Abstract
DNA methylation profiles of the serotonin transporter gene (SLC6A4) have been shown to alter SLC6A4 expression, drive antidepressant treatment response and modify brain functions. This study investigated whether methylation of an AluJb element in the SLC6A4 promotor was associated with major depressive disorder (MDD), amygdala reactivity to emotional faces, 5-HTTLPR/rs25531 polymorphism, and recent stress. MDD patients (n=122) and healthy controls (HC, n=176) underwent fMRI during an emotional face-matching task. Individual SLC6A4 AluJb methylation profiles were ascertained and associated with MDD, amygdala reactivity, 5-HTTLPR/rs25531, and stress. SLC6A4 AluJb methylation was significantly lower in MDD compared to HC and in stressed compared to less stressed participants. Lower AluJb methylation was particularly found in 5-HTTLPR/rs25531 risk allele carriers under stress and correlated with less depressive episodes. fMRI analysis revealed a significant interaction of AluJb methylation and diagnosis in the amygdala, with MDD patients showing lower AluJb methylation associated with decreased amygdala reactivity. While no joint effect of AluJb methylation and 5-HTTLPR/rs25531 existed, risk allele carriers showed significantly increased bilateral amygdala activation. These findings suggest a role of SLC6A4 AluJb methylation in MDD, amygdala reactivity, and stress reaction, partly interwoven with 5-HTTLPR/rs25531 effects. Patients with low methylation in conjunction with a shorter MDD history and decreased amygdala reactivity might feature a more stress-adaptive epigenetic process, maybe via theoretically possible endogenous antidepressant-like effects. In contrast, patients with higher methylation might possibly suffer from impaired epigenetic adaption to chronic stress. Further, the 5-HTTLPR/rs25531 association with amygdala activation was confirmed in our large sample.
Similar content being viewed by others
Introduction
Major depressive disorder (MDD) is one of the most common mental illnesses worldwide with estimations about 350 million people affected (Marcus et al, 2012). Whereas antidepressant medication with selective serotonin reuptake inhibitors (SSRIs) is among the most prevalent treatment forms (Olfson and Marcus, 2009), the role of the serotonergic system and its interactions with environmental factors has not been conclusively determined. The main target of SSRIs, the serotonin transporter (5-HTT, SERT), broadly influences serotonergic neurotransmission and is encoded by the SLC6A4 gene. In a polymorphic SLC6A4 promoter region, the functional insertion/deletion polymorphism 5-HTTLPR and the functionally related single nucleotide polymorphism rs25531 are discussed as risk factors for the psychopathology of MDD (Xia and Yao, 2015) and frequently associated with robust neural correlates of MDD, like reduced hippocampal volumes (eg, Frodl et al, 2008). Starting with the seminal study of Hariri and colleagues (Hariri et al, 2002), numerous imaging genetic studies also investigated the association of the 5-HTTLPR region and amygdala activity, with several replications (eg, Dannlowski et al, 2010) but, however, also controversially discussed results (cf. Bastiaansen et al, 2015; Kaufman, 2015). This underlines the importance of an expanded perspective and more complex study approaches including also epigenetics (Kaufman, 2015).
Epigenetic modifications like DNA methylation impact gene transcription in response to environmental influences, revealing a link between nature (genetics) and nurture (life experience) that shapes physical responses to, eg, stressful stimuli (Zannas and West, 2014). Alexander et al have recently shown an association between the 5-HTTLPR genotype, SLC6A4 DNA methylation, and stress sensitivity, where S allele carriers in combination with low SLC6A4 methylation revealed increased cortisol stress reactivity (eg, Alexander et al, 2014; Frodl et al, 2015). Further research suggests that in addition to the genetic makeup, epigenetic mechanisms play a major role in the susceptibility to acute, chronic, and developmental stressors. Wilkinson et al (2009) showed different neuronal methylation patterns in stress-vulnerable compared to resilient mice at several genes, offering the hypothesis that resilient individuals reveal an antidepressant-like epigenetic response to chronic stress. Recent studies on DNA methylation in the SLC6A4 promoter revealed associations with acute stress (Kang et al, 2013), 5-HTT mRNA levels (Olsson et al, 2010; Philibert et al, 2008), history of lifetime depression (Philibert et al, 2008), depression severity (Okada et al, 2014), childhood maltreatment (Frodl et al, 2015; Kang et al, 2013), and antidepressant response (Domschke et al, 2014). In addition, a large imaging epigenetic study linked amygdala hyperactivity with increased SLC6A4 promoter methylation in healthy participants (Nikolova et al, 2014). Given the heterogeneous susceptibility to stress-related disorders in humans (Galea et al, 2005), improved understanding of how epigenetically modified genes are associated with adaptive stress responses, psychiatric disorders, and neurobiological changes could have tremendous implications for therapeutic interventions.
Recently, another functional region in the SLC6A4 promoter awakened interest, a retrotransposonal Alu element of subtype AluJb. The primate specific Alus are ~300 bp in length, genome-wide prevalent (occupying 10% of the human genome) and function either as genomic elements or as transcribed RNA (Deininger, 2011; Wang and Huang, 2014). As genomic elements, Alus contain several transcription-factor-binding sites and preferentially interact with nearby promoters (Su et al, 2014). Further, Alus can regulate through editing and exonization the diversity, stability and translatability of mRNA when embedded in pre/mRNA. Finally, free transcribed Alu ncRNAs in sense orientation have been shown to regulate gene expression via blocking RNA pol II, while free transcribed Alu ncRNA in reverse orientation have been shown to act through antisense RNA or small RNA interference mechanisms (Wang and Huang, 2014). Alus have great potential to regulate nearby gene expression (cf. Kaer and Speek, 2013) and their methylation status was already associated with pathology of numerous diseases and shown to be regulated by environmental factors (Miousse et al, 2015). For the AluJb in the SLC6A4 promoter, located between 5-HTTLPR/rs25531 and the promoter associated CpG island (Supplementary Figure S1), we previously reported a striking relationship between its methylation status and hippocampus and amygdala volumes in two independent and large samples of healthy participants (Dannlowski et al, 2014).
