Abstract
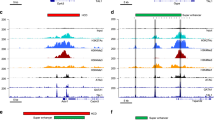
ChIP-on-chip has emerged as a powerful tool to dissect the complex network of regulatory interactions between transcription factors and their targets. However, most ChIP-on-chip analysis methods use conservative approaches aimed to minimize false-positive transcription factor targets. We present a model with improved sensitivity in detecting binding events from ChIP-on-chip data. Biochemically validated analysis in human T-cells reveals that three transcription factor oncogenes, NOTCH1, MYC, and HES1, bind one order of magnitude more promoters than previously thought. Gene expression profiling upon NOTCH1 inhibition shows broad-scale functional regulation across the entire range of predicted target genes, establishing a closer link between occupancy and regulation. Finally, the resolution of a more complete map of transcriptional targets reveals that MYC binds nearly all promoters bound by NOTCH1. Overall, these results suggest an unappreciated complexity of transcriptional regulatory networks and highlight the fundamental importance of genome-scale analysis to represent transcriptional programs.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Margolin, A., Palomero, T., Sumazin, P. et al. ChIP-on-chip significance analysis reveals large-scale binding and regulation by human transcription factor oncogenes. Nat Prec (2007). https://doi.org/10.1038/npre.2007.1364.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2007.1364.1