Abstract
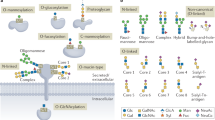
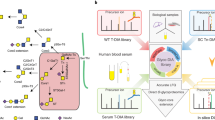
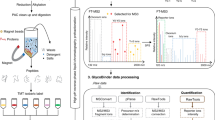
The comprehensive analysis of protein glycosylation is a major requirement for understanding glycoprotein function in biological systems, and is a prerequisite for producing recombinant glycoprotein therapeutics. This protocol describes workflows for the characterization of glycopeptides and their site-specific heterogeneity, showing examples of the analysis of recombinant human erythropoietin (rHuEPO), α1-proteinase inhibitor (A1PI) and immunoglobulin (IgG). Glycoproteins of interest can be proteolytically digested either in solution or in-gel after electrophoretic separation, and the (glyco)peptides are analyzed by capillary/nano-liquid chromatography–electrospray ionization tandem mass spectrometry (LC-ESI-MS/MS). If required, specific glycopeptide enrichment steps, such as hydrophilic interaction liquid chromatography (HILIC), can also be performed. Particular emphasis is placed on data interpretation and the determination of site-specific glycan heterogeneity. The described workflow takes approximately 3–5 d, including sample preparation and data analysis. The data obtained from analyzing released glycans of rHuEPO and IgG, described in the second protocol of this series (10.1038/nprot.2012.063), provide complementary detailed glycan structural information that facilitates characterization of the glycopeptides.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Morell, A.G., Gregoriadis, G., Scheinberg, I.H., Hickman, J. & Ashwell, G. The role of sialic acid in determining the survival of glycoproteins in the circulation. J. Biol. Chem. 246, 1461–1467 (1971).
Ma, B.Y. et al. CD28 T cell co-stimulatory receptor function is negatively regulated by N-linked carbohydrates. Biochem. Biophys. Res. Commun. 317, 60–67 (2004).
Takahashi, M., Tsuda, T., Ikeda, Y., Honke, K. & Taniguchi, N. Role of N-glycans in growth factor signaling. Glycoconj. J. 20, 207–212 (2004).
Kolarich, D., Lepenies, B. & Seeberger, P.H. Glycomics, glycoproteomics and the immune system. Curr. Opin. Chem. Biol. 16, 214–220 (2012).
Kolarich, D. et al. Biochemical, molecular characterization, and glycoproteomic analyses of α(1)-proteinase inhibitor products used for replacement therapy. Transfusion 46, 1959–1977 (2006).
Jensen, P.H., Karlsson, N.G., Kolarich, D. & Packer, N.H. Structural analysis of N- and O-glycans released from glycoproteins. Nat. Protoc. 7, 1299–1310 (2012).
Van Droogenbroeck, B. et al. Aberrant localization and underglycosylation of highly accumulating single-chain Fv-Fc antibodies in transgenic Arabidopsis seeds. Proc. Natl. Acad. Sci. USA 104, 1430–1435 (2007).
Deshpande, N., Jensen, P.H., Packer, N.H. & Kolarich, D. GlycoSpectrumScan: fishing glycopeptides from MS spectra of protease digests of human colostrum sIgA. J. Proteome Res. 9, 1063–1075 (2010).
Kolarich, D., Weber, A., Turecek, P.L., Schwarz, H.P. & Altmann, F. Comprehensive glyco-proteomic analysis of human α1-antitrypsin and its charge isoforms. Proteomics 6, 3369–3380 (2006).
Stadlmann, J., Pabst, M., Kolarich, D., Kunert, R. & Altmann, F. Analysis of immunoglobulin glycosylation by LC-ESI-MS of glycopeptides and oligosaccharides. Proteomics 8, 2858–2871 (2008).
Christiansen, M.N., Kolarich, D., Nevalainen, H., Packer, N.H. & Jensen, P.H. Challenges of determining O-glycopeptide heterogeneity: a fungal glucanase model system. Anal. Chem. 82, 3500–3509 (2010).
Kolarich, D. et al. Glycoproteomic characterization of butyrylcholinesterase from human plasma. Proteomics 8, 254–263 (2008).
Kolarich, D., Leonard, R., Hemmer, W. & Altmann, F. The N-glycans of yellow jacket venom hyaluronidases and the protein sequence of its major isoform in Vespula vulgaris. FEBS J. 272, 5182–5190 (2005).
