Abstract
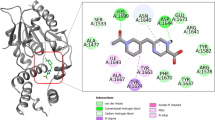
Tuberculosis causes more than two million deaths per year. Faced with this global threat it is crucial to better understand the physiology of the causative organism, Mycobacterium Tuberculosis, in order to develop efficient therapeutic strategies. PKnB from Mycobacterium tuberculosis is a crucial receptor-like protein kinase involved in signal transduction. M. tuberculosis PKnB is a trans-membrane Ser/Thr protein kinase (STPK) highly conserved in Gram-positive bacteria and apparently essential for Mycobacterial viability. We have attempted with the help of virtual screening and docking approaches to expound the extent of specificity of protein kinase B towards different classes of Thiadiazoles (an anti-tubercular agent). The selected Thiadiazoles were selected on the basis of the structural specificity to the enzyme towards its substrate and inhibitors. Total number of Thiadiazoles were 5000 in number with the minimum binding energy of -10.46 kcal/mol with 10 molecules showing hydrogen bonds with the active site residue. The protein kinase B peptide contains two types of structural elements (Valine 95, Arginine 97) and the basic residue ring consisted of glycine rich residues. The structure of the protein-ligand complex reveals that Thiadiazoles partially occupies the adenine-binding pocket in PKnB, providing a framework for the design of compounds with potential therapeutic applications. The study provides hints for the future design of new derivatives with higher potency and specificity.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Gupta, V., Naqvi, A., Tanwar, J. et al. Molecular Modeling study of Protein Kinase PKnB from Mycobacterium Tuberculosis with derivatives of 1, 3, 4- Thiadiazoles. Nat Prec (2011). https://doi.org/10.1038/npre.2011.6136.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2011.6136.1