Based on these promising findings the question arises, whether a similar relation might also exist between AluJb methylation and amygdala reactivity in MDD patients. Further, we were interested in associations between AluJb methylation and gene (5-HTTLPR region) and environment (stressors) effects since studies have shown associations between SLC6A4 DNA methylation, diagnosis of major depression, the 5-HTTLPR genotype, stress sensitivity as well as an interaction between 5-HTTLPR genotype and stress sensitivity predicting SLC6A4 DNA methylation (eg, Alexander et al, 2014; Frodl et al, 2015; Iga et al, 2016). Thus, for the present study we hypothesized that (1) the diagnosis of MDD, 5-HTTLPR/rs25531, stressful life events, and a gene × environment interaction (5-HTTLPR/rs25531 × stressful life events) would predict SLC6A4 promoter AluJb methylation. Stressful life events were exploratory operationalized using the List of Threatening Experiences Questionnaire (LTE-Q,(Brugha and Cragg, 1990)). Further, we hypothesized that (2) amygdala reactivity to negative emotional faces was associated with the diagnosis of MDD and SLC6A4 promoter AluJb methylation, and that (3) a joint effect of AluJb methylation and 5-HTTLPR/rs25531 on amygdala reactivity might exist.
Materials and methods
Subjects
A total of n=137 individuals with acute MDD, for whom fMRI data were available, participated in the present study as part of an ongoing project (Münster Neuroimaging Cohort). Patients were under current in-patient treatment at the Department of Psychiatry and Psychotherapy at the University Hospital of Münster with a large majority taking antidepressants (see Supplementary Table S1 for details). All patients were diagnosed with the Structured Clinical Interview for DSM-IV Axis I Disorders (SCID-I; (Wittchen et al, 1997)). Severity of depressive symptoms was measured with the Beck Depression Inventory (BDI, (Beck, 1987)) and the occurrence of 12 different stressful life events during the last year with the LTE-Q (Brugha and Cragg, 1990). Exclusion criteria were any neurological abnormalities, history of seizures, head trauma or unconsciousness, substance abuse, and the usual MRI-contraindications. In addition, patients with former electroconvulsive therapy and acute benzodiazepine treatment were excluded. A medication load index (MLI) was calculated as described previously (Hassel et al, 2008; Redlich et al, 2014) with medications grouped into absent, low, or high, and a composite measure of total medication reflecting dose and variety of different medications taken (for details see Supplementary Material).
The healthy control (HC) sample of n=189 individuals was initially recruited for our preceding study and described there in detail (Dannlowski et al, 2014). Briefly, they were free from any life-time history of psychiatric disorders according to DSM-IV criteria (American Psychiatric Association, 1994), as diagnosed with the SCID interview (Wittchen et al, 1997) and were characterized with the same questionnaires and followed the same exclusion criteria as abovementioned for the patients (for details and a comparison of MDD vs HC see also Table 1).
All subjects were of European ancestry and had normal or corrected-to-normal vision. Due to excess motion in the fMRI session, 28 participants (13 HCs, 15 MDD patients) were excluded from all analysis after data collection resulting in a final sample of n=122 MDD patients and n=176 HCs. The study was approved by the Ethics Committee of the University of Münster. After a complete description of the study to the participants, written informed consent was obtained. All participants received a financial compensation.
DNA Analysis
Venous blood sample collection was carried out by default between 1700 and 1900 hours. DNA extraction methods, genotyping of the functional promoter polymorphisms 5-HTTLPR (long (L) and short (S) allele) and rs25531 (common A and rare G allele), and bisulfite specific sequencing analyses of the AluJb methylation rates in the SLC6A4 promoter region were all performed as described previously (Dannlowski et al, 2014). Focus was put on the same eight of nine CpG sites (bp80, bp86, bp102, bp110, bp161, bp163, bp173, bp177) spanning the AluJb as before (Dannlowski et al, 2014; Supplementary Figure S1) for comparison reasons. Quality controls were extended to three independent repetitions of standard PCR and sequencing reactions with concordance checks of triplicate data (exclusion of outlier in case of standard deviation (SD)>0.05). Further, ten random patient DNAs were used for additional independent bisulfite conversions followed by triplicate PCRs and sequencing, resulting in high concordance rates (overall SD=0.018; max. SD=0.055) except for CpG site bp102 (SD=0.085, SD(max)=0.112), which therefore was excluded from further analysis.
The genotype distributions of 5-HTTLPR and rs25531 did not differ significantly from the expected numbers calculated according to the Hardy–Weinberg equilibrium in both, patients and controls (p>0.05). For subsequent analysis, subjects were grouped into carriers of no risk allele (LALA homozygotes) vs carriers of one risk (S or LG) allele vs carriers of two risk alleles (for details see Table 1), with the LG allele treated as risk allele based on comparable levels of serotonin transporter expression between S and LG alleles (Hu et al, 2006). To maximize statistical power, groups were also dichotomized into no risk allele (LALA homozygotes) vs one or two risk (S/LG) allele carriers. 5-HTTLPR/rs25531 genotype group frequencies were compared between patients and controls using a chi square test and did not differ statistically between them (see Table 1).
fMRI Methods
The experimental fMRI paradigm was frequently used to elicit a robust and replicable amygdala response across an array of imaging genetics studies (eg, Dannlowski et al, 2011; Nikolova et al, 2014). The paradigm, which utilized a face-processing task (faces with anger or fear expressions), alternating with a sensorimotor control task was conducted as described previously (Dannlowski et al, 2011; see Supplementary Material and Figure S2). Functional images were realigned and unwarped, spatially normalized to standard MNI space (Montreal Neurological Institute), and smoothed using a Gaussian kernel (6 mm FWHM). As noted above, 28 subjects had to be excluded due to excessive head movements (exclusion criterion >3 mm and/or 3°). The remaining n=298 datasets were free from movement effects. The onsets and durations of the two experimental conditions (faces and shapes) were modelled using a canonical hemodynamic response function in the context of the general linear model (GLM). The model was corrected for serial correlations and a high-pass filter of 128 seconds was applied to reduce low frequency noise. An individual contrast image was generated in each fixed-effects first-level analysis comparing activation in response to fear-relevant faces with the control condition as baseline. The resulting contrast images were further used in second-level random-effects group analyses.
