Abstract
The single nucleotide polymorphism rs17070145 within the KIBRA gene (kidney and brain expressed protein) has been associated with variations in memory functions and related brain areas. However, previous studies yielded conflicting results, which might be due to divergent sample characteristics or task-specific effects. Therefore, we aimed to determine the impact of KIBRA genotype on learning and memory formation, and volume, microstructural integrity and functional connectivity (FC) of the hippocampus and its subfields in a well-characterized cohort of healthy older adults. One-hundred and forty subjects (72 women, age 50–80) were KIBRA genotyped and memory was tested using the Auditory Verbal Learning Task. Also, subjects underwent structural and resting-state functional magnetic resonance imaging at 3T. Subfields were delineated using automated segmentation (FreeSurfer software). Microstructural integrity was measured using mean diffusivity (MD) derived from diffusion tensor images. Seed-based analyses were used to assess FC patterns of the hippocampus. KIBRA T-allele carriers showed a trend for better memory performance, and in the hippocampus significantly higher volumes and partly lower MD, indicative for better microstructure, compared with non-T-allele carriers in the cornu ammonis (CA)2/3 and CA4/dentate gyrus subfields (all P⩽0.008, Bonferroni corrected). Also, T-allele carriers exhibited lower FC of the left hippocampus with areas outside the synchronized HC network. In sum, we could show for the first time that older T-allele carriers exhibited larger volumes and better microstructure within those hippocampus subfields that are implicated in long-term potentiation and neurogenesis, key features of memory processes. Moreover, T-allele carriers showed a more selective FC network of the hippocampus.
Similar content being viewed by others
Introduction
A single nucleotide polymorphism (SNP) in the gene encoding KIBRA (kidney and brain expressed protein), also referred to as WWC1 (WW and C2 domain containing 1), might contribute to interindividual differences in memory performance (Milnik et al, 2012; Papassotiropoulos et al, 2006; Schneider et al, 2010). Such variations are known to magnify during aging and may also influence timing and rate of late-life cognitive changes (Deary et al, 2004; Lindenberger et al, 2008). Understanding the basis of interindividual differences could help to develop novel strategies to maintain memory functions in old age (Lindenberger et al, 2008).
In a hypothesis-free genome-wide association study, Papassotiropoulos et al (2006) could demonstrate higher memory performance in KIBRA T-allele carriers compared with non-T-allele carriers. Although others could not confirm these results (eg, Need et al, 2008; Wersching et al, 2011), and one study even showed poorer performance in T-allele carriers (Nacmias et al, 2008), a recent meta-analysis supported the beneficial effect of the T-allele for memory (Milnik et al, 2012). A recent study provided evidence for better spatial memory especially in older T-allele carriers using a virtual reality navigation paradigm (Schuck et al, 2013). Also, a protective effect of T-allele carrier status for the risk of Alzheimer’s disease has been observed in most, but not all investigated cohorts so far (Burgess et al, 2011; Corneveaux et al, 2010; Hayashi et al, 2010; Wang et al, 2013b).
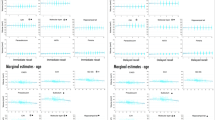
KIBRA has been suggested to act as a postsynaptic scaffold protein implicated in vesicle transport, transcriptional regulation and synaptogenesis (reviewed in Schneider et al, 2010), yet the exact underlying mechanisms of how KIBRA T-allele carrier status may influence memory performance is not fully understood. Using structural magnetic resonance imaging (MRI) in humans, Palombo et al (2013) recently found larger volumes of the hippocampus, a key region implicated in memory functioning, in young T-allele carriers compared with non-T-allele carriers. In addition, several task-related functional (f)MRI studies demonstrated differences in brain activation of memory-related brain networks between KIBRA genotype groups (Kauppi et al, 2011; Muse et al, 2013; Papassotiropoulos et al, 2006). For example, a weaker blood oxygen-level-dependent (BOLD) response during memory retrieval in the hippocampus was found for T-allele carriers (Papassotiropoulos et al, 2006), yet others found the opposite pattern (Kauppi et al, 2011), or a genotype-selective correlation with age (Muse et al, 2013). Considering large-scale brain networks, assessed using resting-state fMRI, a single study observed stronger synchronization patterns in the default mode and executive network in KIBRA C-allele carriers (Wang et al, 2013a). Taken together, similar to the behavioral results described above, neuroimaging studies yielded in part conflicting results (Kauppi et al, 2011; Muse et al, 2013; Papassotiropoulos et al, 2006).
Taken together, a putative beneficial effect of the KIBRA T-allele for memory performance is still a matter of debate, and whether T-allele carriers at older age exhibit differences in hippocampal volume, microstructure or synchronization patterns compared with non-T-allele carriers is unknown. Therefore, we analyzed memory performance as well as volume, microstructure and resting-state functional connectivity (FC) of the hippocampus and its subfields dependent on KIBRA T-allele carrier status in a homogenous sample of healthy older adults, which was balanced for sex and well characterized for cardiovascular risk factors.
