Abstract
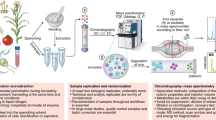
metaXCMS is a software program for the analysis of liquid chromatography/mass spectrometry–based untargeted metabolomic data. It is designed to identify the differences between metabolic profiles across multiple sample groups (e.g., 'healthy' versus 'active disease' versus 'inactive disease'). Although performing pairwise comparisons alone can provide physiologically relevant data, these experiments often result in hundreds of differences, and comparison with additional biologically meaningful sample groups can allow for substantial data reduction. By performing second-order (meta-) analysis, metaXCMS facilitates the prioritization of interesting metabolite features from large untargeted metabolomic data sets before the rate-limiting step of structural identification. Here we provide a detailed step-by-step protocol for going from raw mass spectrometry data to metaXCMS results, visualized as Venn diagrams and exported Microsoft Excel spreadsheets. There is no upper limit to the number of sample groups or individual samples that can be compared with the software, and data from most commercial mass spectrometers are supported. The speed of the analysis depends on computational resources and data volume, but will generally be less than 1 d for most users. metaXCMS is freely available at http://metlin.scripps.edu/metaxcms/.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Weckwerth, W. Unpredictability of metabolism—the key role of metabolomics science in combination with next-generation genome sequencing. Anal. Bioanal. Chem. 400, 1967–1978 (2011).
Baker, M. Metabolomics: from small molecules to big ideas. Nat. Meth. 8, 117–121 (2011).
Yanes, O. et al. Metabolic oxidation regulates embryonic stem cell differentiation. Nat. Chem. Biol. 6, 411–417 (2010).
Wikoff, W.R. et al. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc. Natl. Acad. Sci. USA 106, 3698–3703 (2009).
Wikoff, W.R., Pendyala, G., Siuzdak, G. & Fox, H.S. Metabolomic analysis of the cerebrospinal fluid reveals changes in phospholipase expression in the CNS of SIV-infected macaques. J. Clin. Invest. 118, 2661–2669 (2008).
Vinayavekhin, N. & Saghatelian, A. Untargeted metabolomics. Curr. Protoc. Mol. Biol. 90, 30.1.1–30.1.24 (2010).
Wikoff, W.R., Nagle, M.A., Kouznetsova, V.L., Tsigelny, I.F. & Nigam, S.K. Untargeted metabolomics identifies enterobiome metabolites and putative uremic toxins as substrates of organic anion transporter 1 (Oat1). J. Proteome Res. 10, 2842–2851 (2011).
Vinayavekhin, N., Homan, E.A. & Saghatelian, A. Exploring disease through metabolomics. ACS Chem. Biol. 5, 91–103 (2010).
McKnight, S.L. On getting there from here. Science 330, 1338–1339 (2010).
Wang, T.J. et al. Metabolite profiles and the risk of developing diabetes. Nat. Med. 17, 448–453 (2011).
Wang, Z. et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–63 (2011).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009).
Olszewski, K.L. et al. Branched tricarboxylic acid metabolism in Plasmodium falciparum. Nature 466, 774–778 (2010).
Fernie, A.R., Trethewey, R.N., Krotzky, A.J. & Willmitzer, L. Metabolite profiling: from diagnostics to systems biology. Nat. Rev. Mol. Cell Biol. 5, 763–769 (2004).
Lu, W. et al. Metabolomic analysis via reversed-phase ion-pairing liquid chromatography coupled to a stand-alone Orbitrap mass spectrometer. Anal. Chem. 82, 3212–3221 (2010).
Buscher, J.M., Czernik, D., Ewald, J.C., Sauer, U. & Zamboni, N. Cross-platform comparison of methods for quantitative metabolomics of primary metabolism. Anal. Chem. 81, 2135–2143 (2009).
Smith, C.A., Want, E.J., O′Maille, G., Abagyan, R. & Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Yanes, O., Tautenhahn, R., Patti, G.J. & Siuzdak, G. Expanding coverage of the metabolome for global metabolite profiling. Anal. Chem. 83, 2152–2161 (2011).
Want, E.J. et al. Solvent-dependent metabolite distribution, clustering, and protein extraction for serum profiling with mass spectrometry. Anal. Chem. 78, 743–752 (2006).
Lommen, A. MetAlign: interface-driven, versatile metabolomics tool for hyphenated full-scan mass spectrometry data preprocessing. Anal. Chem. 81, 3079–3086 (2009).
Katajamaa, M., Miettinen, J. & Oresic, M. MZmine: toolbox for processing and visualization of mass spectrometry based molecular profile data. Bioinformatics 22, 634–636 (2006).
Böttcher, C. et al. The multifunctional enzyme CYP71B15 (PHYTOALEXIN DEFICIENT3) converts cysteine-indole-3-acetonitrile to camalexin in the indole-3-acetonitrile metabolic network of Arabidopsis thaliana. Plant Cell 21, 1830–1845 (2009).
