excrete chronic wasting disease prions
Scientific Reports (2023)
-
PMCA screening of retropharyngeal lymph nodes in white-tailed deer and comparisons with ELISA and IHC
Scientific Reports (2023)
-
PMCA for ultrasensitive detection of prions and to study disease biology
Cell and Tissue Research (2023)
-
BSE can propagate in sheep co-infected or pre-infected with scrapie
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.