Materials and methods
Participants
We recruited 141 healthy older volunteers (aged 50–80 years) via advertisements in Berlin, Germany. Inclusion criteria were no severe untreated medical, neurological or psychiatric disease, no history of diabetes mellitus type 2, a body mass index between 25 and 30 kg/m2, daily consumption of <50 g alcohol, <10 cigarettes, <6 cups of coffee, native German speaking and right handedness. The Mini Mental State Examination (Folstein et al, 1975) was used to rule out preexisting cognitive impairment (a score <26 points led to exclusion). All subjects underwent fasting blood sampling, neuropsychological testings including memory performance, and cerebral MRI for exclusion of brain pathologies and for subsequent brain analyses. Also, cardiovascular risk factors including systolic blood pressure and physical activity, as well as other genetic factors (APOE rs429358 and rs7412, BDNF rs6265, COMT rs4680) that have been previously implicated in episodic memory performance were assessed (Corder et al, 1993; Egan et al, 2003; Witte and Floel, 2012). Psychiatric comorbidity was monitored using the Beck’s Depression Inventory (BDI; Kuhner et al, 2007) and the State-Trait Anxiety Inventory (STAI X1; Laux et al, 1981). Individuals were stratified for the KIBRA-SNP rs17070145 genotype into T-allele carriers and non-T-allele carriers in line with previous studies (Milnik et al, 2012). One subject could not be genotyped for the KIBRA-SNP rs17070145 due to technical problems, leaving 140 for analysis (71 women, 69 men, mean age 63 years). In total, the sample comprised 69 T-allele carriers (54 CT and 15 TT) and 71 non-T-allele carriers (CC), see Table 1.
The study was approved by the Ethics Committee of the Charité University Medicine Berlin, Germany, and was in accordance with the Declaration of Helsinki. All subjects gave informed written consent before participation in the study, and received a small reimbursement at the end.
Blood Sampling and SNP Genotyping
Venous blood was collected after an overnight fast of at least 10 h. Blood measurements included glycated hemoglobin (HbA1c) as long-term marker of glucose control. DNA was extracted from whole blood using a blood mini-kit (Qiagen, Hilden, Germany) and stored at −80 °C until analysis. Genotyping of KIBRA (rs17070145, GenBank accession number NM_015238), BDNF (rs6265, GenBank accession number 2174122), COMT (rs4680, GenBank accession number AY341246) and APOE (rs429358, rs7412, GenBank accession number AF261279) were performed on a Sequenom MassARRAY iPLEX, Taqman Assay at the laboratory of Prof. Dr Dan Rujescu (University of Halle, Germany) following procedures described previously (O’Dwyer et al, 2012). Due to methodological failure, the genotype of COMT (rs4680) could not be conducted for one subject.
Memory Performance
Subjects were tested using the German version of the Rey Auditory Verbal Learning Test (AVLT; Lezak, 2004). Participants had to learn a list of 15 words within five immediate recall trials, followed by a 30-min delayed recall and recognition trial. We evaluated the following three subtasks: memory consolidation, delayed recall, and learning ability (Kerti et al, 2013). Memory consolidation was defined as the number of correct words recalled after the fifth trial subtracted by those correctly recalled after 30 min delay. The values were multiplied by −1 to create positive relations. Delayed recall represented the total number of remembered words after 30 min, whereas learning ability corresponds to the sum of words learned in all five trials. Tests were conducted following standard procedures by trained staff members who were blinded to individual genotype group status.
MRI Acquisition
MRI was performed on a Siemens Trio system operating at 3 T using a 12-channel head coil. We obtained high resolution 3D T1-weighted scans (3D magnetization prepared rapid acquisition with gradient echoes (MPRAGE); TR=1900 ms, TE=2.52 ms, 192 sagittal slices, voxel-size of 1.0 × 1.0 × 1.0 mm3, flip angle=9°), and a diffusion-weighted spin-echo echo-planar imaging sequence (TR=7500 ms, TE=86 ms, 61 axial slices, voxel size of 2.3 × 2.3 × 2.3 mm3; 64 directions with a b-value of 1000 s/mm2 and one or ten b0). Functional scans were obtained at rest using a T2*-weighted echo-planar imaging sequence (TR=2300 ms, TE=30 ms, 34 slices, voxel size of 3.0 × 3.0 × 4.0 mm3, flip angle=90°). Subjects were instructed to keep their eyes closed and not to think of anything in particular during this 6 min scan. The order of MRI sequences was as follows: localizer, resting-state fMRI, diffusion tensor imaging (DTI), T1-magnetization prepared rapid acquisition with gradient echoes, fluid-attenuated inversion recovery (for medical rating), angiography with contrast agent (for medical rating).
Hippocampal Volume
To estimate hippocampal volume, high-resolution T1-weighted images were first registered to MNI space by rigid body transformation using the software FSL 4.1 (www.fmrib.ox.ac.uk/fsl). Cortical and subcortical reconstruction and volumetric segmentation, including the left and right hippocampus, was performed by FreeSurfer 5.1.0 (http://surfer.nmr.mgh.harvard.edu/). Briefly, this process included motion correction, intensity normalization, and skull stripping using a watershed algorithm (more technical details are described in Fischl et al, 2002). FreeSurfer-based automated subfield segmentation was carried out according to Van Leemput et al, 2009 in order to extract individual volumes of cornu ammonis (CA) fields 1–4, dentate gyrus (DG) and subiculum. For the whole hippocampus and for each subfield (ie, CA1, CA2/3, CA4/DG, subiculum, tail), individual volumes were adjusted for intracranial volume (ICV) according to previous studies (Raz et al, 2005; Kerti et al, 2013) using the following formula: adjusted volume=raw individual volume − b × (individual ICV–whole group mean ICV). The coefficient b represents the slope of a whole-group-wise regression (enter mode) of the respective raw volume (ie, whole hippocampus or CA1 or CA2/3 or CA4/DG or subiculum or tail) on ICV.