Tautenhahn, R. et al. metaXCMS: second-order analysis of untargeted metabolomics data. Anal. Chem. 83, 696–700 (2011).
Normand, S.L. Meta-analysis: formulating, evaluating, combining, and reporting. Stat. Med. 18, 321–359 (1999).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
de Bakker, P.I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
de Bakker, P.I. et al. Efficiency and power in genetic association studies. Nat. Genet. 37, 1217–1223 (2005).
Chu, Y.C. et al. Effect of genetic knockout or pharmacologic inhibition of neuronal nitric oxide synthase on complete Freund′s adjuvant-induced persistent pain. Pain 119, 113–123 (2005).
Bolcskei, K., Petho, G. & Szolcsanyi, J. Noxious heat threshold measured with slowly increasing temperatures: novel rat thermal hyperalgesia models. Methods Mol. Biol. 617, 57–66 (2010).
Kyburz, D. & Corr, M. The KRN mouse model of inflammatory arthritis. Springer Semin. Immunopathol. 25, 79–90 (2003).
Goo, Y.A. et al. Proteomic analysis of an extreme halophilic archaeon, Halobacterium sp. NRC-1. Mol. Cell Proteomics 2, 506–524 (2003).
Kaur, A. et al. A systems view of haloarchaeal strategies to withstand stress from transition metals. Genome Res. 16, 841–854 (2006).
Branca, F., Hanley, A.B., Pool-Zobel, B. & Verhagen, H. Biomarkers in disease and health. Br. J. Nutr. 86 (suppl. 1): S55–S92 (2001).
Cantor, R.M., Lange, K. & Sinsheimer, J.S. Prioritizing GWAS results: a review of statistical methods and recommendations for their application. Am. J. Hum. Genet. 86, 6–22 (2010).
Crews, B. et al. Variability analysis of human plasma and cerebral spinal fluid reveals statistical significance of changes in mass spectrometry–based metabolomics data. Anal. Chem. 81, 8538–8544 (2009).
Kalisiak, J. et al. Identification of a new endogenous metabolite and the characterization of its protein interactions through an immobilization approach. J. Am. Chem. Soc. 131, 378–386 (2009).
Wise, L.H., Lanchbury, J.S. & Lewis, C.M. Meta-analysis of genome searches. Ann. Hum. Genet. 63, 263–272 (1999).
Evangelou, E., Maraganore, D.M. & Ioannidis, J.P. Meta-analysis in genome-wide association datasets: strategies and application in Parkinson disease. PLoS ONE 2, e196 (2007).
Want, E.J., Nordstrom, A., Morita, H. & Siuzdak, G. From exogenous to endogenous: the inevitable imprint of mass spectrometry in metabolomics. J. Proteome Res. 6, 459–468 (2007).
Tautenhahn, R., Böttcher, C. & Neumann, S. Highly sensitive feature detection for high-resolution LC/MS. BMC Bioinformatics 9, 504 (2008).
Prince, J.T. & Marcotte, E.M. Chromatographic alignment of ESI-LC-MS proteomics data sets by ordered bijective interpolated warping. Anal. Chem. 78, 6140–6152 (2006).
Masson, P., Spagou, K., Nicholson, J.K. & Want, E.J. Technical and biological variation in UPLC-MS–based untargeted metabolic profiling of liver extracts: application in an experimental toxicity study on galactosamine. Anal. Chem. 83, 1116–1123 (2011).
Acknowledgements
This work was supported by the California Institute of Regenerative Medicine (grant TR1-01219), the US National Institutes of Health (grants R24 EY017540-04, P30 MH062261-10 and P01 DA026146-02) and a US National Institutes of Health/National Institute on Aging grant (L30 AG0 038036; to G.J.P.). Financial support was also received from the US Department of Energy (grants FG02-07ER64325 and DE-AC0205CH11231).
Author information
Authors and Affiliations
Contributions
G.J.P., R.T. and G.S. contributed to the development of the protocol and the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Patti, G., Tautenhahn, R. & Siuzdak, G. Meta-analysis of untargeted metabolomic data from multiple profiling experiments. Nat Protoc 7, 508–516 (2012). https://doi.org/10.1038/nprot.2011.454
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2011.454
This article is cited by
-
Metabolome panels as potential noninvasive biomarkers for primary glomerulonephritis sub-types: meta-analysis of profiling metabolomics studies
Scientific Reports (2023)
-
Small molecule metabolites: discovery of biomarkers and therapeutic targets
Signal Transduction and Targeted Therapy (2023)
-
Quick tips for re-using metabolomics data
Nature Cell Biology (2022)
-
Cognitive analysis of metabolomics data for systems biology
Nature Protocols (2021)
-
Salivary metabolomics for the diagnosis of periodontal diseases: a systematic review with methodological quality assessment
Metabolomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.