Abstract
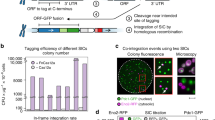
The availability of a near-complete (96%) collection of gene-deletion mutants in Saccharomyces cerevisiae greatly facilitates the systematic analyses of gene function in yeast. The unique 20 bp DNA 'barcodes' or 'tags' in each deletion strain enable the individual fitness of thousands of deletion mutants to be resolved from a single pooled culture. Here, we present protocols for the study of pooled cultures of tagged yeast deletion mutants with a tag microarray. This process involves five main steps: pooled growth, isolation of genomic DNA, PCR amplification of the barcodes, array hybridization and data analysis. Pooled deletion screening can be used to study gene function, uncover a compound's mode of action and identify drug targets. In addition to these applications, the general method of studying pooled samples with barcode arrays can also be adapted for use with other types of samples, such as mutant collections in other organisms, short interfering RNA vectors and molecular inversion probes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout











Similar content being viewed by others
References
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Winzeler, E.A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Shoemaker, D.D., Lashkari, D.A., Morris, D., Mittmann, M. & Davis, R.W. Quantitative phenotypic analysis of yeast deletion mutants using a highly parallel molecular bar-coding strategy. Nat. Genet. 14, 450–456 (1996).
Birrell, G.W. et al. Transcriptional response of Saccharomyces cerevisiae to DNA-damaging agents does not identify the genes that protect against these agents. Proc. Natl. Acad. Sci. USA 99, 8778–8783 (2002).
Deutschbauer, A.M. et al. Mechanisms of haploinsufficiency revealed by genome-wide profiling in yeast. Genetics 169, 1915–1925 (2005).
Giaever, G. et al. Chemogenomic profiling: identifying the functional interactions of small molecules in yeast. Proc. Natl. Acad. Sci. USA 101, 793–798 (2004).
Giaever, G. et al. Genomic profiling of drug sensitivities via induced haploinsufficiency. Nat. Genet. 21, 278–283 (1999).
Kastenmayer, J.P. et al. Functional genomics of genes with small open reading frames (sORFs) in S. cerevisiae. Genome Res. 16, 365–373 (2006).
Lee, W. et al. Genome-wide requirements for resistance to functionally distinct DNA-damaging agents. PLoS Genet. 1, e24 (2005).
Lum, P.Y. et al. Discovering modes of action for therapeutic compounds using a genome-wide screen of yeast heterozygotes. Cell 116, 121–137 (2004).
Ooi, S.L., Shoemaker, D.D. & Boeke, J.D. A DNA microarray-based genetic screen for nonhomologous end-joining mutants in Saccharomyces cerevisiae. Science 294, 2552–2556 (2001).
Parsons, A.B. et al. Integration of chemical-genetic and genetic interaction data links bioactive compounds to cellular target pathways. Nat. Biotechnol. 22, 62–69 (2004).
Parsons, A.B. et al. Exploring the mode-of-action of bioactive compounds by chemical-genetic profiling in yeast. Cell 126, 611–625 (2006).
Steinmetz, L.M. et al. Systematic screen for human disease genes in yeast. Nat. Genet. 31, 400–404 (2002).
Workman, C.T. et al. A systems approach to mapping DNA damage response pathways. Science 312, 1054–1059 (2006).
Jensen, L.J., Jensen, T.S., de Lichtenberg, U., Brunak, S. & Bork, P. Co-evolution of transcriptional and post-translational cell-cycle regulation. Nature 443, 594–597 (2006).
Pollack, J.R. et al. Microarray analysis reveals a major direct role of DNA copy number alteration in the transcriptional program of human breast tumors. Proc. Natl. Acad. Sci. USA 99, 12963–12968 (2002).
Groh, J.L., Luo, Q., Ballard, J.D. & Krumholz, L.R. A method adapting microarray technology for signature-tagged mutagenesis of Desulfovibrio desulfuricans G20 and Shewanella oneidensis MR-1 in anaerobic sediment survival experiments. Appl. Environ. Microbiol. 71, 7064–7074 (2005).
