Abstract
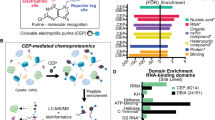
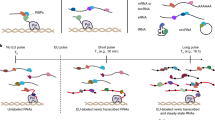
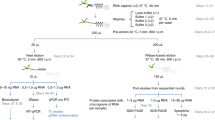
All aspects of RNA metabolism are regulated by RNA-binding proteins (RBPs) that directly associate with the RNA. Some aspects of RNA biology such as RNA abundance can be readily assessed using standard hybridization technologies. However, identification of RBPs that specifically associate with selected RNAs has been more difficult, particularly when attempting to assess this in live cells. The peptide nucleic acid (PNA)-assisted identification of RBPs (PAIR) technology has recently been developed to overcome this issue. The PAIR technology uses a cell membrane–penetrating peptide (CPP) to efficiently deliver into the cell its linked PNA oligomer that complements the target mRNA sequence. The PNA will then anneal to its target mRNA in the living cell, and then covalently couple to the mRNA-RBP complexes subsequent to an ultraviolet (UV) cross-linking step. The resulting PNA-RNA-RBP complex can be isolated using sense oligonucleotide magnetic beads, and the RBPs can then be identified by mass spectrometry (MS). This procedure can usually be completed within 3 d. The use of the PAIR procedure promises to provide insight into the dynamics of RNA processing, transport, degradation and translation in live cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Wagner, R.W., Smith, J.E., Cooperman, B.S. & Nishikura, K. A double-stranded RNA unwinding activity introduces structural alterations by means of adenosine to inosine conversions in mammalian cells and Xenopus eggs. Proc. Natl. Acad. Sci. USA 86, 2647–2651 (1989).
Gingras, A.C., Raught, B. & Sonenberg, N. eIF4 initiation factors: effectors of mRNA recruitment to ribosomes and regulators of translation. Annu. Rev. Biochem. 68, 913–963 (1999).
Kiebler, M.A. et al. The mammalian staufen protein localizes to the somatodendritic domain of cultured hippocampal neurons: implications for its involvement in mRNA transport. J. Neurosci. 19, 288–297 (1999).
Gebauer, F. & Hentze, M.W. Molecular mechanisms of translational control. Nat. Rev. Mol. Cell Biol. 5, 827–835 (2004).
Ule, J. & Darnell, R.B. RNA binding proteins and the regulation of neuronal synaptic plasticity. Curr. Opin. Neurobiol. 16, 102–110 (2006).
Blencowe, B.J., Bowman, J.A., McCracken, S. & Rosonina, E. SR-related proteins and the processing of messenger RNA precursors. Biochem. Cell Biol. 77, 277–291 (1999).
Tenenbaum, S.A., Carson, C.C., Lager, P.J. & Keene, J.D. Identifying mRNA subsets in messenger ribonucleoprotein complexes by using cDNA arrays. Proc. Natl. Acad. Sci. USA 97, 14085–14090 (2000).
Brown, V. et al. Microarray identification of FMRP-associated brain mRNAs and altered mRNA translational profiles in fragile X syndrome. Cell 107, 477–487 (2001).
Ule, J. et al. CLIP identifies Nova-regulated RNA networks in the brain. Science 302, 1212–1215 (2003).
Peritz, T. et al. Immunoprecipitation of mRNA-protein complexes. Nat. Protocols doi: 10.1038/nprot.2006.82 (2006).
Zielinski, J. et al. In vivo identification of ribonucleoprotein-RNA interactions. Proc. Natl. Acad. Sci. USA 103, 1557–1562 (2006).
Pooga, M. et al. Cell penetrating PNA constructs regulate galanin receptor levels and modify pain transmission in vivo. Nat. Biotechnol. 16, 857–861 (1998).
Nielsen, P.E., Egholm, M., Berg, R.H. & Buchardt, O. Sequence-selective recognition of DNA by strand displacement with a thymine-substituted polyamide. Science 254, 1497–1500 (1991).
Xi, C., Balberg, M., Boppart, S.A. & Raskin, L. Use of DNA and peptide nucleic acid molecular beacons for detection and quantification of rRNA in solution and in whole cells. Appl. Environ. Microbiol. 69, 5673–5678 (2003).
Pooga, M. & Langel, Ü. Synthesis of cell-penetrating peptides for cargo delivery. in Peptide Synthesis and Applications (ed. Howl, J.) 77–89 (Humana Press, Totowa, NJ, 2005).
Soomets, U. et al. Deletion analogues of transportan. Biochim. Biophys. Acta 1467, 165–176 (2000).
Wessel, D. & Flugge, U.I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal. Biochem. 138, 141–143 (1984).
Acknowledgements
We thank Jennifer Zielinski for helping to pioneer this technology. We thank Margie Maronski for help with culturing the neurons, and Dr. Chao Xing Yuan and Elena Blagoi for help with mass spectrometry. This work was funded by National Institutes of Health grants AG9900 and MH58561 to J.E.; and the Swedish Science Foundation Grants Med and NT, and European Community Grant QLRT-2001-01989 to U.L.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
J.E., K.K. and U.L. are co-inventors on the Pair Technology, and a patent has been submitted.
Supplementary information
Supplementary Table 1
PAIR protocol checklist. (DOC 86 kb)
Rights and permissions
About this article
Cite this article
Zeng, F., Peritz, T., Kannanayakal, T. et al. A protocol for PAIR: PNA-assisted identification of RNA binding proteins in living cells. Nat Protoc 1, 920–927 (2006). https://doi.org/10.1038/nprot.2006.81
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.81
This article is cited by
-
Methods to study RNA–protein interactions
Nature Methods (2019)
-
Transcriptome in vivo analysis (TIVA) of spatially defined single cells in live tissue
Nature Methods (2014)
-
Modulation of dADAR-dependent RNA editing by the Drosophila fragile X mental retardation protein
Nature Neuroscience (2011)
-
Optimization of immunoprecipitation–western blot analysis in detecting GW182-associated components of GW/P bodies
Nature Protocols (2009)
-
Immunoprecipitation of mRNA-protein complexes
Nature Protocols (2006)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.