Hippocampal Microstructure
Microstructure of the hippocampus was assessed by mean diffusivity (MD) estimated using DTI, in line with previous studies (den Heijer et al, 2012; Kerti et al, 2013). Therefore, a tensor model was fitted to the motion-corrected DTI data to create individual 3D images of fractional anisotropy and MD, this was done using FMRIBs Diffusion Toolbox (FDT) in FSL 4.1. Then, individual T1-weighted images were co-registered with FSL to the fractional anisotropy maps of the preprocessed DTI data using rigid-body transformations. These registrations were used to transform masks of the left and right hippocampus and hippocampus subfields (derived by FreeSurfer segmentation on the T1 images) to the MD maps. For extraction of the mean individual hippocampal MD values we used statistical analysis tools in FSL. Ten subjects had to be excluded from MD analysis because of image artifacts.
Hippocampal FC
To assess potential differences in FC of the hippocampus, we used a customized processing stream based on the 1000 Functional Connectomes Project (www.nitrc.org/projects/fcon_1000). Co-registered masks of the left and right hippocampus derived from FreeSurfer served as seeds for FC analyses, in line with previous studies (Andrews-Hanna et al, 2010; Rombouts et al, 2003; Witte et al, 2014). Preprocessing of individual functional scans comprised slice time correction, motion correction, spatial smoothing with a 6-mm full-width-half-maximum Gaussian kernel, temporal filtering (0.01–0.1 Hz), and de-trending using the software packages FSL and AFNI 2011 (http://afni.nimh.nih.gov/afni). The functional scans were normalized to the anatomical image using affine co-registrations in FSL. Noise due to motion, white matter, cerebrospinal fluid, and global change was removed from the functional signal by multiple regressions in FSL using commands of the fMRI Expert Analaysis Tool (FEAT), creating standardized residual BOLD-signal time series. Then, mean time series of both hippocampus seeds were correlated with times series of all other gray matter voxels using a general linear model approach within FSL (FMRIB’s local analysis of mixed effects, FLAME). The resulting Pearson’s r correlation coefficient values were then Fisher’s z-transformed using statistical analysis tools in FSL.
Statistical Analysis
SPSS 20 (PASW, SPSS; IBM, Armonk, NY) was used for statistical analysis unless indicated otherwise. Normal distributions were tested using the Kolmogorov–Smirnov test. We used the chi-squared test to test whether the distribution of KIBRA alleles was within Hardy–Weinberg equilibrium. The two KIBRA genotype groups (T-allele carriers vs non-T-allele carriers) were matched for age, sex, education, cardiovascular risk factors and APOE-, BDNF- and COMT-SNP carrier status (all P>0.05, see Table 1 for details). Accordingly, we calculated independent t-tests (normal distribution, two-sided) or Mann–Whitney U-tests (non-normal distribution) to detect significant differences between KIBRA genotype groups in memory performance, as well as hippocampal volume and microstructure. Corrections for multiple comparisons were applied using a Bonferroni threshold of P=0.05/3=0.015 for memory performance and P=0.05/5=0.01 for volume and microstructure of the hippocampus, respectively. A threshold of P=0.05/10=0.005 was applied when looking at the subfields separately for each hemispheres. FC group analyses included gray matter-voxelwise GLM statistics implemented in FSL (FLAME) of hippocampal FC between KIBRA T-carriers and non-T-carriers. Gaussian random field theory was used to correct multiple comparisons at the cluster level (clusterwise correction, z=2.3, P<0.05). In addition, we explored whether memory performance would be correlated with volumetric, microstructural and FC measures using bivariate Spearman’s correlations.
Results
The allelic distribution for the KIBRA-SNP rs17070145 was in Hardy–Weinberg equilibrium (10.7% TT carriers, 38.6% CT carriers and 50.7% TT carriers; χ2=0.93, 1 df, P=0.33). KIBRA genotype groups did not differ with regard to age, sex, education, cardiovascular risk factors and memory-associated gene variations including APOE-, BDNF- and COMT-SNP carrier status (all P>0.05, see Table 1 for details).
Memory Performance
We observed a trend for better memory consolidation (P=0.032, uncorrected) and better delayed recall (P=0.18, uncorrected) in healthy older KIBRA T-allele carriers compared with non-T-allele carriers (Figure 1, Table 2). These differences however did not reach statistical significance after correction for multiple testing.
Episodic memory performance of healthy older KIBRA T-allele carriers and non-T-allele carriers. T-allele carriers (gray squares, n=69) showed a trend for higher consolidation memory (P=0.032, uncorrected) and delayed recall performance (P=0.18, uncorrected) compared with non-T-allele carriers (black circles, n=71). No difference was seen in learning ability (P>0.6). Error bars indicate standard error.
Volume, Microstructure and FC of the Hippocampus
Considering the structure of the hippocampus, T-allele carriers exhibited larger hippocampus volumes than non-T-allele carriers within the CA2/3 and CA4/DG subfields (both t(138)<−2.7, Cohen’s d>0.45, P<0.008; Figure 2, Table 3).