Statistical Analysis
Analyses were conducted using SPSS (Version 23.0, IBM, Chicago, IL, USA), R programming language (Version R3.3.2, Vienna, Austria) and Statistical Parametric Mapping (SPM8, http://www.fil.ion.ucl.ac.uk/spm). Beta regression analyses were performed using algorithms implemented in the R package betareg (Cribari-Neto and Zeileis, 2010).
Effects of SLC6A4 AluJb Methylation
For AluJb methylation analyses, principal component analysis (PCA) was used for data reduction of the seven CpG sites. The PCA yielded one principal component with an eigenvalue > 1. This explained 55.4% of the total variance. To facilitate the analysis, a factor-based score was constructed by taking the average of CpG sites with a recommended cut-off factor loading of >.35 (Hair et al, 1998). Since methylation values are bounded between 0 and 1, we used beta regression modeled on the logit scale to model DNA methylation (Ferrari and Cribari-Neto, 2004). According to our first study goal, we performed a beta regression model predicting mean AluJb methylation by diagnosis (MDD vs HC), 5-HTTLPR/rs25531 genotype group (carriers of 0 vs 1 and carriers of 0 vs 2 risk S or LG alleles), LTE-Q, and the interaction term 5-HTTLPR/rs25531 × LTE-Q. We included two additional confounders, sex and age, based on previous significant associations with SLC6A4 methylation (Philibert et al, 2008; Domschke et al, 2014). Since beta-regressions do not allow omnibus F-tests, we focused on the differences between carriers of no risk allele vs one and no risk allele vs two risk alleles, based on former research showing significant differences mainly between no-risk allele carriers and carriers of one or two risk alleles (eg, Alexander et al, 2014; Olsson et al, 2010). However, for comparison reasons, all analyses were also conducted using an analysis of covariance (ANCOVA) (Supplementary Table S5 and S7).
For MDD patients the regression model was extended by number of depressive episodes and MLI since the duration of depression and medication effects have already been associated with SLC6A4 methylation (eg, Kang et al, 2013; Domschke et al, 2014). To explore potentially confounding effects we added body mass index (BMI) as regressor in additional regression models. Continuous predictors were standardized in order for their coefficients to be more comparable. All analyses were performed using a cut-off p-value of 0.05.
fMRI Analyses
According to our second study goal, the effects of diagnosis and AluJb methylation on amygdala reactivity were investigated using an ANCOVA in SPM with AluJb methylation as covariate and diagnosis and 5-HTTLPR/rs25531 as between-subject factors, sex as regressor of no interest, and diagnosis × AluJb methylation as interaction term. To meet our third goal, we further investigated the joint effect of AluJb methylation × 5-HTTLPR/rs25531 within the model. Given the study’s primary focus on the amygdala, all calculations were restricted to the bilateral amygdala as defined by Tzourio-Mazoyer et al. (2002) using an anatomical mask created with the Wake Forest University (WFU) Pick Atlas (Maldjian et al, 2003). A rigorous p<0.05 family-wise error (FWE) correction on the voxel-level was applied for all analyses. For each subject the mean contrast values of the peak voxel from significant results of the fMRI ANCOVA analysis were extracted from SPM and further analyzed in SPSS. An ANCOVA was performed to investigate whether other factors influenced possible associations of AluJb methylation and 5-HTTLPR/rs25531 with amygdala reactivity by adding LTE-Q, age, and BMI as nuisance covariates. Again, for MDD patients the nuisance covariates were further expanded by number of depressive episodes and MLI.
Results
SLC6A4 AluJb Methylation Rates
Analyses of individual AluJb methylation profiles revealed different methylation rates for all seven CpG sites (Supplementary Table S2). Except one CpG site (bp110) all other six CpG sites were approximately normally distributed (Shapiro Wilk Tests, all p>0.05) and had factor loadings >0.35 and thus, were used to calculate the mean methylation rate as an estimate of SLC6A4 promoter AluJb methylation status (Supplementary Table S2).
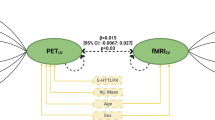
The regression model predicting mean AluJb methylation by diagnosis, 5-HTTLPR/rs25531, LTE-Q, the interaction 5-HTTLPR/rs25531 × LTE-Q, sex, and age explained 18.2% of the variance (see Supplementary Table S4 for beta-regression coefficients). Regarding our first study goal, mean AluJb methylation rate was significantly lower in MDD as compared to HC (z=−2.61, p=0.009, Figure 1a) and lower AluJb methylation was associated with higher LTE-Q scores (z=2.35, p=0.019). 5-HTTLPR/rs25531 was not associated with AluJb methylation, however, there was a significant interaction of 5-HTTLPR/rs25531 and LTE-Q, revealing that individuals carrying risk (S/LG)-alleles had different associations with LTE-Q compared to LALA homozygotes (1xS/LG vs LALA: z=−2.35, p=0.019, 2 × S/LG vs LALA: z=−2.93, p=0.003, Figure 1b): When confronted with stressors, carriers of one risk allele had lower and carriers of two risk alleles had lowest AluJb methylation compared to non-risk allele carriers, while carriers of one vs two risk alleles did not differ significantly (data not shown). Further, AluJb methylation was significantly lower in males (z=6.03, p<0.001), while age and BMI were not significantly associated. A correlation matrix with all variables as well as a sensitivity analysis including higher order interaction terms are provided in Supplementary Table S3 and S8 in the Supplementary Material.
