Abstract
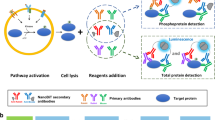
This protocol describes a 'mix-and-measure' procedure for the analysis of interactions of endogenous proteins in microliters of crude cell lysates. The proteins of interest are labeled by indirect immunofluorescence through simple addition of all primary and secondary antibodies to the lysate. Detection is based on fluorescence cross-correlation spectroscopy. Due to the minimal number of handling steps for sample preparation and the need of only microliters of sample, the approach enables the parallel and miniaturized analysis of protein-protein interactions. No heterologous expression of proteins with detection tags is required. For this reason, the cellular processes leading to protein-protein interactions are not skewed by overexpression of individual components. This makes the approach particularly suitable for the parallel monitoring of interactions in signaling networks. Additionally, the approach enables the screening and titration of compounds interfering with interactions, especially for those interactions based on signaling-dependent post-translational modifications. This protocol can be completed in approximately 22 h, including a 16-h incubation phase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Stoevesandt, O., Köhler, K., Fischer, R., Johnston, I.C. & Brock, R. One-step analysis of protein complexes in microliters of cell lysate. Nat. Methods 2, 833–835 (2005).
Schwille, P., Meyer-Almes, F.J. & Rigler, R. Dual-color fluorescence cross-correlation spectroscopy for multicomponent diffusional analysis in solution. Biophys. J. 72, 1878–1886 (1997).
Bacia, K., Kim, S.A. & Schwille, P. Fluorescence cross-correlation spectroscopy in living cells. Nat. Methods 3, 83–89 (2006).
Clegg, R.M. Fluorescence resonance energy transfer. Curr. Opin. Biotechnol. 6, 103–110 (1995).
Schoeberl, B., Eichler-Jonsson, C., Gilles, E.D. & Muller, G. Computational modeling of the dynamics of the MAP kinase cascade activated by surface and internalized EGF receptors. Nat. Biotechnol. 20, 370–375 (2002).
Stoevesandt, O. et al. Peptide microarrays for the detection of molecular interactions in cellular signal transduction. Proteomics 5, 2010–2017 (2005).
Gaide, O. et al. Carma1 is a critical lipid raft-associated regulator of TCR-induced NF-kappa B activation. Nat. Immunol. 3, 836–843 (2002).
Jahnz, M. & Schwille, P. An ultrasensitive site-specific DNA recombination assay based on dual-color fluorescence cross-correlation spectroscopy. Nucleic Acids Res. 33, e60 (2005).
Acknowledgements
The authors thank N. Fischer for practical help with the experiments. R.B. gratefully acknowledges financial support from the Volkswagen Foundation ('Nachwuchsgruppen an Universitäten' I/77 472).
Author information
Authors and Affiliations
Contributions
O.S. developed and validated the protocol, performed data analysis, and wrote the manuscript; R.B. developed the protocol and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Stoevesandt, O., Brock, R. One-step analysis of protein complexes in microliters of cell lysate using indirect immunolabeling & fluorescence cross-correlation spectroscopy. Nat Protoc 1, 223–229 (2006). https://doi.org/10.1038/nprot.2006.34
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.34
This article is cited by
-
Practical guidelines for dual-color fluorescence cross-correlation spectroscopy
Nature Protocols (2007)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.