Abstract
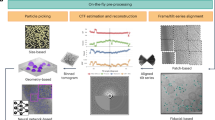
We describe a collection of standardized image processing protocols for electron microscopy single-particle analysis using the XMIPP software package. These protocols allow performing the entire processing workflow starting from digitized micrographs up to the final refinement and evaluation of 3D models. A particular emphasis has been placed on the treatment of structurally heterogeneous data through maximum-likelihood refinements and self-organizing maps as well as the generation of initial 3D models for such data sets through random conical tilt reconstruction methods. All protocols presented have been implemented as stand-alone, executable python scripts, for which a dedicated graphical user interface has been developed. Thereby, they may provide novice users with a convenient tool to quickly obtain useful results with minimum efforts in learning about the details of this comprehensive package. Examples of applications are presented for a negative stain random conical tilt data set on the hexameric helicase G40P and for a structurally heterogeneous data set on 70S Escherichia coli ribosomes embedded in vitrified ice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Henderson, R. Realizing the potential of electron cryo-microscopy. Q. Rev. Biophys. 37, 3–13 (2004).
Subramaniam, S. & Milne, J.L.S. Three-dimensional electron microscopy at molecular resolution. Annu. Rev. Biophys. Biomol. Struct. 33, 141–155 (2004).
Carragher, B . & Smith, P.R. Special issue on Advances in Computational Image Processing for Microscopy. J. Struct. Biol. 116, 2–8 (1996).
Chiu, S.L.A.W. Special Issue on Single Particle Processing (eds. Ludtke, S.J. & Chiu, W.) J. Struct. Biol. 173 (2001).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Heymann, J.B. & Belnap, D.M. Bsoft: image processing and molecular modeling for electron microscopy. J. Struct. Biol. 157, 3–18 (2007).
van Heel, M., Harauz, G., Orlova, E.V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996).
Hohn, M. et al. SPARX, a new environment for Cryo-EM image processing. J. Struct. Biol. 157, 47–55 (2007).
Sorzano, C.O.S. et al. XMIPP: a new generation of an open-source image processing package for electron microscopy. J. Struct. Biol. 148, 194–204 (2004).
Software tools for molecular microscopy. http://en.wikipedia.org/wiki/Software_tools_for_molecular_microscopy (Wikipedia, 2007).
Marabini, R. et al. Xmipp: an image processing package for electron microscopy. J. Struct. Biol. 116, 237–240 (1996).
Nickell, S. et al. Automated cryoelectron microscopy of 'single particles' applied to the 26S proteasome. FEBS Lett. 581, 2751–2756 (2007).
Martin-Benito, J. et al. Divergent substrate-binding mechanisms reveal an evolutionary specialization of eukaryotic prefoldin compared to its archaeal counterpart. Structure 15, 101–110 (2007).
Busselez, J. et al. Structural basis for the PufX-mediated dimerization of bacterial photosynthetic core complexes. Structure 15, 1674–1683 (2007).
Tato, I. et al. The ATPase activity of the DNA transporter TrwB is modulated by protein TrwA: implications for a common assembly mechanism of DNA translocating motors. J. Biol. Chem. 282, 25569–25576 (2007).
Agirrezabala, X. et al. Quasi-atomic model of bacteriophage t7 procapsid shell: insights into the structure and evolution of a basic fold. Structure 15, 461–472 (2007).
Nunez-Ramirez, R. et al. Loading a ring: structure of the Bacillus subtilis DnaB protein, a co-loader of the replicative helicase. J. Mol. Biol. 367, 764–769 (2007).
Rivera-Calzada, A., Spagnolo, L., Pearl, L.H. & Llorca, O. Structural model of full-length human Ku70-Ku80 heterodimer and its recognition of DNA and DNA-PKcs. EMBO Rep. 8, 56–62 (2007).
Arias-Palomo, E., Recuero-Checa, M.A., Bustelo, X.R. & Llorca, O. 3D structure of Syk kinase determined by single-particle electron microscopy. Biochim. Biophys. Acta 1774, 1493–1499 (2007).
Van Heel, M. Angular reconstitution: a posteriori assignment of projection directions for 3D reconstruction. Ultramicroscopy 21, 111–123 (1987).
Crowther, R.A. & Amos, L.A. Harmonic analysis of electron microscope images with rotational symmetry. J. Mol. Biol. 60, 123–130 (1971).
Pascual-Montano, A. et al. A novel neural network technique for analysis and classification of EM single-particle images. J. Struct. Biol. 133, 233–245 (2001).
Scheres, S.H.W. et al. Maximum-likelihood multi-reference refinement for electron microscopy images. J. Mol. Biol. 348, 139–149 (2005).
Radermacher, M., Wagenknecht, T., Verschoor, A. & Frank, J. Three-dimensional reconstruction from a single-exposure, random conical tilt series applied to the 50S ribosomal subunit of Escherichia coli. J. Microsc. 146, 113–136 (1987).