Association of mean AluJb methylation rate in the serotonin transporter gene (SLC6A4) promotor with major depressive disorder (MDD), List of Threatening Experiences Questionnaire (LTE-Q), and 5-HTTLPR/rs25531. (a) AluJb mean methylation rates were significant lower in n=122 MDD patients compared to n=176 healthy controls (HC) (MDD vs HC: b=−0.05, z=−2.61, p=0.009). (b) The interaction of 5-HTTLPR/rs25531 and LTE-Q shows that when confronted with high threatening experiences during the past year, carriers of one risk allele appeared to have lower AluJb methylation as compared to non-risk allele carriers (1 × S/LG vs LALA: b=−0.05, z=−2.35, p=0.019). Similarly, carriers of two risk alleles appear to have lower AluJb methylation as compared to non-risk allele carriers (2 × S/LG vs LALA: b=−0.07, z=−2.94, p=0.003). The coefficient of the standardized LTE-Q was positive for LALA (b=0.04, z=2.35, p=0.019), but negative for 1 × S/LG (b=−0.01, z=−0.72, p=0.474) and 2 × S/LG (b=−0.03, z=−1.65, p=0.099). Error bars and shaded areas depict 95% CIs. All reported coefficients are on the logit-scale.
In MDD patients only, a positive association between AluJb methylation and number of depressive episodes was found (z=2.08, p=0.038) (Supplementary Table S6). Furthermore, the interaction of the dichotomized HTTLPR/rs25531 with LTE-Q (z=−2.21, p=0.027) and the association with sex (z=−3.95, p<0.001) remained significant, while MLI, LTE-Q, age, and BMI revealed no significant associations.
fMRI Results
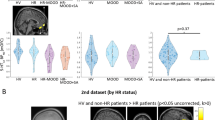
The ROI analysis of the bilateral amygdala reactivity revealed no significant main effect of diagnosis and, concerning our second study goal, no significant main effect of AluJb methylation. However, a significant interaction of AluJb methylation and diagnosis for the right amygdala reactivity appeared (x=34, y=−2, z=-20, F(1,289)=12.23, p=0.00054, pFWE=0.039, k=2, Figure 2a and b). Post hoc analysis revealed a strong positive association of AluJb methylation and the right amygdala reactivity for MDD patients (right: x=34, y=2, z=−22, t(119)=3.70, p=0.00016, pFWE=0.01, k=24, Figures 2c and d), while there was no significant association for HCs.
Association of mean AluJb methylation rate and amygdala reactivity. (a) Statistical parametric map illustrating diagnosis × mean AluJb methylation rate interaction in the right amygdala (x=34, y=−2, z=−20, F(1,289)=12.23, p=0.00054 pFWE=0.039, k=2). Color bar, F values (df=1,289). (b) Scatter plot from the peak voxel of (a) calculated on the contrast estimates from the right amygdala illustrating the interaction of the diagnosis × mean AluJb methylation rate for both, patients with major depressive disorder (MDD) and healthy controls (HC). (c) Results from the association of AluJb methylation rate and amygdala reactivity for MDD patients only (right: x=34, y=2, z=−22, t(119)=3.70, p=0.00016, pFWE=0.01, k=24). Color bar, t values (df=118). (d) Scatter plot for MDD patients from the right amygdala peak voxel of (c), illustrating the positive correlation of the mean AluJb methylation rate with the right amygdala reactivity (r=0.22). (a,c) Coronal view (y=−2), radiological convention, for display reasons the voxel threshold was set to p<0.05, uncorrected.
Regarding our third study goal, no significant interaction effect of 5-HTTLPR/rs25531 and AluJb methylation emerged in all participants as well as in both groups separately. However, in accordance with literature (Murphy et al, 2013a) risk allele carriers revealed a higher reactivity in the right amygdala compared to LALA homozygotes (x=34, y=−2, z=−28, t(289)=3.30, p=0.00054, pFWE=0.036, k=2). All abovementioned associations with amygdala reactivity were not affected by sex, LTE-Q, BMI, age, number of depressive episodes or MLI as analyzed by subsequent multiple regression with t-values ranging from t=−1.48 to t=1.95.
Discussion
In the present study, we found significant differences in SLC6A4 promoter methylation between MDD patients and HCs in a large sample. This is in line with recent work showing differences in SLC6A4 methylation level between a small sample of MDD patients and controls (Iga et al, 2016). Furthermore, as assumed in our previous study (Dannlowski et al, 2014) patients with MDD indeed revealed lower SLC6A4 promotor AluJb methylation compared to HCs (Figure 1a). In addition to MDD, AluJb methylation was lower in participants with higher experiences of threatening events during the past year (higher LTE-Q). Since mean LTE-Q was distinctly higher in MDD patients (Table 1), this might explain their distinctly lower mean methylation. A reaction of lowered AluJb methylation when confronted with psychological stressors might lead to the assumption that SLC6A4 AluJb hypomethylation in MDD could represent an underlying endogenous response mechanism to stress. However, since mediation analysis for beta-regression models have not been invented yet to our knowledge, the assumption could not be further tested and stays hypothetical. Global Alu hypomethylation has already been associated with stress exposure, such as for night shift workers (Bollati et al, 2010), individuals with socially disadvantaged status (Subramanyam et al, 2013), and the development of post-traumatic stress disorder after military service (Rusiecki et al, 2012). Interestingly, low Alu methylation was also associated with reduced DNA methyltransferase (DNMT) expression (Deplus et al, 2014) and such reduced DNMT was found in amygdala tissue of postmortem depressed suicides compared to controls (Poulter et al, 2008). Thus, the MDD patients in our present study might reveal lower AluJb methylation compared to HCs also based on reduced DNMT expression. However, this remains speculative since no DNMT levels were examined.
