Abstract
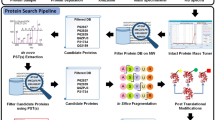
There is growing interest in the overall study of post-translational modifications (PTMs) of proteins. Beside phosphorylation and glycosylation, truncations of the nascent polypeptide chain at the N or C termini are by far the most common types of PTMs found in proteins. However, little attention has been paid to the development of approaches that allow a systematic analysis of these proteolytic processing events. Here we present a protocol that allows the identification of the C-terminal sequences of proteins. A peptide mixture is generated by cleavage of the protein with cyanogen bromide and is incubated with carboxypeptidase Y. The enzyme is only able to act on the C-terminal fragment, because this is the only peptide without a homoserine lactone residue at its C terminus. The resulting fragments, forming a peptide ladder, are analyzed by matrix-assisted laser desorption/ionization mass spectrometry (MALDI-MS). The entire protocol, including the CNBr cleavage, takes 21 h and can be applied to proteins purified either by SDS-PAGE or by 2D PAGE or in solution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Boyd, V., Bozzini, M., Zon, G., Noble, R.L. & Mattaliano, R.J. Sequencing of peptides and proteins from the carboxy terminus. Anal. Biochem. 206, 344–352 (1992).
Bailey, J.M. & Shively, J.E. A chemical method for the C-terminal sequence analysis of proteins. Methods Companion Methods Enzymol. 6, 334–350 (1994).
Samyn, B., Hardeman, K., Van der Eycken, J. & Van Beeumen, J. Applicability of the alkylation chemistry for chemical C-terminal protein sequence analysis. Anal. Chem. 72, 1389–1399 (2000).
Hardeman, K., Samyn, B., Van der Eycken, J. & Van Beeumen, J. An improved chemical approach toward the C-terminal sequence analysis of proteins containing all natural amino acids. Protein Sci. 7, 1593–1602 (1998).
Sechi, S. & Chait, B.T. A method to define the carboxyl terminal of proteins. Anal. Chem. 72, 3374–3378 (2000).
Kosaka, T., Takazawa, T. & Nakamura, T. Identification and C-terminal characterization of proteins from two-dimensional polyacrylamide gels by a combination of isotopic labeling and nano-electrospray Fourier transform ion cyclotron resonance mass spectrometry. Anal. Chem. 72, 1179–1185 (2000).
Zhou, X.W., Blackman, M.J., Howell, S.A. & Carruthers, V.B. Proteomic analysis of cleavage events reveals a dynamic two-step mechanism for proteolysis of a key parasite adhesive complex. Mol. Cell. Proteomics 3, 565–576 (2004).
Chait, B.T., Wang, R., Beavis, R.C. & Kent, S.B.H. Protein ladder sequencing. Science 262, 89–92 (1993).
Samyn, B., Sergeant, K., Castanheira, P., Faro, C. & Van Beeumen, J. A novel method for C-terminal sequence analysis in the proteomic era. Nat. Methods 2, 193–200 (2005).
Gross, E. & Witkop, B. Nonenzymatic cleavage of peptide bonds: the methionine residues in bovine pancreatic ribonuclease. J. Biol. Chem. 237, 1856–1860 (1962).
Gross, E. The cyanogen bromide reaction. Methods Enzymol. 11, 27.238–27.255 (1967).
Wilkins, M.R et al. Protein identification with N- and C-terminal sequence tags in proteome projects. J. Mol. Biol. 278, 599–608 (1998).
Chung, J.-J., Yang, H. & Li, M. Genome-wide analysis of carboxyl-terminal sequences. Mol. Cell. Proteomics 2, 173–181 (2003).
Chung, J.-J., Shikano, S., Hanyu, Y. & Li, M. Functional diversity of protein C-termini: more than zipcoding? Trends Cell. Biol. 12, 146–150 (2002).
Castanheira, P. et al. Activation, proteolytic processing and peptide specificity of recombinant cardosin A. J. Biol. Chem. 280, 13047–13054 (2005).
Patterson, D.H., Tarr, G.E., Regnier, F.E. & Martin, S.A. C-terminal ladder sequencing via matrix-assisted laser desorption mass spectrometry coupled with carboxypeptidase Y time dependent and concentration-dependent digestions. Anal. Chem. 67, 3971–3978 (1995).
Ambler, R.P. Enzymatic hydrolysis with carboxypeptidases. Methods Enzymol. 11, 14.155–14.166 (1967).
Jones, J.J. et al. Investigation of MALDI-TOF and FT-MS techniques for analysis of Escherichia coli whole cells. Anal. Chem. 75, 1340–1347 (2003).
Quadroni, M. & James, P. Proteomics and automation. Electrophoresis 20, 664–677 (1999).
Caputo, E., Moharram, R. & Martin, B.M. Methods for on-chip protein analysis. Anal. Biochem. 321, 116–124 (2003).
Acknowledgements
B.S. is a Postdoctoral fellow of the Fund for Scientific Research–Flanders (FWO.-Vlaanderen), Belgium. K.S. is funded by a PhD grant of the Institute for the Promotion of Innovation through Science and Technology in Flanders (IWT-Vlaanderen).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
C-terminal sequence analysis of the test peptide ACTH 18–39 Human. (PDF 4149 kb)
Supplementary Fig. 2
C-terminal sequence analysis of cytochrome c (pigeon) in solution. (PDF 5100 kb)
Rights and permissions
About this article
Cite this article
Samyn, B., Sergeant, K. & Van Beeumen, J. A method for C-terminal sequence analysis in the proteomic era (proteins cleaved with cyanogen bromide). Nat Protoc 1, 318–323 (2006). https://doi.org/10.1038/nprot.2006.50
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.50
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.