Adjusted total and subfield volume of the hippocampal formation dependent on KIBRA genotype groups. KIBRA T-allele carriers (n=71) showed a significant larger volume of the CA2/3 and CA4/DG hippocampus subfields (all P=0.008, Bonferroni corrected) compared with non-T-allele carriers (n=69). CA, cornu ammonis; DG, dentate gyrus. Error bars indicate SE. **Significant at P<0.01 (Bonferroni-corrected), according to independent t-test.
To elucidate whether the differences in hippocampus subfield volume were subfield specific, we additionally conducted a repeated measures analysis of variance (ANOVA rm) with hippocampus subfields as dependent variables and KIBRA genotype group as independent factor. Accordingly, the interaction term genotype X subfield reached significance (F(4,135)=2.52, P=0.044), suggesting a selective effect of KIBRA genotype on CA2/3 and CA4/DG subfield volume.
With regard to microstructure, mean hippocampal MD was similar in both KIBRA genotype groups (P>0.05; Table 4). However, considering the subfields, lower MD was noted in the CA1, CA2/3, and CA4/DG for T-allele carriers compared with non-T-allele carriers, reaching significance for the right DG/CA4 region (t(128)=2.8, P=0.005, Bonferroni corrected; Table 4).
Resting-state FC analyses revealed a bilateral synchronization network of the right and left hippocampus seed with the temporal lobe and limbic structures, but also involving medial (pre)frontal areas, precuneus, and auditory cortices that were identical between groups (Figure 3, yellow–red, similar to previous studies, eg, Rombouts et al, 2003; Witte et al, 2014). Outside this network, KIBRA non-T-allele carriers showed a significant increase in FC between the left hippocampus seed and a cluster within visual areas, compared with T-carriers (Figure 3, blue (548 voxel), hot voxel: x=0, y=−82, z=10, P<0.001).
Differences in seed-based functional connectivity (FC) networks between KIBRA genotype groups using the left and right hippocampus as seeds. Both left hippocampus (upper panel) and right hippocampus (lower panel) seeds showed a bilateral synchronization network with the temporal lobe, limbic structures, medial (pre)frontal areas, precuneus, and auditory cortices (yellow–red). Compared with KIBRA T-allele carriers (n=71), non-T-allele carriers (n=69) showed increased correlation between the left hippocampus seed and a cluster in visual areas (blue), outside the mean hippocampal network. Color bars indicate t-values of significant voxels (cluster-based thresholding, P<0.05) superimposed on a study-specific template. Images are displayed in neurological convention, coordinates in millimeters according to MNI space. A, anterior; L, left; P, posterior; R, right.
According to bivariate correlations, learning and delayed recall performance was partly associated with larger hippocampal volumes, most pronounced in CA2/3 and CA4/DG subfields (see Supplementary Table S1 for r- and P-values). In addition, better mean microstructural integrity was associated with better memory performance (consolidation, delayed recall, learning ability, all P<0.019). No reliable correlations were found between FC changes and memory performance (all P>0.05).
Discussion
In this cohort of 140 healthy older adults, well characterized for cognitive functions and cardiovascular risk factors, we observed a trend for a better memory performance and significantly higher volumes and better microstructural integrity in the subfields comprising the CA2/3 and CA4/DG of the hippocampus in KIBRA T-allele carriers compared with non-T-carriers. In addition, KIBRA-T-carriers exhibited lower FC (ie, less de-differentiation) of the left hippocampus seed with areas that lie outside the synchronized hippocampus network.
KIBRA Genotype and Memory
Our results of trendwise better memory performance in KIBRA T-allele carriers are in line with previous studies in younger and older adults (Almeida et al, 2008; Kauppi et al, 2011; Papassotiropoulos et al, 2006; Schaper et al, 2008; Yasuda et al, 2010), but not all (Nacmias et al, 2008; Need et al, 2008; Wersching et al, 2011). Conflicting findings could be a result of slight variations in episodic memory tasks and outcome measures between studies, leading to potential over- or underestimation of global KIBRA-associated effects. Also, potential interacting effects with other genetic polymorphisms could help to explain divergent results (see eg, Preuschhof et al, 2010). In a recent meta-analysis including 17 independent samples (total n=8909), Milnik et al (2012) reported a significant effect of the KIBRA T allele on global measures of episodic memory that explained 0.5% of the variance, pointing to rather small overall effect sizes at the behavioral level in normal cognition.
KIBRA Genotype and the Hippocampus
Turning to the hippocampus, we found that healthy older T-allele carriers exhibit significantly larger volumes in comparison with non-T-allele carriers, similar to previous reports in young adults (Palombo et al, 2013), but not to the original GWAS study (Papassotiropoulos et al, 2006). However, note the relatively small structural MR-sample size in both studies, n=30 (Palombo et al, 2013) and n=28 (Papassotiropoulos et al, 2006), compared with our sample size of n=140. More specifically, we observed larger volumes of the CA2/3 and CA4/DG subfields in older T-allele carriers compared with non-T-allele carriers. This is again largely in line with Palombo et al (2013), who used manual delineations of the CA- and DG-areas. The subfield specificity of genotype effects to CA and DG subfields might be due to the high expression of KIBRA in these areas, compared with, for example, the subiculum (Johannsen et al, 2008). Note that the freesurfer method of defining hippocampal subfields might underestimate the CA1 subfield extent (De Flores et al, 2015), which could have contributed to a lack of significant effects between KIBRA genotype in this subfield in the current study compared with Palombo et al (2013).
