Abstract
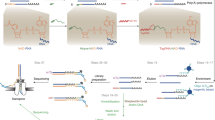
Cap-analysis gene expression (CAGE) provides accurate high-throughput measurement of RNA expression. CAGE allows mapping of all the initiation sites of both capped coding and noncoding RNAs. In addition, transcriptional start sites within promoters are characterized at single-nucleotide resolution. The latter allows the regulatory inputs driving gene expression to be studied, which in turn enables the construction of transcriptional networks. Here we provide an optimized protocol for the construction of CAGE libraries on the basis of the preparation of 27-nt-long tags corresponding to initial bases at the 5′ ends of capped RNAs. We have optimized the methods using simple steps based on filtration, which altogether takes 4 d to complete. The CAGE tags can be readily sequenced with Illumina sequencers, and upon modification they are also amenable to sequencing using other platforms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Schena, M., Shalon, D., Davis, R.W. & Brown, P.O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Cheng, J. et al. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science 308, 1149–1154 (2005).
Forrest, A.R. & Carninci, P. Whole genome transcriptome analysis. RNA Biol. 6, 107–112 (2009).
Velculescu, V.E., Zhang, L., Vogelstein, B. & Kinzler, K.W. Serial analysis of gene expression. Science 270, 484–487 (1995).
Kodzius, R. et al. CAGE: cap analysis of gene expression. Nat. Methods 3, 211–222 (2006).
Kanamori-Katayama, M. et al. Unamplified cap analysis of gene expression on a single-molecule sequencer. Genome Res. 21, 1150–1159 (2011).
Kawaji, H. et al. Dynamic usage of transcription start sites within core promoters. Genome Biol. 7, R118 (2006).
Ponjavic, J. et al. Transcriptional and structural impact of TATA-initiation site spacing in mammalian core promoters. Genome Biol. 7, R78 (2006).
Frith, M.C. et al. Evolutionary turnover of mammalian transcription start sites. Genome Res. 16, 713–2 (2006).
Hoskins, R.A. et al. Genome-wide analysis of promoter architecture in Drosophila melanogaster. Genome Res. 21, 182–192 (2011).
Carninci, P. et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nat. Genet. 38, 626–635 (2006).
Gustincich, S. et al. The complexity of the mammalian transcriptome. J. Physiol. 575, 321–332 (2006).
Vitezic, M. et al. Building promoter aware transcriptional regulatory networks using siRNA perturbation and deepCAGE. Nucleic Acids Res. 38, 8141–8148 (2010).
Frith, M.C. et al. A code for transcription initiation in mammalian genomes. Genome Res. 18, 1–12 (2008).
Suzuki, H. et al. The transcriptional network that controls growth arrest and differentiation in a human myeloid leukemia cell line. Nat. Genet. 41, 553–562 (2009).
Faulkner, G.J. et al. The regulated retrotransposon transcriptome of mammalian cells. Nat. Genet. 41, 563–571 (2009).
Hestand, M.S. et al. Tissue-specific transcript annotation and expression profiling with complementary next-generation sequencing technologies. Nucleic Acids Res. 38, e165 (2010).
Wei, C.L. et al. 5′ Long serial analysis of gene expression (LongSAGE) and 3′ LongSAGE for transcriptome characterization and genome annotation. Proc. Natl. Acad. Sci. USA. 101, 11701–11706 (2004).
Valen, E. et al. Genome-wide detection and analysis of hippocampus core promoters using DeepCAGE. Genome Res. 19, 255–265 (2009).
Shiraki, T. et al. Cap analysis gene expression for high-throughput analysis of transcriptional starting point and identification of promoter usage. Proc. Natl. Acad. Sci. USA 100, 15776–15781 (2003).
Myers, R.M. et al. A user's guide to the encyclopedia of DNA elements (ENCODE). PLoS Biol. 9, e1001046 (2011).
Takahashi, H., Kato, S., Murata, M. & Carninci, P. CAGE (Cap Analysis of Gene Expression): a protocol for the detection of promoter and transcriptional networks. Methods Mol. Biol. 786, 181–200 (2012).
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005).
Carninci, P. et al. Thermostabilization and thermoactivation of thermolabile enzymes by trehalose and its application for the synthesis of full length cDNA. Proc. Natl. Acad. Sci. USA 95, 520–524 (1998).
Carninci, P., Shiraki, T., Mizuno, Y., Muramatsu, M. & Hayashizaki, Y. Extra-long first-strand cDNA synthesis. Biotechniques 32, 984–985 (2002).