Hypomethylation of Alu elements was generally associated with decreased protein synthesis of nearby genes (cf. Landry et al, 2001), presumably via their function as genomic elements or as transcribed RNA (Wang and Huang, 2014). The AluJb element in the SLC6A4 promoter studied here belongs to the evolutionary early subfamily of AluJb and contains, as expected for this group, numerous transcription-factor-binding sites, particularly that for the repressive PAX6 (Dannlowski et al, 2014). Further, AluJb is located in antisense direction near the transcription start site, allowing possibly exonization, which would lead to alternative SLC6A4 mRNA competing with the original mRNA (Schmitz and Brosius, 2011) and as already shown for an AluJb element in the leptin receptor gene (Huh et al, 2010). Lower SLC6A4 AluJb methylation might allow, eg, more repression of gene expression via PAX6 binding or more alternative (but less original) SLC6A4 mRNA via exonization, both possibly resulting in lower 5-HTT levels (for a detailed discussion see (Dannlowski et al, 2014). Another possibility is the active Alu transcription via RNA polymerase III resulting in free transcribed Alu ncRNAs, which have been demonstrated to inhibit gene expression, and which can be highly adaptive and sensitive to stress (reviewed in (Wang and Huang, 2014). This might also hold true for our SLC6A4 AluJb in case of low or no methylation allowing RNA polymerase III binding. Taken together, recent findings suggest that lower SLC6A4 AluJb methylation could cause inhibition of SLC6A4 gene expression and thus declined levels of 5-HTT. In line with this, MDD patients reveal less 5-HTT-binding compared to HCs (Yeh et al, 2015). However, these considerations remain speculative since no experimental data exists for the impact of AluJb methylation on SLC6A4 expression.
Our results show in agreement with previous studies (Bollati et al, 2010; Subramanyam et al, 2013) lower AluJb methylation associated with high stressful experiences. Interestingly, we also found a gene × environment interaction predicting AluJb methylation: when confronted with high stressful experiences in the past year HTTLPR/rs25531 risk allele carriers revealed lower AluJb methylation compared LALA homozygotes (Figure 1b). Considering the opposing effect of SLC6A4 AluJb methylation on gene expression (lower methylation related to declined 5-HTT levels as suggested above) compared to CpG island promoter methylation (higher methylation related to declined 5-HTT levels), our results are in line with studies reporting high CpG island methylation associated with high acute as well as chronic stress (Duman and Canli, 2015) in risk allele carriers. In contrast to such current stress, IJzendoorn and colleagues reported responses to past stress: here less unresolved loss or trauma was associated with high SLC6A4 CpG island methylation in risk allele carriers (IJzendoorn et al, 2010), indicating possibly adaptive processes as longterm outcome of stress-related epigenetic changes. The less unresolved loss or trauma might be due to the abovementioned declined 5HTT-levels, which should lead to increased synaptic 5HT availability, comparable to an endogenous SSRI-like effect. Such an adaptive effect might also exist in our MDD patients, since decreased number of depressive episodes were associated with lower AluJb methylation. However, taking an additional performed 4-way interaction with diagnosis × 5-HTTLPR/rs25533 × LTE-Q × sex into account it appeared that sex and diagnosis further impact the gene × environment interaction (see supplementary Table S8 and Supplementary Figure S3). Further, we could replicate a finding of higher SLC6A4 methylation in female compared to male participants (Philibert et al, 2008). Gender specific DNA methylation might reflect hormonal influences during early development stages and possibly contributes to gender specific prevalence of MDD (Uddin et al, 2013). Future studies are needed to deepen the understanding of epigenetic effects on gender differences in the vulnerability for psychiatric disorders.
A possible connecting link between AluJb methylation, stress and MDD clinical appearance are DNMTs. DNMT expression in the nucleus accumbens promotes depression-like behavior in rodents (LaPlant et al, 2010) and increased global DNA methylation as well as genetic variation in DNMTs has already been associated with suicide attempts in psychiatric patients (Murphy et al, 2013b). Furthermore, DNMT inhibition has been shown to downregulate Alu methylation (Deplus et al, 2014), enhance resilience to chronic stress, induce antidepressant effects, and changes in neuronal stress-induced DNA methylation in rodents (Sales and Joca, 2016). However, it remains possible, that not DNMTs but antidepressant treatment changed the AluJb methylation, although extensive subsequent analysis of type and duration of medication revealed no significant association neither with AluJb methylation nor amygdala reactivity.
On a neurofunctional level, we observed an interaction of AluJb methylation and the diagnosis of MDD in the right amygdala responsiveness to emotional faces. Again, assuming the opposing effect of AluJb methylation on gene expression compared to CpG island methylation in the SLC6A4 promoter, our findings are in line with recent studies reporting SLC6A4 CpG island methylation as a predictor for amygdala reactivity in HC and MDD patients (Frodl et al, 2015; Nikolova et al, 2014). Further, lower AluJb methylation in MDD patients was strongly associated with reduced amygdala reactivity. Our results are pointing in the same direction as studies in rodents reporting changes of retrotransposon expression in the basolateral amygdala after stress exposure (Ponomarev et al, 2010). However, the spatial resolution of BOLD fMRI together with individual variations in amygdala morphometry does not allow direct conclusions about single nuclei. Endogenous SSRI-like effects, as the abovementioned possible consequence of lower AluJb methylation, might cause such decreased amygdala reactivity in response to emotional faces, an effect already shown after pharmacological treatment with SSRIs in MDD patients (Ruhé et al, 2014). In HCs, an association between amygdala reactivity and AluJb methylation might have been absent due to lower LTE-Q levels and/or more successful coping strategies in combination with sufficient demethylation when confronted with stress. However, since no neuronal 5-HTT mRNA levels can be collected in living participants, this interpretation remains speculative.
Analysis of joint effects of 5-HTTLPR/rs25531 and AluJb methylation on amygdala reactivity revealed no significant associations. However, we could replicate a controversially discussed finding of risk allele carriers exhibiting greater threat-related amygdala reactivity relative to non-carriers (Murphy et al, 2013a). Based on a large sample, meeting the need for high statistical power (Murphy et al, 2013a), our data adds notable evidence to the hypothesis that 5-HTTLPR/rs25531 represents an important player in the neural activity during emotion processing.
Our results predicting amygdala reactivity were limited to the right amygdala since weaker association in the left amygdala did not withstand FWE correction. This corroborates the fMRI paradigm used in our study, which mainly increases activation in the right amygdala (Lanteaume et al, 2007). Further it might reflect a functional asymmetry of the amygdala demanding further investigations (Baas et al, 2004), since the majority of studies analyzing the amygdala reported lateralized results.
