Abstract
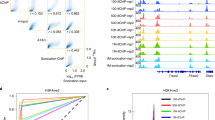
Chromatin immunoprecipitation (ChIP) combined with high-throughput sequencing (ChIP-seq) has become the gold standard for whole-genome mapping of protein-DNA interactions. However, conventional ChIP protocols necessitate the use of large numbers of cells, and library preparation steps associated with current high-throughput sequencing platforms require substantial amounts of DNA; both of these factors preclude the application of ChIP-seq technology to many biologically important but rare cell types. Here we describe a nano-ChIP-seq protocol that combines a high-sensitivity small-scale ChIP assay and a tailored procedure for generating high-throughput sequencing libraries from scarce amounts of ChIP DNA. In terms of the numbers of cells required, the method provides two to three orders of magnitude of improvement over the conventional ChIP-seq method and the entire procedure can be completed within 4 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Park, P.J. ChIP-seq: advantages and challenges of a maturing technology. Nat. Rev. Genet. 10, 669–680 (2009).
Dahl, J.A. & Collas, P. Q2ChIP, a quick and quantitative chromatin immunoprecipitation assay, unravels epigenetic dynamics of developmentally regulated genes in human carcinoma cells. Stem Cells 25, 1037–1046 (2007).
Acevedo, L.G. et al. Genome-scale ChIP-chip analysis using 10,000 human cells. Biotechniques 43, 791–797 (2007).
O'Neill, L.P., VerMilyea, M.D. & Turner, B.M. Epigenetic characterization of the early embryo with a chromatin immunoprecipitation protocol applicable to small cell populations. Nat. Genet. 38, 835–841 (2006).
Attema, J.L. et al. Epigenetic characterization of hematopoietic stem cell differentiation using miniChIP and bisulfite sequencing analysis. Proc. Natl. Acad. Sci. USA 104, 12371–12376 (2007).
Goren, A. et al. Chromatin profiling by directly sequencing small quantities of immunoprecipitated DNA. Nat. Methods 7, 47–49 (2009).
Adli, M., Zhu, J. & Bernstein, B.E. Genome-wide chromatin maps derived from limited numbers of hematopoietic progenitors. Nat. Methods 7, 615–618 (2010).
Shankaranarayanan, P. et al. Single-tube linear DNA amplification (LinDA) for robust ChIP-seq. Nat. Methods 8, 565–567 (2011).
Dahl, J.A. & Collas, P. A rapid micro chromatin immunoprecipitation assay (microChIP). Nat. Protoc. 3, 1032–1045 (2008).
Lieb, J.D., Liu, X., Botstein, D. & Brown, P.O. Promoter-specific binding of Rap1 revealed by genome-wide maps of protein-DNA association. Nat. Genet. 28, 327–334 (2001).
Acknowledgements
We acknowledge all the members of the Bernstein Lab for constructive discussions during the method development. We thank M. Coyne, S. Furuyama, S. Gillespie and N.A. Okan for their critical reading of the manuscript and J. Zhu, N. Shoresh and T. Mikkelsen for computational assistance. This research was supported by funds from the Starr Cancer Consortium, a Charles E. Culpeper Scholarship, the US National Human Genome Research Institute, the National Institutes of Health Roadmap for Epigenomics and the National Heart, Lung and Blood Institute.
AUTHOR CONTRIBUTIONS
M.A. and B.E.B. designed the method and analyzed the data. M.A. performed the experiments and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
M.A. and B.E.B. have filed a patent application describing the methods presented here. Patent application #: 12 / 699508 (USA)
Rights and permissions
About this article
Cite this article
Adli, M., Bernstein, B. Whole-genome chromatin profiling from limited numbers of cells using nano-ChIP-seq. Nat Protoc 6, 1656–1668 (2011). https://doi.org/10.1038/nprot.2011.402
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2011.402
This article is cited by
-
Transcription factor chromatin profiling genome-wide using uliCUT&RUN in single cells and individual blastocysts
Nature Protocols (2021)
-
Rapid method for chromatin immunoprecipitation (ChIP) assay in a dimorphic fungus, Candida albicans
Journal of Microbiology (2020)
-
Profiling chromatin states using single-cell itChIP-seq
Nature Cell Biology (2019)
-
Purification of nanogram-range immunoprecipitated DNA in ChIP-seq application
BMC Genomics (2017)
-
Time-resolved analysis of DNA-protein interactions in living cells by UV laser pulses
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.