Abstract
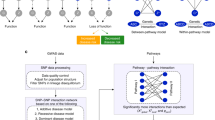
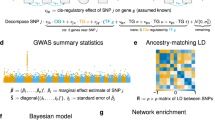
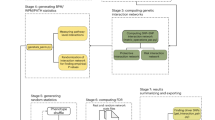
After completion of a number of large scale Genome-Wide Association Studies (GWAS), there is still a significant amount of trait and disease variance that cannot be explained by existing genetic variability. This review introduces new, Integrative Network-based Association Study (INAS) approaches that aim to minimize the impact from multiple hypothesis testing statistics, thus allowing the identification of rare variants/alterations and epistatic interactions. In particular we discuss methods that rely on the de novo computational, experimental, and integrative dissection of context specific molecular interaction networks (or interactomes, for short). We provide several examples of how these approaches may be used to tackle discovery of genetic variants and somatic alterations causally related to the presentation of specific traits and diseases. We also discuss how more complex systems, including a variety of non-cell-autonomous traits and diseases will require new multicellular networks that explicitly represent short distance paracrine and long distance endocrine interactions.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Califano, A., Butte, A., Friend, S. et al. Integrative Network-based Association Studies: Leveraging cell regulatory models in the post-GWAS era . Nat Prec (2011). https://doi.org/10.1038/npre.2011.5732.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2011.5732.1