Some limitations have to be acknowledged. First, DNA methylation was measured via whole EDTA-blood. Inter-subject heterogeneity in blood cell type proportions might potentially confound methylation levels and the following PCA analysis in our study (Jaffe and Irizarry, 2014). Furthermore, measuring methylation in the periphery of the body, a direct correlation to methylation levels in the brain cannot be determined. However, recent work indicates substantial correlations between SLC6A4 methylation of CpG sites in peripheral blood leukocytes and SLC6A4 methylation in post mortem amygdala tissue in healthy controls and patients with MDD (Riese et al, 2014). In addition, SLC6A4 promoter methylation based on peripheral blood cells has already been mapped onto individual differences in serotonin biosynthesis in vivo (Wang et al, 2012). Besides, influences of adjacent CpG sites and other functional polymorphisms could be biologically relevant and potentially confound our results. BDNF, for instance, has frequently been shown to interact with SLC6A4 on behavioral, transcriptional, and epigenetic levels (Ignácio et al, 2014). More research is needed to examine further genetic and epigenetic factors involved in the association of MDD and SLC6A4 AluJb methylation rates in detail. Although physical parameters like smoking status or physical exercises revealed no effects on SLC6A4 methylation previously (Alexander et al, 2014; Olsson et al, 2010) interfering influences cannot be excluded. Methylation rates did not differ with regard to medication status in our MDD sample, yet, influences of several drugs could not be conclusively clarified regarding the high heterogeneity of psychotropic drugs. Besides, amygdala reactivity has frequently been shown to be influenced by medication intake (Ruhé et al, 2014), which could not be determined in our sample, but should be taken into account in future replication studies.
Despite these limitations, our study provides new insights in the role of SLC6A4 AluJb methylation in MDD and amygdala reactivity and its associations with 5-HTTLPR/rs25531 and stress. Lowered AluJb methylation in MDD patients might represent an underlying endogenous response mechanism to higher stress levels. In conjunction with a shorter history of MDD and decreased amygdala reactivity, AluJb hypomethylation might even point to an adaptive epigenetic process, maybe via theoretically possible endogenous antidepressant-like effects by decreasing 5-HTT expression rates. The present study therefore is the first, which highlights the possible role of repetitive (Alu) elements in MDD and which might indicate new promising targets for therapeutic interventions.
Funding and disclosure
The study was supported by grants of Innovative Medizinische Forschung (IMF) of the Medical Faculty of Münster (DA120903 to UD, DA111107 to UD, and HO221003 to CH and UD), the German Research Foundation (DFG, grant FOR2107 DA1151/5-1 to UD; SFB-TRR58, Project C09 to UD) and the Interdisciplinary Center for Clinical Research (IZKF) of the medical faculty of Münster (grant Dan3/012/17 to UD). VA is member of advisory boards and/or gave presentations for the following companies: Astra Zeneca, Janssen-Organon, Lilly, Lundbeck, Servier, Pfizer, Otsuka, Trommsdorff, and Wyeth. He also receives funds from the German Ministry of Education and Research (BMBF) and from the European Union (EU-FP7). BTB is member of advisory boards, received funding and/or gave presentations for the following companies: AstraZeneca, Lundbeck, Pfizer, Servier, and Wyeth. He receives funding from the National Health and Medical Research Council (NHMRC) Australia. HK has received consultation fees from MR:comp GmbH, Testing Services for MR Safety. These affiliations have no relevance to the work covered in the manuscript. The remaining authors declare no conflict of interest.
References
Alexander N, Wankerl M, Hennig J, Miller R, Zänkert S, Steudte-Schmiedgen S et al (2014). DNA methylation profiles within the serotonin transporter gene moderate the association of 5-HTTLPR and cortisol stress reactivity. Transl Psychiatry 4: e443.
American Psychiatric Association Diagnostic and Statistical Manual of Mental Disorders. American Psychiatric Association: Washington, DC; (1994).
Baas D, Aleman A, Kahn RS (2004). Lateralization of amygdala activation: a systematic review of functional neuroimaging studies. Brain Res Rev 45: 96–103.
Bastiaansen JA, Vries YA, de, Munafò MR (2015). Citation distortions in the literature on the serotonin-transporter-linked polymorphic region and amygdala activation. Biol Psychiatry 78: e35–e36.
Beck AT BDI, Beck Depression Inventory: Manual. Psychological Corp.: San Antonio, TX and Harcourt Brace Jovanovich: New York, NY, (1987).
Bollati V, Baccarelli A, Sartori S, Tarantini L, Motta V, Rota F et al (2010). Epigenetic effects of shiftwork on blood DNA methylation. Chronobiol Int 27: 1093–1104.
Brugha TS, Cragg D (1990). The list of threatening experiences: the reliability and validity of a brief life events questionnaire. Acta Psychiatr Scand 82: 77–81.
Cribari-Neto F, Zeileis A (2010). Beta regression in R. J Stat Softw 34: 1–24.
Dannlowski U, Konrad C, Kugel H, Zwitserlood P, Domschke K, Schöning S et al (2010). Emotion specific modulation of automatic amygdala responses by 5-HTTLPR genotype. Neuroimage 53: 893–898.
Dannlowski U, Kugel H, Franke F, Stuhrmann A, Hohoff C, Zwanzger P et al (2011). Neuropeptide-S (NPS) receptor genotype modulates basolateral amygdala responsiveness to aversive stimuli. Neuropsychopharmacology 36: 1879–1885.
Dannlowski U, Kugel H, Redlich R, Halik A, Schneider I, Opel N et al (2014). Serotonin transporter gene methylation is associated with hippocampal gray matter volume. Hum Brain Mapp 35: 5356–5367.
Deininger P (2011). Alu elements: know the SINEs. Genome Biol 12: 236.
Deplus R, Blanchon L, Rajavelu A, Boukaba A, Defrance M, Luciani J et al (2014). Regulation of DNA methylation patterns by CK2-mediated phosphorylation of Dnmt3a. Cell Rep 8: 743–753.
Domschke K, Tidow N, Schwarte K, Deckert J, Lesch K-P, Arolt V et al (2014). Serotonin transporter gene hypomethylation predicts impaired antidepressant treatment response. Int J Neuropsychopharmacol 17: 1167–1176.
