Abstract
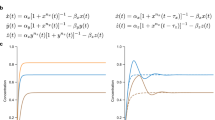
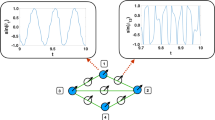
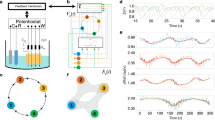
The mammalian cell can be represented as a large modular network that consists of a central signal network that interacts with and regulates multiple cellular machines that are responsible for phenotypic behavior. We have used graph-theory approaches to analyze signal flow through a network representing the hippocampal neuron and find that signal-induced connectivity results in the formation of many regulatory motifs. Information flow through the central signaling network is initiated by extra-cellular signals such as hormones binding to their receptors. The flow of information through the signaling network results in the appearance of regulatory motifs such as feedback loops, feedforward and bifan motifs. Within the large cellular networks, these regulatory motifs are juxtaposed next to each other in several formats such as the stacked configuration or the nested configuration. We have studied the dynamics of regulatory motifs by biochemical computation using ordinary differential equation models. Positive feedback loops can function as bistable switches. Nested feed-forward motifs can give rise to two emergent properties: coincidence detection and prolonged outputs for short inputs. Bifan motifs can control response times, with some configurations working as delays and others promoting rapid responses. Bifan motifs can also act as filters. Feed-forward motifs lead to signal prolongation and thus function as a switch to alter cell state. The functional consequences of organization of motifs within networks as well as the properties of feedback, feedforward and bifans motifs are presented.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Iyengar, R. Dynamic Topology of Biological Networks. Nat Prec (2007). https://doi.org/10.1038/npre.2007.24.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2007.24.1