Abstract
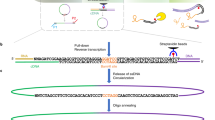
Understanding gene regulatory networks in mammalian cells requires detailed knowledge of protein–DNA interactions. Commonly used methods for genome-wide mapping of these interactions are based on chromatin immunoprecipitation. However, these methods have some drawbacks, such as the use of crosslinking reagents, the need for highly specific antibodies and relatively large amounts of starting material. We present DamID, an alternative technique to map genome-wide occupancy of interaction sites in vivo, that bypasses these limitations. DamID is based on the expression of a fusion protein consisting of a protein of interest and DNA adenine methyltransferase (Dam). This leads to methylation of adenines near sites where the protein of interest interacts with the DNA. These methylated sequences are subsequently amplified by a methylation-specific PCR protocol and identified by hybridization to microarrays. Using DamID, genome-wide maps of the binding of DNA-interacting proteins in mammalian cells can be constructed efficiently. Depending on the strategy used for expression of the Dam-fusion proteins, genome-wide binding maps can be obtained in as little as 2 weeks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
06 September 2007
The version of this article originally published indicated that the authors had no competing financial interests. The Competing Interests Statement should have read: “B.V.S. is listed as an inventor on a patent application for the DamID technology.” The error has been corrected in the HTML and PDF versions of the article.
References
Oberley, M.J., Tsao, J., Yau, P. & Farnham, P.J. High-throughput screening of chromatin immunoprecipitates using CpG-island microarrays. Methods Enzymol. 376, 315–334 (2004).
Lee, T.I., Johnstone, S.E. & Young, R.A. Chromatin immunoprecipitation and microarray-based analysis of protein location. Nat. Protocols 1, 729–748 (2006).
Ren, B. & Dynlacht, B.D. Use of chromatin immunoprecipitation assays in genome-wide location analysis of mammalian transcription factors. Methods Enzymol. 376, 304–315 (2004).
Nelson, J.D., Denisenko, O., Sova, P. & Bomsztyk, K. Fast chromatin immunoprecipitation assay. Nucleic Acids Res. 34, e2 (2006).
Wei, C.L. et al. A global map of p53 transcription-factor binding sites in the human genome. Cell 124, 207–219 (2006).
van Steensel, B. & Henikoff, S. Identification of in vivo DNA targets of chromatin proteins using tethered dam methyltransferase. Nat. Biotechnol. 18, 424–428 (2000).
van Steensel, B., Delrow, J. & Henikoff, S. Chromatin profiling using targeted DNA adenine methyltransferase. Nat. Genet. 27, 304–308 (2001).
Vogel, M.J. et al. Human heterochromatin proteins form large domains containing KRAB-ZNF genes. Genome Res. 16, 1493–1504 (2006).
Greil, F., Moorman, C. & van Steensel, B. DamID: mapping of in vivo protein–genome interactions using tethered DNA adenine methyltransferase. Methods Enzymol. 410, 342–359 (2006).
Orian, A. Chromatin profiling, DamID and the emerging landscape of gene expression. Curr. Opin. Genet. Dev. 16, 157–164 (2006).
Moorman, C. et al. Hotspots of transcription factor colocalization in the genome of Drosophila melanogaster . Proc. Natl. Acad. Sci. USA 103, 12027–12032 (2006).
Pickersgill, H. et al. Characterization of the Drosophila melanogaster genome at the nuclear lamina. Nat. Genet. 38, 1005–1014 (2006).
Tolhuis, B. et al. Genome-wide profiling of PRC1 and PRC2 Polycomb chromatin binding in Drosophila melanogaster . Nat. Genet. 38, 694–699 (2006).
Germann, S., Juul-Jensen, T., Letarnec, B. & Gaudin, V. DamID, a new tool for studying plant chromatin profiling in vivo, and its use to identify putative LHP1 target loci. Plant J. 48, 153–163 (2006).
Sun, L.V. et al. Protein–DNA interaction mapping using genomic tiling path microarrays in Drosophila . Proc. Natl. Acad. Sci. USA 100, 9428–9433 (2003).
Buck, M.J. & Lieb, J.D. ChIP-chip: considerations for the design, analysis, and application of genome-wide chromatin immunoprecipitation experiments. Genomics 83, 349–360 (2004).
Boyer, L.A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Negre, N. et al. Chromosomal distribution of PcG proteins during Drosophila development. PLoS Biol. 4, e170 (2006).
Tiscornia, G., Singer, O. & Verma, I.M. Production and purification of lentiviral vectors. Nat. Protocols 1, 241–245 (2006).
Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004).
Huber, W., von Heydebreck, A., Sultmann, H., Poustka, A. & Vingron, M. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 18 (suppl. 1): S96–S104 (2002).
Kreil, D.P. & Russell, R.R. There is no silver bullet—a guide to low-level data transforms and normalisation methods for microarray data. Brief Bioinform. 6, 86–97 (2005).
Wit, E. & McClure, J. Statistics for Microarrays: Design, Analysis, and Inference (John Wiley & Sons Ltd, Chichester, England, 2004).
Acknowledgements
We thank Lars Guelen for development of DamID using lentiviruses and critical reading of the manuscript; Martin Lodén, Richard Heideman and Anja Duursma for useful suggestions and sharing results; and members of our laboratory for help with the development of DamID in mammalian cells. We thank the staff of the NKI Central Microarray Facility for extensive help with protocol development and technical support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
B.V.S. is listed as an inventor on a patent application for the DamID technology.
Rights and permissions
About this article
Cite this article
Vogel, M., Peric-Hupkes, D. & van Steensel, B. Detection of in vivo protein–DNA interactions using DamID in mammalian cells. Nat Protoc 2, 1467–1478 (2007). https://doi.org/10.1038/nprot.2007.148
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.148
This article is cited by
-
Computational methods for analysing multiscale 3D genome organization
Nature Reviews Genetics (2024)
-
Mena regulates nesprin-2 to control actin–nuclear lamina associations, trans-nuclear membrane signalling and gene expression
Nature Communications (2023)
-
B1 SINE-binding ZFP266 impedes mouse iPSC generation through suppression of chromatin opening mediated by reprogramming factors
Nature Communications (2023)
-
Nuclear architecture and the structural basis of mitotic memory
Chromosome Research (2023)
-
Comprehensive assessment of differential ChIP-seq tools guides optimal algorithm selection
Genome Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.