Duman EA, Canli T (2015). Influence of life stress, 5-HTTLPR genotype, and SLC6A4 methylation on gene expression and stress response in healthy Caucasian males. Biol Mood Anxiety Disord 5: 2.
Ekman P, Friesen WV. Pictures of Facial Affect. Consulting Psychologists Press: Palo Alto, CA; (1976).
Ferrari S, Cribari-Neto F (2004). Beta regression for modelling rates and proportions. J Appl Stat 31: 799–815.
Frodl T, Koutsouleris N, Bottlender R, Born C, Jäger M, Mörgenthaler M et al (2008). Reduced gray matter brain volumes are associated with variants of the serotonin transporter gene in major depression. Mol Psychiatry 13: 1093–1101.
Frodl T, Szyf M, Carballedo A, Ly V, Dymov S, Vaisheva F et al (2015). DNA methylation of the serotonin transporter gene (SLC6A4) is associated with brain function involved in processing emotional stimuli. J Psychiatry Neurosci 40: 140180.
Galea S, Nandi A, Vlahov D (2005). The epidemiology of post-traumatic stress disorder after disasters. Epidemiol Rev 27: 78–91.
Hair JF, Tatham RL, Anderson RE, Black W Multivariate Data Analysis. Prentice Hall: Upper Saddle River, NJ; (1998).
Hariri AR, Mattay VS, Tessitore A, Kolachana B, Fera F, Goldman D et al (2002). Serotonin transporter genetic variation and the response of the human amygdala. Science 297: 400–403.
Hassel S, Almeida JRC, Kerr N, Nau S, Ladouceur CD, Fissell K et al (2008). Elevated striatal and decreased dorsolateral prefrontal cortical activity in response to emotional stimuli in euthymic bipolar disorder: no associations with psychotropic medication load. Bipolar Disord 10: 916–927.
Hu X-Z, Lipsky RH, Zhu G, Akhtar LA, Taubman J, Greenberg BD et al (2006). Serotonin transporter promoter gain-of-function genotypes are linked to obsessive-compulsive disorder. Am J Hum Genet 78: 815–826.
Huh J-W, Kim Y-H, Kim D-S, Park S-J, Lee S-R, Kim S-H et al (2010). Alu-derived old world monkeys exonization event and experimental validation of the LEPR gene. Mol Cells 30: 201–207.
Iga J, Watanabe S, Numata S, Umehara H, Nishi A, Kinoshita M et al (2016). Association study of polymorphism in the serotonin transporter gene promoter, methylation profiles, and expression in patients with major depressive disorder. Hum Psychopharmacol Clin Exp 31: 193–199.
Ignácio ZM, Réus GZ, Abelaira HM, Quevedo J (2014). Epigenetic and epistatic interactions between serotonin transporter and brain-derived neurotrophic factor genetic polymorphism: insights in depression. Neuroscience 275: 455–468.
IJzendoorn MH, van, Caspers K, Bakermans-Kranenburg MJ, Beach SRH, Philibert R (2010). Methylation matters: interaction between methylation density and serotonin transporter genotype predicts unresolved loss or trauma. Biol Psychiatry 68: 405–407.
Jaffe AE, Irizarry RA (2014). Accounting for cellular heterogeneity is critical in epigenome-wide association studies. Genome Biol 15: R31.
Kaer K, Speek M (2013). Retroelements in human disease. Gene 518: 231–241.
Kang H-J, Kim J-M, Stewart R, Kim S-Y, Bae K-Y, Kim S-W et al (2013). Association of SLC6A4 methylation with early adversity, characteristics and outcomes in depression. Prog Neuropsychopharmacology Biol Psychiatry 44: 23–28.
Kaufman J (2015). Reply to: citation distortions in the literature on the serotonin-transporter-linked polymorphic region and amygdala activation. Biol Psychiatry 78: e37–e38.
Landry JR, Medstrand P, Mager DL (2001). Repetitive elements in the 5’ untranslated region of a human zinc-finger gene modulate transcription and translation efficiency. Genomics 76: 110–116.
Lanteaume L, Khalfa S, Regis J, Marquis P, Chauvel P, Bartolomei F (2007). Emotion induction after direct intracerebral stimulations of human amygdala. Cereb Cortex 17: 1307–1313.
LaPlant Q, Vialou V, Covington HE, Dumitriu D, Feng J, Warren BL et al (2010). Dnmt3a regulates emotional behavior and spine plasticity in the nucleus accumbens. Nat Neurosci 13: 1137–1143.
Maldjian JA, Laurienti PJ, Kraft RA, Burdette JH (2003). An automated method for neuroanatomic and cytoarchitectonic atlas-based interrogation of fMRI data sets. NeuroImage 19: 1233–1239.
Marcus M, Yasamy MT, Ommeren M, van, Chisholm D, Saxena S (2012) Depression: A Global Public Health Concern. World Health Organization: Geneva, Switzerland.
Miousse IR, Chalbot M-CG, Lumen A, Ferguson A, Kavouras IG, Koturbash I (2015). Response of transposable elements to environmental stressors. Mutat Res Mutat Res 765: 19–39.
Murphy SE, Norbury R, Godlewska BR, Cowen PJ, Mannie ZM, Harmer CJ et al (2013a). The effect of the serotonin transporter polymorphism (5-HTTLPR) on amygdala function: a meta-analysis. Mol Psychiatry 18: 512–520.
Murphy TM, Mullins N, Ryan M, Foster T, Kelly C, McClelland R et al (2013b). Genetic variation in DNMT3B and increased global DNA methylation is associated with suicide attempts in psychiatric patients. Genes Brain Behav 12: 125–132.
Nikolova YS, Koenen KC, Galea S, Wang C-M, Seney ML, Sibille E et al (2014). Beyond genotype: serotonin transporter epigenetic modification predicts human brain function. Nat Neurosci 17: 1153–1155.
Okada S, Morinobu S, Fuchikami M, Segawa M, Yokomaku K, Kataoka T et al (2014). The potential of SLC6A4 gene methylation analysis for the diagnosis and treatment of major depression. J Psychiatr Res 53: 47–53.
Olfson M, Marcus SC (2009). National patterns in antidepressant medication treatment. Arch Gen Psychiatry 66: 848–856.