Considering microstructural integrity, our study provides first-time evidence for a trend toward lower MD within the hippocampus gray matter in T-allele carriers compared with non-T-allele carriers, reaching statistical significance in the right CA4/DG subfield. DTI-based quantification of MD is considered as an inverse measure of neuronal integrity, ie, lower MD has been linked with better microstructure (den Heijer et al, 2012; Kerti et al, 2013), thus our findings point to a superior microstructure linked with the T allele. Non-significant results for mean hippocampus MD values according to T-allele carrier status might be explained by lateralization effects or microstructural heterogeneity between hippocampal subfields.
Using resting-state fMRI, we observed increased FC of the hippocampus with regions outside the ‘positively’ synchronized hippocampus network in KIBRA non-T-allele carriers. This might indicate a larger de-differentiation or lower selectivity of the hippocampus network in this group (Antonenko and Floel, 2014), possibly due to differences in KIBRA-related neuronal functioning in the CA and DG areas of the hippocampus (Johannsen et al, 2008). Our finding of selective effects of KIBRA genotype on hippocampal subfield volume supports this hypothesis.
So far, only one study investigated resting-state fMRI with regard to KIBRA genotype. Here, Wang et al (2013a) reported in a Chinese cohort of young adults increased synchronization patterns in the default mode and executive control networks in KIBRA C-allele carriers (eg, carriers of 1 or 0 T-alleles), paralleled by regional reductions in cortical gray matter volume. The increased synchronization pattern was discussed as a potential compensation mechanism, yet age- and genotype-related differences might limit a direct comparison between this and the current study. Using task-based fMRI, a recent study did not find overall genotype-related differences in hippocampal activation during an episodic memory task (Muse et al, 2014), which may be linked with genotype-independent FC patterns within the hippocampus synchronization network, as found in the current study (given we did not find significant differences within the network itself). However, opposing task-based fMRI-results were found by others in young (eg, weaker activation in T-allele carriers, Papassotiropoulos et al, 2006) and old samples (eg, stronger activation in T-allele carriers, Kauppi et al, 2011). Moreover, Muse et al (2014) observed that hippocampal activation in non-T-allele carriers showed a negative correlation with age that was not present in T-allele carriers. This latter finding could help to explain differential effects of KIBRA regarding hippocampal task activation between studies; however, more research is needed to clarify the direction and extent of KIBRA genotype on hippocampal FC.
Functional Implications and Underlying Mechanisms
Larger volume and better microstructure of the hippocampus in T-allele carriers might have translated into behavioral advantages, as these MR-based measures are thought to reflect better neuronal functioning due to, for example, enlarged axons, dendrites and cell bodies, improved vasculature and possibly even higher neurogenesis (Draganski and May, 2008; Thomas and Baker, 2012) as well as superior integrity of neuronal cell bodies, axons, and dendrites in cerebral gray matter (Beaulieu, 2002; Kantarci et al, 2005). Enlarged neuronal and vascular structures and higher neurogenesis might induce hippocampal growth, reflected by larger volumes. On the other hand, loss of membrane integrity or synaptic contacts would lead to an expansion of the extracellular space where water diffusivity is faster and subsequently MD is higher (for further discussion, see Beaulieu, 2002; Kantarci et al, 2005; Kerti et al, 2013). Indeed, larger volumes and better microstructural integrity of the hippocampus were correlated with learning and delayed recall performance in our cohort. Thus, structural differences between KIBRA genotypes might indicate neuronal correlates for subtle behavioural differences, trendwise only in our study.
Moreover, neuronal activity within specific subfields of the hippocampus might serve distinct aspects of memory encoding and retrieval (reviewed in Mueller et al, 2011; Squire, 2004). Therefore, subfield-specific effects of KIBRA genotype might further underline a proposed role for KIBRA genotype in particular aspects of memory formation. For example, the CA3 and dentate gyrus, characterized by long-term potentiation and adult neurogenesis, are thought to promote higher capacities for pattern separation and completion, crucial for encoding and retrieval of memories (Bakker et al, 2008; Bruel-Jungerman et al, 2007). Recently, larger volumes of the CA3 and 4 and DG region have been associated with long-term gains in associative memory in humans (Bender et al, 2013). Improved volume and microstructure in these regions in T-allele carriers might thus be associated with better consolidation and delayed recall memory in T-allele carriers, but may not relate to immediate learning, in line with our current trendwise results. Others, however, linked primarily the CA1 region (Mueller et al, 2011) or the subiculum (Zeineh et al, 2003), only marginally associated with KIBRA genotype in our cohort, with late retrieval and consolidation memory, which might also account for a lack of strong behavioural differences with regard to KIBRA genotype. Also, given the intrinsic functional and structural circuits of the medial temporal lobe including the hippocampal formation, attributing unique memory aspects to specific hippocampal subfields might result in oversimplification (Squire, 2004).
With regard to resting-state FC, previous studies suggested a link between larger de-differentiation of FC networks and worse cognitive performance (Andrews-Hanna et al, 2007; Antonenko and Floel, 2014). Thus, the observed increases in hippocampal FC with areas outside the hippocampal network could have contributed to poorer memory results in non-T-allele carriers. However, note that memory effects were trendwise only, and that we did not find significant correlations of increased de-differentiation with worse memory performance in the current study. Also, it cannot be excluded that a different FC network in individuals not carrying a KIBRA T-allele might have served as a compensatory mechanism regarding memory functions. A compensatory increase in FC has, for example, been discussed for patients with mild cognitive impairment, here, increased frontal-parietal resting-state FC might compensate for decreases in synchronity of the hippocampus network (Qi et al, 2010).