Kolarich, D., Altmann, F. & Sunderasan, E. Structural analysis of the glycoprotein allergen Hev b 4 from natural rubber latex by mass spectrometry. Biochim. Biophys. Acta. 1760, 715–720 (2006).
Wada, Y. et al. Comparison of the methods for profiling glycoprotein glycans—HUPO Human Disease Glycomics/Proteome Initiative multi-institutional study. Glycobiology 17, 411–422 (2007).
Wada, Y. et al. Comparison of methods for profiling O-glycosylation: human proteome organisation human disease glycomics/proteome initiative multi-institutional study of IgA1. Mol. Cell Proteomics 9, 719–727 (2010).
Ito, S., Hayama, K. & Hirabayashi, J. Enrichment strategies for glycopeptides. Methods Mol. Biol. 534, 195–203 (2009).
Huhn, C., Selman, M.H., Ruhaak, L.R., Deelder, A.M. & Wuhrer, M. IgG glycosylation analysis. Proteomics 9, 882–913 (2009).
Selman, M.H. et al. Immunoglobulin G glycopeptide profiling by matrix-assisted laser desorption ionization Fourier transform ion cyclotron resonance mass spectrometry. Anal. Chem. 82, 1073–1081 (2010).
Mysling, S., Palmisano, G., Hojrup, P. & Thaysen-Andersen, M. Utilizing ion-pairing hydrophilic interaction chromatography solid phase extraction for efficient glycopeptide enrichment in glycoproteomics. Anal. Chem. 82, 5598–5609 (2010).
Thaysen-Andersen, M., Mysling, S. & Hojrup, P. Site-specific glycoprofiling of N-linked glycopeptides using MALDI-TOF MS: strong correlation between signal strength and glycoform quantities. Anal. Chem. 81, 3933–3943 (2009).
Peltoniemi, H., Joenvaara, S. & Renkonen, R. De novo glycan structure search with the CID MS/MS spectra of native N-glycopeptides. Glycobiology 19, 707–714 (2009).
Demelbauer, U.M., Zehl, M., Plematl, A., Allmaier, G. & Rizzi, A. Determination of glycopeptide structures by multistage mass spectrometry with low-energy collision-induced dissociation: comparison of electrospray ionization quadrupole ion trap and matrix-assisted laser desorption/ionization quadrupole ion trap reflectron time-of-flight approaches. Rapid Commun. Mass Spectrom. 18, 1575–1582 (2004).
Morelle, W. & Michalski, J.C. Analysis of protein glycosylation by mass spectrometry. Nat. Protoc. 2, 1585–1602 (2007).
O'Connor, P.B. & Costello, C.E. A high pressure matrix-assisted laser desorption/ionization Fourier transform mass spectrometry ion source for thermal stabilization of labile biomolecules. Rapid Commun. Mass Spectrom. 15, 1862–1868 (2001).
Wang, H. et al. Integrated mass spectrometry–based analysis of plasma glycoproteins and their glycan modifications. Nat. Protoc. 6, 253–269 (2011).
Yu, K. et al. Ultra-performance liquid chromatography/tandem mass spectrometric quantification of structurally diverse drug mixtures using an ESI-APCI multimode ionization source. Rapid Commun. Mass Spectrom. 21, 893–902 (2007).
Castellanos-Serra, L., Proenza, W., Huerta, V., Moritz, R.L. & Simpson, R.J. Proteome analysis of polyacrylamide gel-separated proteins visualized by reversible negative staining using imidazole-zinc salts. Electrophoresis 20, 732–737 (1999).
An, H.J., Peavy, T.R., Hedrick, J.L. & Lebrilla, C.B. Determination of N-glycosylation sites and site heterogeneity in glycoproteins. Anal. Chem. 75, 5628–5637 (2003).
Nwosu, C.C. et al. Simultaneous and extensive site-specific N- and O-glycosylation analysis in protein mixtures. J. Proteome Res. 10, 2612–2624 (2011).
Zauner, G., Koeleman, C.A., Deelder, A.M. & Wuhrer, M. Protein glycosylation analysis by HILIC-LC-MS of Proteinase K–generated N- and O-glycopeptides. J. Sep. Sci. 33, 903–910 (2010).
Dodds, E.D., Seipert, R.R., Clowers, B.H., German, J.B. & Lebrilla, C.B. Analytical performance of immobilized pronase for glycopeptide footprinting and implications for surpassing reductionist glycoproteomics. J. Proteome Res. 8, 502–512 (2009).