Karlyshev, A.V. et al. Application of high-density array-based signature-tagged mutagenesis to discover novel Yersinia virulence-associated genes. Infect. Immun. 69, 7810–7819 (2001).
Berns, K. et al. A large-scale RNAi screen in human cells identifies new components of the p53 pathway. Nature 428, 431–437 (2004).
Brummelkamp, T.R. et al. An shRNA barcode screen provides insight into cancer cell vulnerability to MDM2 inhibitors. Nat. Chem. Biol. 2, 202–206 (2006).
Fischer, K.D. et al. Defective T-cell receptor signalling and positive selection of Vav-deficient CD4+ CD8+ thymocytes. Nature 374, 474–474 (1995).
Fraser, A. RNA interference: human genes hit the big screen. Nature 428, 375–375 (2004).
Kolfschoten, I.G. et al. A genetic screen identifies PITX1 as a suppressor of RAS activity and tumorigenicity. Cell 121, 849–858 (2005).
Ngo, V.N. et al. A loss-of-function RNA interference screen for molecular targets in cancer. Nature 441, 106–110 (2006).
Westbrook, T.F. et al. A genetic screen for candidate tumor suppressors identifies REST. Cell 121, 837–848 (2005).
Akhras, M.S. et al. PathogenMip assay: a multiplex pathogen detection assay. PLoS ONE 2, e223 (2007).
Clayton, D.G. et al. Population structure, differential bias and genomic control in a large-scale, case-control association study. Nat. Genet. 37, 1243–1246 (2005).
Hardenbol, P. et al. Multiplexed genotyping with sequence-tagged molecular inversion probes. Nat. Biotechnol. 21, 673–678 (2003).
Hardenbol, P. et al. Highly multiplexed molecular inversion probe genotyping: over 10,000 targeted SNPs genotyped in a single tube assay. Genome Res. 15, 269–275 (2005).
Ooi, S.L., Shoemaker, D.D. & Boeke, J.D. A DNA microarray-based genetic screen for nonhomologous end-joining mutants in Saccharomyces cerevisiae. Science 294, 2552–2556 (2001).
Pan, X. et al. A robust toolkit for functional profiling of the yeast genome. Mol. Cell. 16, 487–496 (2004).
Yuan, D.S. et al. Improved microarray methods for profiling the Yeast Knockout strain collection. Nucleic Acids Res. 33, e103 (2005).
Bolstad, B.M., Irizarry, R.A., Astrand, M. & Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Pierce, S.E. et al. A unique and universal molecular barcode array. Nat. Methods 3, 601–603 (2006).
Tusher, V.G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Tong, A.H. et al. Global mapping of the yeast genetic interaction network. Science 303, 808–813 (2004).
Davierwala, A.P. et al. The synthetic genetic interaction spectrum of essential genes. Nat. Genet. 37, 1147–1152 (2005).
Schuldiner, M. et al. Exploration of the function and organization of the yeast early secretory pathway through an epistatic miniarray profile. Cell 123, 507–519 (2005).
Stark, C. et al. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 34, D535–D539 (2006).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Data 1
Matlab function for modeling samling error (PDF 10 kb)
Supplementary Data 2
Barcode sequences on the Affymetrix TAG4 array (PDF 1105 kb)
Rights and permissions
About this article
Cite this article
Pierce, S., Davis, R., Nislow, C. et al. Genome-wide analysis of barcoded Saccharomyces cerevisiae gene-deletion mutants in pooled cultures. Nat Protoc 2, 2958–2974 (2007). https://doi.org/10.1038/nprot.2007.427
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.427
This article is cited by
-
The cellular response to drug perturbation is limited: comparison of large-scale chemogenomic fitness signatures
BMC Genomics (2022)
-
Identification of novel genes involved in neutral lipid storage by quantitative trait loci analysis of Saccharomyces cerevisiae
BMC Genomics (2021)
-
Inactivating histone deacetylase HDA promotes longevity by mobilizing trehalose metabolism
Nature Communications (2021)
-
A genome-wide portrait of pervasive drug contaminants
Scientific Reports (2021)
-
Jawsamycin exhibits in vivo antifungal properties by inhibiting Spt14/Gpi3-mediated biosynthesis of glycosylphosphatidylinositol
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.