Limitations
Our study has several limitations. First, definite conclusions about causalities cannot be drawn, as our analyses are cross-sectional. Second, due to limited resolution of the MR system, partial volume effects could have biased the calculation of small hippocampus subfield volumes. Third, we did not acquire task-related fMRI measures. Fourth, we cannot exclude that other parameters could have contributed to the observed effects. However, note that KIBRA genotype groups did not differ with regard to education, MMSE, body mass index, vascular risk factors, APOE e4-carrier status and a list of other parameters that have been previously associated with neuropsychological performance, thus a potential confounding seems unlikely.
Conclusion and Outlook
Differences in memory performance, decline of cognitive functioning as well as the vulnerability of the hippocampus increase with age and is in part modulated by genetic background (Craik and Rose, 2012; Raz et al, 2014). Thus, to elucidate the mechanisms underlying the association of KIBRA-SNP rs17070145 with brain function and structure, we here for the first time assessed behavioral, microstructural and functional MRI data in a large cohort of healthy older adults, well characterized for potential confounding factors. We observed a trend for better memory performance in KIBRA T-allele carriers compared with non-T-allele carriers. In addition, T-allele carriers showed significantly larger total hippocampal volume and selective volumetric and microstructural improvements in the CA2/3 and CA4/DG hippocampal subfields, known to be involved in memory formation. In addition, resting-state FC analyses showed a larger de-differentiation of the hippocampus network in KIBRA non-T-allele carriers, which might display a compensatory mechanism. Although the underlying mechanisms are still unclear, our findings support and extend existing evidence for a beneficial role of the KIBRA T-allele on hippocampal structural and functional plasticity, which might eventually translate into better memory performance. Future studies are needed to replicate these findings, particularly with regard to differences in hippocampal microstructure and FC, and to further identify the effects of the KIBRA polymorphism on protein expression in the hippocampus subfields, for example, using in vitro assays in postmortem brain slices (Hock et al, 2000).
In sum, our study further strengthens the hypothesis that the common KIBRA-SNP rs17070145 has a distinct role for structural plasticity in memory-related brain areas in older age. Thus, the findings contribute to a better understanding of mechanisms underlying interindividual differences in cognitive function during aging, and may help to develop new preventive and therapeutic strategies against age-associated cognitive decline.
Funding and disclosure
This work was supported by grants from the Deutsche Forschungsgemeinschaft (Fl 379-8/1, Fl 379-10/1; Fl 379-11/1, and DFG-Exc 257), the Else-Kröner Fresenius Stiftung (2009-141; 2011-119), and the Bundesministerium für Bildung und Forschung (FKZ 0315673A, 01EO0801, 01 GY 1144). AF received consulting fees from Schwabe, honoraria for oral presentations from Novartis, Böhringer-Ingelheim and Souvenaid. The remaining authors declare no conflict of interest.
References
Almeida OP, Schwab SG, Lautenschlager NT, Morar B, Greenop KR, Flicker L et al (2008). KIBRA genetic polymorphism influences episodic memory in later life, but does not increase the risk of mild cognitive impairment. J Cell Mol Med 12: 1672–1676.
Andrews-Hanna JR, Reidler JS, Sepulcre J, Poulin R, Buckner RL (2010). Functional-anatomic fractionation of the brain's default network. Neuron 65: 550–562.
Andrews-Hanna JR, Snyder AZ, Vincent JL, Lustig C, Head D, Raichle ME et al (2007). Disruption of large-scale brain systems in advanced aging. Neuron 56: 924–935.
Antonenko D, Floel A (2014). Healthy aging by staying selectively connected: a mini-review. Gerontology 60: 3–9.
Bakker A, Kirwan CB, Miller M, Stark CE (2008). Pattern separation in the human hippocampal CA3 and dentate gyrus. Science 319: 1640–1642.
Beaulieu C (2002). The basis of anisotropic water diffusion in the nervous system—a technical review. NMR Biomed 15: 435–455.
Bender AR, Daugherty AM, Raz N (2013). Vascular risk moderates associations between hippocampal subfield volumes and memory. J Cogn Neurosci 25: 1851–1862.
Bruel-Jungerman E, Rampon C, Laroche S (2007). Adult hippocampal neurogenesis, synaptic plasticity and memory: facts and hypotheses. Rev Neurosci 18: 93–114.
Burgess JD, Pedraza O, Graff-Radford NR, Hirpa M, Zou F, Miles R et al (2011). Association of common KIBRA variants with episodic memory and AD risk. Neurobiol Aging 32: 557 e551–557 e559.
Corder EH, Saunders AM, Strittmatter WJ, Schmechel DE, Gaskell PC, Small GW et al (1993). Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer's disease in late onset families. Science 261: 921–923.
Corneveaux JJ, Liang WS, Reiman EM, Webster JA, Myers AJ, Zismann VL et al (2010). Evidence for an association between KIBRA and late-onset Alzheimer's disease. Neurobiol Aging 31: 901–909.
Craik FI, Rose NS (2012). Memory encoding and aging: a neurocognitive perspective. Neurosci Biobehav Rev 36: 1729–1739.