Taouatas, N., Drugan, M.M., Heck, A.J. & Mohammed, S. Straightforward ladder sequencing of peptides using a Lys-N metalloendopeptidase. Nat. Methods 5, 405–407 (2008).
Calvano, C.D., Zambonin, C.G. & Jensen, O.N. Assessment of lectin and HILIC based enrichment protocols for characterization of serum glycoproteins by mass spectrometry. J. Proteomics 71, 304–317 (2008).
Wang, Y., Wu, S.L. & Hancock, W.S. Monitoring of glycoprotein products in cell culture lysates using lectin affinity chromatography and capillary HPLC coupled to electrospray linear ion trap-Fourier transform mass spectrometry (LTQ/FTMS). Biotechnol. Prog. 22, 873–880 (2006).
Lee, A. et al. The lectin riddle: glycoproteins fractionated from complex mixtures have similar glycomic profiles. OMICS 14, 487–499 (2010).
Lee, A. et al. Rat liver membrane glycoproteome: enrichment by phase partitioning and glycoprotein capture. J. Proteome Res. 8, 770–781 (2009).
Larsen, M.R., Cordwell, S.J. & Roepstorff, P. Graphite powder as an alternative or supplement to reversed-phase material for desalting and concentration of peptide mixtures prior to matrix-assisted laser desorption/ionization-mass spectrometry. Proteomics 2, 1277–1287 (2002).
Larsen, M.R., Hojrup, P. & Roepstorff, P. Characterization of gel-separated glycoproteins using two-step proteolytic digestion combined with sequential microcolumns and mass spectrometry. Mol. Cell Proteomics 4, 107–119 (2005).
Lewandrowski, U. & Sickmann, A. Online dual gradient reversed-phase/porous graphitized carbon nanoHPLC for proteomic applications. Anal. Chem. 82, 5391–5396 (2010).
Wagner-Rousset, E. et al. The way forward, enhanced characterization of therapeutic antibody glycosylation: comparison of three level mass spectrometry–based strategies. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 872, 23–37 (2008).
Pan, S., Chen, R., Aebersold, R. & Brentnall, T.A. Mass spectrometry–based glycoproteomics—from a proteomics perspective. Mol. Cell Proteomics 10, R110 003251 (2011).
Jensen, O.N., Shevchenko, A. & Mann, M. in Protein Structure: A Practical Approach (ed. Creighton, T.E.) 29–57 (IRL Press, 1997).
Hagglund, P. et al. An enzymatic deglycosylation scheme enabling identification of core fucosylated N-glycans and O-glycosylation site mapping of human plasma proteins. J. Proteome Res. 6, 3021–3031 (2007).
Cooper, C.A., Gasteiger, E. & Packer, N.H. GlycoMod—a software tool for determining glycosylation compositions from mass spectrometric data. Proteomics 1, 340–349 (2001).
Wada, Y. et al. Comparison of methods for profiling O-glycosylation. Mol. Cell Proteomics 9, 719–727 (2010).
Acknowledgements
D.K. was supported by an Erwin Schrödinger Fellowship from the Austrian Science Fund (grant J2661) and Macquarie University. P.H.J. was supported by the Danish Agency for Science, Technology and Innovation (grant 272-07-0066). The technical help of T. Dalik is as always highly appreciated.
Author information
Authors and Affiliations
Contributions
D.K. performed the experiments on the therapeutic proteins and developed the initial protocol. F.A. and D.K. crucially validated MS data interpretation. D.K. and P.H.J. performed the experiments on the protein complex mixtures and optimized the protocol. D.K. and N.H.P. wrote the paper and prepared it for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kolarich, D., Jensen, P., Altmann, F. et al. Determination of site-specific glycan heterogeneity on glycoproteins. Nat Protoc 7, 1285–1298 (2012). https://doi.org/10.1038/nprot.2012.062
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.062
This article is cited by
-
Machine Learning Classifies Core and Outer Fucosylation of N-Glycoproteins Using Mass Spectrometry
Scientific Reports (2020)
-
Measurement of the total angiotensinogen and its reduced and oxidised forms in human plasma using targeted LC-MS/MS
Analytical and Bioanalytical Chemistry (2019)
-
Cell-free protein synthesis as a novel tool for directed glycoengineering of active erythropoietin
Scientific Reports (2018)
-
Global site-specific analysis of glycoprotein N-glycan processing
Nature Protocols (2018)
-
Global site-specific N-glycosylation analysis of HIV envelope glycoprotein
Nature Communications (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.