Carninci, P. et al. High-efficiency full-length cDNA cloning by biotinylated CAP trapper. Genomics 37, 327–336 (1996).
Shibata, K. et al. RIKEN integrated sequence analysis (RISA) system—384-format sequencing pipeline with 384 multicapillary sequencer. Genome Res. 10, 1757–1771 (2000).
Plessy, C. et al. Linking promoters to functional transcripts in small samples with nanoCAGE and CAGEscan. Nat. Methods 7, 528–534 (2010).
Maeda, N. et al. Development of a DNA barcode tagging method for monitoring dynamic changes in gene expression by using an ultra high-throughput sequencer. Biotechniques 45, 95–97 (2008).
Janscak, P., Sandmeier, U., Szczelkun, M.D. & Bickle, T.A. Subunit assembly and mode of DNA cleavage of the type III restriction endonucleases EcoP1I and EcoP15I. J. Mol. Biol. 306, 417–431 (2001).
Raghavendra, N.K. & Rao, D.N. Exogenous AdoMet and its analogue sinefungin differentially influence DNA cleavage by R.EcoP15I–usefulness in SAGE. Biochem. Biophys. Res. Commun. 334, 803–811 (2005).
Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Lassmann, T., Hayashizaki, Y. & Daub, C.O. TagDust—a program to eliminate artifacts from next generation sequencing data. Bioinformatics 25, 2839–2840 (2009).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Fujita, P.A. et al. The UCSC Genome Browser database: update 2011. Nucleic Acids Res. 39, D876–D882 (2011).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Carninci, P. Cap-analysis Gene Expression (CAGE): The Science of Decoding Gene Transcription (Pan Stanford, 2010).
Li, Q., Brown, J.B., Huang, H. & Bickel, P.J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011).
Acknowledgements
This work was founded by a Research Grant for the RIKEN Omics Science Center from the Japanese Ministry of Education, Culture, Sports, Science and Technology (to Y.H.) This project was also supported by the US National Human Genome Research Institute grant no. U54 HG004557. We thank S. Kato for experimental support, J. Severin for the genome browser, the RIKEN Genome Network Analysis Service for sequencing and basic bioinformatics analysis, and all our colleagues at the Omics Science Center for valuable feedback during the development of the methodology.
Author information
Authors and Affiliations
Contributions
H.T. performed most experiments. M.M. performed the background reduction experiment. T.L. performed computations analysis. H.T. and P.C. wrote the manuscript. P.C. designed the project.
Corresponding author
Ethics declarations
Competing interests
P.C. is an inventor on various patents owned by RIKEN and Dnaform on the Cap-trapper technology, full-length cDNA cloning technologies and the CAGE technology.
Supplementary information
Supplementary Fig. 1
Oligo-dT priming enhances the capture of CAGE tags on exons and 3′ UTRs. CAGE libraries made from THP-1 cells. Data was displayed with the ZENBU genome browser (J. Severin, unpublished data). (a) The Actin beta gene is transcribed from right to left (violet arrow) on chromosome 7. (b) GAPDH gene is transcribed from left to right (green arrow) on chromosome 12. CAGE libraries were primed RT reaction with (1) random and oligodT (ratio 4:1) primers. (2) oligodT primers only and (3) random primers only. Both panels indicate that oligodT primers could enhance the capture of transcripts on 3′ exons and on internal exons, compared to random primer alone. (PNG 652 kb)
Supplementary Data 1
The make_ctss script, which is used to cluster the CTSS (Step 65). (TXT 1 kb)
Rights and permissions
About this article
Cite this article
Takahashi, H., Lassmann, T., Murata, M. et al. 5′ end–centered expression profiling using cap-analysis gene expression and next-generation sequencing. Nat Protoc 7, 542–561 (2012). https://doi.org/10.1038/nprot.2012.005
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.005
This article is cited by
-
The neuropsychopharmacology of acetyl-L-carnitine (LAC): basic, translational and therapeutic implications
Discover Mental Health (2024)
-
A multi-omics dataset for the analysis of frontotemporal dementia genetic subtypes
Scientific Data (2023)
-
Spurious transcription causing innate immune responses is prevented by 5-hydroxymethylcytosine
Nature Genetics (2023)
-
TNRC18 engages H3K9me3 to mediate silencing of endogenous retrotransposons
Nature (2023)
-
Tumor-promoting function and regulatory landscape of PD-L2 in B-cell lymphoma
Leukemia (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.