Scheres, S.H.W. et al. Disentangling conformational states of macromolecules in 3D-EM through likelihood optimization. Nat. Methods 4, 27–29 (2007).
Scheres, S.H.W. et al. Modeling experimental image formation for likelihood-based classification of electron microscopy data. Structure 15, 1167–1177 (2007).
Cheng, R.H. et al. Functional implications of quasi-equivalence in a T = 3 icosahedral animal virus established by cryo-electron microscopy and X-ray crystallography. Structure 2, 271–282 (1994).
Penczek, P.A., Grassucci, R.A. & Frank, J. The ribosome at improved resolution: new techniques for merging and orientation refinement in 3D cryo-electron microscopy of biological particles. Ultramicroscopy 53, 251–270 (1994).
Sorzano, C.O.S. et al. A multiresolution approach to orientation assignment in 3D electron microscopy of single particles. J. Struct. Biol. 146, 381–392 (2004).
Jonic, S. et al. Spline-based image-to-volume registration for three-dimensional electron microscopy. Ultramicroscopy 103, 303–317 (2005).
Sorzano, C.O.S., Marabini, R., Herman, G.T., Censor, Y. & Carazo, J.M. Transfer function restoration in 3D electron microscopy via iterative data refinement. Phys. Med. Biol. 49, 509–522 (2004).
Marabini, R., Herman, G.T. & Carazo, J.M. 3D reconstruction in electron microscopy using ART with smooth spherically symmetric volume elements (blobs). Ultramicroscopy 72, 53–65 (1998).
Sorzano, C.O.S. et al. The effect of overabundant projection directions on 3D reconstruction algorithms. J. Struct. Biol. 133, 108–118 (2001).
Jonic, S., Sorzano, C.O.S., Cottevieille, M., Larquet, E. & Boisset, N. A novel method for improvement of visualization of power spectra for sorting cryo-electron micrographs and their local areas. J. Struct. Biol. 157, 156–167 (2007).
Zhu, Y. et al. Automatic particle selection: results of a comparative study. J. Struct. Biol. 145, 3–14 (2004).
Pascual, A., Barcena, M., Merelo, J.J. & Carazo, J.M. Mapping and fuzzy classification of macromolecular images using self-organizing neural networks. Ultramicroscopy 84, 85–99 (2000).
Pascual-Montano, A., Taylor, K.A., Winkler, H., Pascual-Marqui, R.D. & Carazo, J.M. Quantitative self-organizing maps for clustering electron tomograms. J. Struct. Biol. 138, 114–122 (2002).
Scheres, S.H.W., Valle, M. & Carazo, J.M. Fast maximum-likelihood refinement of electron microscopy images. Bioinformatics 21 Suppl 2: ii243–ii244 (2005).
Sorzano, C.O.S., Marabini, R., Herman, G.T. & Carazo, J.M. Multiobjective algorithm parameter optimization using multivariate statistics in three-dimensional electron microscopy reconstruction. Pattern Recognit. 38, 2587–2601 (2005).
Van Heel, M. Similarity measures between images. Ultramicroscopy 21, 95–99 (1987).
Unser, M., Aldroubi, A. & Eden, M. The L(2) polynomial spline pyramid. IEEE Trans. Pattern Anal. Mach. Intell. 15, 364–379 (1993).
Unser, M. et al. Spectral signal-to-noise ratio and resolution assessment of 3D reconstructions. J. Struct. Biol. 149, 243–255 (2005).
Nunez-Ramirez, R. et al. Quaternary polymorphism of replicative helicase G40P: structural mapping and domain rearrangement. J. Mol. Biol. 357, 1063–1076 (2006).
Acknowledgements
We thank Haixiao Gao and Joachim Frank for providing the ribosome data, and we thank the Barcelona Supercomputing Center (Centro Nacional de Supercomputación) for providing computer resources. This work was funded by the European Union (FP6-502828 and UE-512092), the US National Institutes of Health (HL740472), the Spanish Comisión Interministerial de Ciencia y Tecnología (BFU2004-00217), the Spanish Ministerio de Educacioón y Ciencias (CSD2006-0023, BIO2007-67150-C03-01 and -03), the Spanish Fondo de Investigación Sanitaria (04/0683) and the Comunidad de Madrid (S-GEN-0166-2006).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Scheres, S., Núñez-Ramírez, R., Sorzano, C. et al. Image processing for electron microscopy single-particle analysis using XMIPP. Nat Protoc 3, 977–990 (2008). https://doi.org/10.1038/nprot.2008.62
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2008.62
This article is cited by
-
Distinct structural modulation of photosystem I and lipid environment stabilizes its tetrameric assembly
Nature Plants (2020)
-
Structure of a minimal photosystem I from the green alga Dunaliella salina
Nature Plants (2020)
-
Structural basis for cooperativity of human monoclonal antibodies to meningococcal factor H-binding protein
Communications Biology (2019)
-
Coherent diffractive imaging of microtubules using an X-ray laser
Nature Communications (2019)
-
Cryo-EM of nucleosome core particle interactions in trans
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.