Abstract
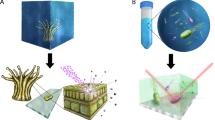
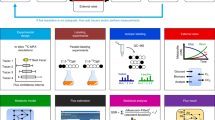
We describe a stable isotope probing (SIP) technique that was developed to link microbe-specific metabolic function to phylogenetic information. Carbon (13C)- or nitrogen (15N)-labeled substrates (typically with >98% heavy label) were used in cultivation experiments and the heavy isotope incorporation into proteins (protein-SIP) on growth was determined. The amount of incorporation provides a measure for assimilation of a substrate, and the sequence information from peptide analysis obtained by mass spectrometry delivers phylogenetic information about the microorganisms responsible for the metabolism of the particular substrate. In this article, we provide guidelines for incubating microbial cultures with labeled substrates and a protocol for protein-SIP. The protocol guides readers through the proteomics pipeline, including protein extraction, gel-free and gel-based protein separation, the subsequent mass spectrometric analysis of peptides and the calculation of the incorporation of stable isotopes into peptides. Extraction of proteins and the mass fingerprint measurements of unlabeled and labeled fractions can be performed in 2–3 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Crick, F. Central dogma of molecular biology. Nature 227, 561–563 (1970).
Radajewski, S., Ineson, P., Parekh, N.R. & Murrell, J.C. Stable-isotope probing as a tool in microbial ecology. Nature 403, 646–649 (2000).
Manefield, M., Whiteley, A.S., Griffiths, R.I. & Bailey, M.J. RNA stable isotope probing, a novel means of linking microbial community function to phylogeny. Appl. Environ. Microbiol. 68, 5367–5373 (2002).
Boschker, H.T.S. et al. Direct linking of microbial populations to specific biogeochemical processes by C-13-labelling of biomarkers. Nature 392, 801–805 (1998).
Dumont, M.G. & Murrell, J.C. Stable isotope probing—linking microbial identity to function. Nat. Rev. Microbiol. 3, 499–504 (2005).
Neufeld, J.D., Dumont, M.G., Vohra, J. & Murrell, J.C. Methodological considerations for the use of stable isotope probing in microbial ecology. Microb. Ecol. 53, 435–442 (2007).
Evershed, R.P. et al. 13C-Labelling of lipids to investigate microbial communities in the environment. Curr. Opin. Biotechnol. 17, 72–82 (2006).
Jehmlich, N. et al. Incorporation of carbon and nitrogen atoms into proteins measured by protein-based stable isotope probing (Protein-SIP). Rapid Commun. Mass Spectrom. 22, 2889–2897 (2008).
Jehmlich, N., Schmidt, F., von Bergen, M., Richnow, H.H. & Vogt, C. Protein-based stable isotope probing (Protein-SIP) reveals active species within anoxic mixed cultures. ISME J. 2, 1122–1133 (2008).
Bastida, F. et al. Elucidating MTBE degradation in a mixed consortium using a multidisciplinary approach. FEMS Microbiol. Ecol. 73, 370–84 (2010).
Hu, L., Ye, M., Jiang, X., Feng, S. & Zou, H. Advances in hyphenated analytical techniques for shotgun proteome and peptidome analysis—a review. Anal. Chim. Acta. 598, 193–204 (2007).
Benndorf, D., Balcke, G.U., Harms, H. & von Bergen, M. Functional metaproteome analysis of protein extracts from contaminated soil and groundwater. ISME J. 1, 224–234 (2007).
Benndorf, D. et al. Improving protein extraction and separation methods for investigating the metaproteome of anaerobic benzene communities within sediments. Biodegradation 20, 737–750 (2009).
Keller, M. & Hettich, R. Environmental proteomics: a paradigm shift in characterizing microbial activities at the molecular level. Microbiol. Mol. Biol. Rev. 73, 62–70 (2009).
Purdy, K.J. Nucleic acid recovery from complex environmental samples. Methods Enzymol. 397, 271–292 (2005).
Jehmlich, N. et al. Comparison of methods for simultaneous identification of bacterial species and determination of metabolic activity by protein-based stable isotope probing (Protein-SIP) experiments. Rapid Commun. Mass Spectrom. 23, 1871–1878 (2009).
Gustavsson, N. et al. A proteomic method for the analysis of changes in protein concentrations in response to systemic perturbations using metabolic incorporation of stable isotopes and mass spectrometry. Proteomics 5, 3563–3570 (2005).
MacCoss, M.J., Wu, C.C., Matthews, D.E. & Yates, J.R. III . Measurement of the isotope enrichment of stable isotope-labeled proteins using high-resolution mass spectra of peptides. Anal. Chem. 77, 7646–7653 (2005).