Deary IJ, Wright AF, Harris SE, Whalley LJ, Starr JM (2004). Searching for genetic influences on normal cognitive ageing. Trends Cogn Sci 8: 178–184.
de Flores RI, La Joie R, Landeau B, Perrotin A, Mézenge F, de La Sayette V et al (2015). Effects of age and Alzheimer's disease on hippocampal subfields: comparison between manual and FreeSurfer volumetry. Hum Brain Mapp 36: 463–474.
den Heijer T, der Lijn F, Vernooij MW, de Groot M, Koudstaal PJ, der Lugt A et al (2012). Structural and diffusion MRI measures of the hippocampus and memory performance. Neuroimage 63: 1782–1789.
Draganski B, May A (2008). Training-induced structural changes in the adult human brain. Behav Brain Res 192: 137–142.
Egan MF, Kojima M, Callicott JH, Goldberg TE, Kolachana BS, Bertolino A et al (2003). The BDNF val66met polymorphism affects activity-dependent secretion of BDNF and human memory and hippocampal function. Cell 112: 257–269.
Fischl B, Salat DH, Busa E, Albert M, Dieterich M, Haselgrove C et al (2002). Whole brain segmentation: automated labeling of neuroanatomical structures in the human brain. Neuron 33: 341–355.
Folstein MF, Folstein SE, McHugh PR (1975). "Mini-mental state". A practical method for grading the cognitive state of patients for the clinician. J Psychiatr Res 12: 189–198.
Hayashi N, Kazui H, Kamino K, Tokunaga H, Takaya M, Yokokoji M et al (2010). KIBRA genetic polymorphism influences episodic memory in Alzheimer's disease, but does not show association with disease in a Japanese cohort. Dement Geriatr Cogn Disord 30: 302–308.
Hock C, Heese K, Hulette C, Rosenberg C, Otten U (2000). Region-specific neurotrophin imbalances in Alzheimer disease: decreased levels of brain-derived neurotrophic factor and increased levels of nerve growth factor in hippocampus and cortical areas. Arch Neurol 57: 846–851.
Johannsen S, Duning K, Pavenstadt H, Kremerskothen J, Boeckers TM (2008). Temporal-spatial expression and novel biochemical properties of the memory-related protein KIBRA. Neuroscience 155: 1165–1173.
Kantarci K, Petersen RC, Boeve BF, Knopman DS, Weigand SD, O'Brien PC et al (2005). DWI predicts future progression to Alzheimer disease in amnestic mild cognitive impairment. Neurology 64: 902–904.
Kauppi K, Nilsson LG, Adolfsson R, Eriksson E, Nyberg L (2011). KIBRA polymorphism is related to enhanced memory and elevated hippocampal processing. J Neurosci 31: 14218–14222.
Kerti L, Witte AV, Winkler A, Grittner U, Rujescu D, Floel A (2013). Higher glucose levels associated with lower memory and reduced hippocampal microstructure. Neurology 81: 1746–1752.
Kuhner C, Burger C, Keller F, Hautzinger M (2007). Reliability and validity of the revised Beck Depression Inventory (BDI-II). Results from German samples. Nervenarzt 78: 651–656.
Laux L, Glanzmann P, Schaffner P, Spielberger C (1981) Das State-Trait-Angstinventar. Theoretische Grundlagen und Handanweisung. Beltz Test GmbH: Weinheim.
Lezak MD (2004) Neuropsychological Assessment, 4th edn. Oxford University Press: New York, Oxford.
Lindenberger U, Nagel IE, Chicherio C, Li SC, Heekeren HR, Backman L (2008). Age-related decline in brain resources modulates genetic effects on cognitive functioning. Front Neurosci 2: 234–244.
Milnik A, Heck A, Vogler C, Heinze HJ, de Quervain DJ, Papassotiropoulos A (2012). Association of KIBRA with episodic and working memory: a meta-analysis. Am J Med Genet B Neuropsychiatr Genet 159B: 958–969.
Mueller SG, Chao LL, Berman B, Weiner MW (2011). Evidence for functional specialization of hippocampal subfields detected by MR subfield volumetry on high resolution images at 4 T. Neuroimage 56: 851–857.
Muse J, Emery M, Sambataro F, Lemaitre H, Tan HY, Chen Q et al (2013). WWC1 genotype modulates age-related decline in episodic memory function across the adult life span. Biol Psychiatry 75: 693–700.
Muse J, Emery M, Sambataro F, Lemaitre H, Tan HY, Chen Q et al (2014). WWC1 genotype modulates age-related decline in episodic memory function across the adult life span. Biol Psychiatry 75: 693–700.
Nacmias B, Bessi V, Bagnoli S, Tedde A, Cellini E, Piccini C et al (2008). KIBRA gene variants are associated with episodic memory performance in subjective memory complaints. Neurosci Lett 436: 145–147.
Need AC, Attix DK, McEvoy JM, Cirulli ET, Linney KN, Wagoner AP et al (2008). Failure to replicate effect of Kibra on human memory in two large cohorts of European origin. Am J Med Genet B Neuropsychiatr Genet 147B: 667–668.
O'Dwyer L, Lamberton F, Matura S, Tanner C, Scheibe M, Miller J et al (2012). Reduced hippocampal volume in healthy young ApoE4 carriers: an MRI study. PLoS One 7: e48895.
Oldfield RC (1971). The assessment and analysis of handedness: the Edinburgh inventory. Neuropsychologia 9: 97–113.