Olsson CA, Foley DL, Parkinson-Bates M, Byrnes G, McKenzie M, Patton GC et al (2010). Prospects for epigenetic research within cohort studies of psychological disorder: a pilot investigation of a peripheral cell marker of epigenetic risk for depression. Biol Psychol 83: 159–165.
Philibert RA, Sandhu H, Hollenbeck N, Gunter T, Adams W, Madan A (2008). The relationship of 5HTT (SLC6A4 methylation and genotype on mRNA expression and liability to major depression and alcohol dependence in subjects from the Iowa Adoption Studies. Am J Med Genet B Neuropsychiatr Genet 147B: 543–549.
Ponomarev I, Rau V, Eger EI, Harris RA, Fanselow MS (2010). Amygdala transcriptome and cellular mechanisms underlying stress-enhanced fear learning in a rat model of posttraumatic stress disorder. Neuropsychopharmacology 35: 1402–1411.
Poulter MO, Du L, Weaver ICG, Palkovits M, Faludi G, Merali Z et al (2008). GABAA receptor promoter hypermethylation in suicide brain: implications for the involvement of epigenetic processes. Biol Psychiatry 64: 645–652.
Redlich R, Almeida JJR, Grotegerd D, Opel N, Kugel H, Heindel W et al (2014). Brain morphometric biomarkers distinguishing unipolar and bipolar depression. A voxel-based morphometry-pattern classification approach. JAMA Psychiatry 71: 1222–1230.
Riese H, Heuvel ER, van den, Snieder H, Dunnen WF, den, Plosch T, Kema IP et al (2014). Association between methylation of the SLC6A4 promoter region in peripheral blood leukocytes and methylation in amygdala tissue. Psychosom Med 76: 244–246.
Ruhé HG, Koster M, Booij J, Herk M, van, Veltman DJ, Schene AH (2014). Occupancy of serotonin transporters in the amygdala by paroxetine in association with attenuation of left amygdala activation by negative faces in major depressive disorder. Psychiatry Res Neuroimaging 221: 155–161.
Rusiecki JA, Chen L, Srikantan V, Zhang L, Yan L, Polin ML et al (2012). DNA methylation in repetitive elements and post-traumatic stress disorder: a case-control study of US military service members. Epigenomics 4: 29–40.
Sales AJ, Joca SRL (2016). Effects of DNA methylation inhibitors and conventional antidepressants on mice behaviour and brain DNA methylation levels. Acta Neuropsychiatr 28: 11–22.
Schmitz J, Brosius J (2011). Exonization of transposed elements: a challenge and opportunity for evolution. Biochimie 93: 1928–1934.
Su M, Han D, Boyd-Kirkup J, Yu X, Han JDJ (2014). Evolution of Alu elements toward enhancers. Cell Rep 7: 376–385.
Subramanyam MA, Diez-Roux AV, Pilsner JR, Villamor E, Donohue KM, Liu Y et al (2013). Social factors and leukocyte DNA methylation of repetitive sequences: the multi-ethnic study of atherosclerosis. PLoS ONE 8: e54018.
Tzourio-Mazoyer N, Landeau B, Papathanassiou D, Crivello F, Etard O, Delcroix N et al (2002). Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. NeuroImage 15: 273–289.
Uddin M, Sipahi L, Li J, Koenen KC (2013). Sex differences in DNA methylation may contribute to risk of PTSD and depression: a review of existing evidence. Depress Anxiety 30: 1151–1160.
Wang C, Huang S (2014). Nuclear function of Alus. Nucleus 5: 131–137.
Wang D, Szyf M, Benkelfat C, Provençal N, Turecki G, Caramaschi D et al (2012). Peripheral SLC6A4 DNA methylation is associated with in vivo measures of human brain serotonin synthesis and childhood physical aggression. PLoS ONE 7: e39501.
Wilkinson MB, Xiao G, Kumar A, LaPlant Q, Renthal W, Sikder D et al (2009). Imipramine treatment and resiliency exhibit similar chromatin regulation in the mouse nucleus accumbens in depression models. J Neurosci 29: 7820–7832.
Wittchen HU, Zaudig M, Fydrich T SKID - Strukturiertes Klinisches Interview DSM IV. Hogrefe: Göttingen; (1997).
Xia L, Yao S (2015). The involvement of genes in adolescent depression: a systematic review. Front Behav Neurosci 9: 329.
Yeh Y-W, Ho P-S, Chen C-Y, Kuo S-C, Liang C-S, Ma K-H et al (2015). Incongruent reduction of serotonin transporter associated with suicide attempts in patients with major depressive disorder: a positron emission tomography study with 4-[18F]-ADAM. Int J Neuropsychopharmacol 18: pyu065.
Zannas AS, West AE (2014). Epigenetics and the regulation of stress vulnerability and resilience. Neuroscience 264: 157–170.
Acknowledgements
We would like to thank Ahmad Hariri for providing the fMRI-paradigm and our reviewers for their very helpful comments that improved our manuscript substantially.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Neuropsychopharmacology website
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Schneider, I., Kugel, H., Redlich, R. et al. Association of Serotonin Transporter Gene AluJb Methylation with Major Depression, Amygdala Responsiveness, 5-HTTLPR/rs25531 Polymorphism, and Stress. Neuropsychopharmacol. 43, 1308–1316 (2018). https://doi.org/10.1038/npp.2017.273
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2017.273
This article is cited by
-
Epigenetic regulation in major depression and other stress-related disorders: molecular mechanisms, clinical relevance and therapeutic potential
Signal Transduction and Targeted Therapy (2023)
-
Losing DNA methylation at repetitive elements and breaking bad
Epigenetics & Chromatin (2021)
-
Epigenetic biotypes of post-traumatic stress disorder in war-zone exposed veteran and active duty males
Molecular Psychiatry (2021)
-
Genotype-dependent associations between serotonin transporter gene (SLC6A4) DNA methylation and late-life depression
BMC Psychiatry (2018)
-
Warning SINEs: Alu elements, evolution of the human brain, and the spectrum of neurological disease
Chromosome Research (2018)