Karty, J.A., Ireland, M.M., Brun, Y.V. & Reilly, J.P. Artifacts and unassigned masses encountered in peptide mass mapping. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 782, 363–383 (2002).
Fetzer, I. et al. Calculation of partial isotope incorporation into peptides measured by mass spectrometry. BMC Res. Notes 3, 178 (2010).
Jehmlich, N. et al. Decimal place slope, a fast and precise method for quantifying 13C incorporation levels for detecting the metabolic activity of microbial species. Mol. Cell Proteomics 9, 1221–1227 (2010).
Lueders, T., Manefield, M. & Friedrich, M.W. Enhanced sensitivity of DNA- and rRNA-based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ. Microbiol. 6, 73–78 (2004).
Webster, G. et al. A comparison of stable-isotope probing of DNA and phospholipid fatty acids to study prokaryotic functional diversity in sulfate-reducing marine sediment enrichment slurries. Environ. Microbiol. 8, 1575–1589 (2006).
Huang, W.E., Griffiths, R.I., Thompson, I.P., Bailey, M.J. & Whiteley, A.S. Raman microscopic analysis of single microbial cells. Anal. Chem. 76, 4452–4458 (2004).
Lechene, C.P., Luyten, Y., McMahon, G. & Distel, D.L. Quantitative imaging of nitrogen fixation by individual bacteria within animal cells. Science 317, 1563–1566 (2007).
Musat, N. et al. A single-cell view on the ecophysiology of anaerobic phototrophic bacteria. Proc. Natl. Acad. Sci. USA 105, 17861–17866 (2008).
Macias, M.T. Use of radionuclides in cancer research and treatment. Clin. Transl. Oncol. 11, 143–153 (2009).
Nikolausz, M. et al. Novel approach using substrate-mediated radiolabelling of RNA to link metabolic function with the structure of microbial communities. FEMS Microbiol. Lett. 274, 154–161 (2007).
Tabb, D.L. et al. Repeatability and reproducibility in proteomic identifications by liquid chromatography-tandem mass spectrometry. J. Proteome Res. 9, 761–776 (2009).
Craig, R. & Beavis, R.C. TANDEM: matching proteins with tandem mass spectra. Bioinformatics 20, 1466–1467 (2004).
Snijders, A.P.L., de Vos, M.G.J., de Koning, B. & Wright, P.C. A fast method for quantitative proteomics based on a combination between two-dimensional electrophoresis and N-15-metabolic labelling. Electrophoresis 26, 3191–3199 (2005).
Senko, M.W., Beu, S.C. & McLafferty, F.W. Determination of monoisotopic masses and ion populations for large biomolecules. Am. Soc. Mass Spectrom. 6, 229–233 (1995).
Widdel, F. & Rabus, R. Anaerobic biodegradation of satured and aromatic hydrocarbons. Curr. Opin. Biotechnol. 12, 259–276 (2001).
Rabus, R. & Widdel, F. Anaerobic degradation of ethylbenzene and other aromatic hydrocarbons by new denitrifying bacteria. Arch. Microbiol. 163, 96–103 (1995).
Acknowledgements
This work was supported by funding from the Helmholtz Association. M.T. was funded by a grant from the Priority Program SPP1319 from the German Research Association. F.B. acknowledges support from Marie Curie Mobility Actions of the European Commission: Host Fellowship for the Transfer of Knowledge (ToK) ISOTONIC project (MTKD-CT-2006-042758).
Author information
Authors and Affiliations
Contributions
N.J. participated in experimental analysis, data analysis and writing; F.S. helped with data analysis and writing; M.T. took part in experimental analysis and writing; J.S. was involved in data analysis and writing; F.B. conducted experimental analysis and data analysis; M.v.B. and H.-H.R. were involved in conceptualizing and designing experiments and writing; C.V. participated in experimental design, experimental analysis, data analysis and writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Jehmlich, N., Schmidt, F., Taubert, M. et al. Protein-based stable isotope probing. Nat Protoc 5, 1957–1966 (2010). https://doi.org/10.1038/nprot.2010.166
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2010.166
This article is cited by
-
Mycelial nutrient transfer promotes bacterial co-metabolic organochlorine pesticide degradation in nutrient-deprived environments
The ISME Journal (2023)
-
Rational design of a microbial consortium of mucosal sugar utilizers reduces Clostridiodes difficile colonization
Nature Communications (2020)
-
Advancing functional and translational microbiome research using meta-omics approaches
Microbiome (2019)
-
Capturing the genetic makeup of the active microbiome in situ
The ISME Journal (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.