Palombo DJ, Amaral RS, Olsen RK, Muller DJ, Todd RM, Anderson AK et al (2013). KIBRA polymorphism is associated with individual differences in hippocampal subregions: evidence from anatomical segmentation using high-resolution MRI. J Neurosci 33: 13088–13093.
Papassotiropoulos A, Stephan DA, Huentelman MJ, Hoerndli FJ, Craig DW, Pearson JV et al (2006). Common KIBRA alleles are associated with human memory performance. Science 314: 475–478.
Preuschhof C, Heekeren HR, Li SC, Sander T, Lindenberger U, Backman L (2010). KIBRA and CLSTN2 polymorphisms exert interactive effects on human episodic memory. Neuropsychologia 48: 402–408.
Qi Z, Wu X, Wang Z, Zhang N, Dong H, Yao L et al (2010). Impairment and compensation coexist in amnestic MCI default mode network. Neuroimage 50: 48–55.
Raz N, Amedi A, Zohary E (2005). V1 activation in congenitally blind humans is associated with episodic retrieval. Cereb Cortex 15: 1459–1468.
Raz N, Daugherty AM, Bender AR, Dahle CL, Land S (2014). Volume of the hippocampal subfields in healthy adults: differential associations with age and a pro-inflammatory genetic variant. Brain Struct Funct. (e-pub ahead of print).
Rombouts SA, Stam CJ, Kuijer JP, Scheltens P, Barkhof F (2003). Identifying confounds to increase specificity during a "no task condition". Evidence for hippocampal connectivity using fMRI. Neuroimage 20: 1236–1245.
Schaper K, Kolsch H, Popp J, Wagner M, Jessen F (2008). KIBRA gene variants are associated with episodic memory in healthy elderly. Neurobiol Aging 29: 1123–1125.
Schneider A, Huentelman MJ, Kremerskothen J, Duning K, Spoelgen R, Nikolich K (2010). KIBRA: a new gateway to learning and memory? Front Aging Neurosci 2: 4.
Schuck NW, Doeller CF, Schjeide BM, Schroder J, Frensch PA, Bertram L et al (2013). Aging and KIBRA/WWC1 genotype affect spatial memory processes in a virtual navigation task. Hippocampus 23: 919–930.
Squire LR (2004). Memory systems of the brain: a brief history and current perspective. Neurobiol Learn Mem 82: 171–177.
Thomas C, Baker CI (2012). Teaching an adult brain new tricks: a critical review of evidence for training-dependent structural plasticity in humans. Neuroimage 73: 225–236.
Van Leemput K, Bakkour A, Benner T, Wiggins G, Wald LL, Augustinack J et al (2009). Automated segmentation of hippocampal subfields from ultra-high resolution in vivo MRI. Hippocampus 19: 549–557.
Wang D, Liu B, Qin W, Wang J, Zhang Y, Jiang T et al (2013a). KIBRA gene variants are associated with synchronization within the default-mode and executive control networks. Neuroimage 69: 213–222.
Wang HF, Tan L, Yu JT, Ma XY, Liu QY, Wang W (2013b). Age-dependent association of KIBRA gene polymorphism with Alzheimer's disease in Han Chinese. Mol Biol Rep 40: 7077–7082.
Wersching H, Guske K, Hasenkamp S, Hagedorn C, Schiwek S, Jansen S et al (2011). Impact of common KIBRA allele on human cognitive functions. Neuropsychopharmacology 36: 1296–1304.
Witte AV, Floel A (2012). Effects of COMT polymorphisms on brain function and behavior in health and disease. Brain Res Bull 88: 418–428.
Witte AV, Kerti L, Margulies DS, Floel A (2014). Effects of resveratrol on memory performance, hippocampal functional connectivity, and glucose metabolism in healthy older adults. J Neurosci 34: 7862–7870.
Yasuda Y, Hashimoto R, Ohi K, Fukumoto M, Takamura H, Iike N et al (2010). Association study of KIBRA gene with memory performance in a Japanese population. World J Biol Psychiatry 11: 852–857.
Zeineh MM, Engel SA, Thompson PM, Bookheimer SY (2003). Dynamics of the hippocampus during encoding and retrieval of face-name pairs. Science 299: 577–580.
Acknowledgements
We specially thank Angela Winkler, Henrike Herrmannstädter and Johanna Richels for help with data acquisition.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Neuropsychopharmacology website
Supplementary information
Rights and permissions
About this article
Cite this article
Witte, A., Köbe, T., Kerti, L. et al. Impact of KIBRA Polymorphism on Memory Function and the Hippocampus in Older Adults. Neuropsychopharmacol 41, 781–790 (2016). https://doi.org/10.1038/npp.2015.203
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2015.203
This article is cited by
-
KIBRA single nucleotide polymorphism is associated with hippocampal subfield volumes and cognition across development
Brain Structure and Function (2023)
-
Brain microstructural alterations in the left precuneus mediate the association between KIBRA polymorphism and working memory in healthy adults: a diffusion kurtosis imaging study
Brain Imaging and Behavior (2022)
-
Cerebral Small Vessel Disease Influences Hippocampal Subfield Atrophy in Mild Cognitive Impairment
Translational Stroke Research (2021)
-
KIBRA polymorphism modulates gray matter volume to influence cognitive ability in the elderly
Brain Imaging and Behavior (2020)
-
KIBRA T allele influences memory performance and progression of cognitive decline: a 7-year follow-up study in subjective cognitive decline and mild cognitive impairment
Neurological Sciences (2019)