Abstract
The discovery of functional cannabinoid receptors 2 (CB2Rs) in brain suggests a potential new therapeutic target for neurological and psychiatric disorders. However, recent findings in experimental animals appear controversial. Here we report that there are significant species differences in CB2R mRNA splicing and expression, protein sequences, and receptor responses to CB2R ligands in mice and rats. Systemic administration of JWH133, a highly selective CB2R agonist, significantly and dose-dependently inhibited intravenous cocaine self-administration under a fixed ratio (FR) schedule of reinforcement in mice, but not in rats. However, under a progressive ratio (PR) schedule of reinforcement, JWH133 significantly increased breakpoint for cocaine self-administration in rats, but decreased it in mice. To explore the possible reasons for these conflicting findings, we examined CB2R gene expression and receptor structure in the brain. We found novel rat-specific CB2C and CB2D mRNA isoforms in addition to CB2A and CB2B mRNA isoforms. In situ hybridization RNAscope assays found higher levels of CB2R mRNA in different brain regions and cell types in mice than in rats. By comparing CB2R-encoding regions, we observed a premature stop codon in the mouse CB2R gene that truncated 13 amino-acid residues including a functional autophosphorylation site in the intracellular C-terminus. These findings suggest that species differences in the splicing and expression of CB2R genes and receptor structures may in part explain the different effects of CB2R-selective ligands on cocaine self-administration in mice and rats.
Similar content being viewed by others
INTRODUCTION
It was previously reported that the cannabinoid receptor 1 (CB1R) is predominantly expressed in brain, whereas the cannabinoid receptor 2 (CB2R) is predominantly expressed in the peripheral immune system (Matsuda, 1997). Therefore, the majority of CB2R-related studies have focused on peripheral tissues. This view of central CB1R and peripheral CB2R cannabinoid systems has been challenged recently by growing evidence demonstrating functional CB2R expression in brain (Atwood and Mackie, 2010; Mechoulam and Parker, 2013; Onaivi et al, 2012). Furthermore, the findings of neuronal CB2Rs in the brain and their postsynaptic localization (Brusco et al, 2008a; Gong et al, 2006; Vinckenbosch et al, 2006) suggest an important role of the CB2R in neuronal synaptic transmission and CNS disorders such as drug abuse, schizophrenia, and stroke (Onaivi et al, 2008; Onaivi et al, 2012).
However, neuronal localization of CB2Rs in the brain has been controversial (Atwood and Mackie, 2010; Van Sickle et al, 2005) because of concerns regarding antibody specificities. Several independent groups have demonstrated a functional role for brain CB2Rs in terms of genetic association with psychiatric diseases (Ishiguro et al, 2010a, 2010b), cellular distributions and neuronal localizations (Lanciego et al, 2011; Suárez et al, 2009; Van Sickle et al, 2005), electrophysiological effects (den Boon et al, 2012; Morgan et al, 2009), and behavioral pharmacological effects using CB2R transgenic mice (Callén et al, 2012; Garcia-Gutiérrez and Manzanares, 2011; Garcia-Gutiérrez et al, 2013; Navarrete et al, 2013; Xi et al, 2011). Strikingly, growing evidence demonstrates that brain CB2Rs may be involved in drug reward and addiction. Xi et al (2011) recently reported that systemic or intracranial local administration of JWH133, a selective CB2R agonist, into the nucleus accumbens significantly inhibits intravenous (i.v.) cocaine self-administration in wild-type (WT) and CB1R-knockout (CB1-KO) mice, but not in CB2-KO mice. Consistent with these findings, systemic administration of the CB2R agonist O-1966 inhibits cocaine-induced conditioned place preference (CPP) in WT mice, but not in CB2R-KO mice (Ignatowska-Jankowska et al, 2013). Transgenic mice with overexpression of CB2R in the brain show decreased cocaine self-administration and cocaine-enhanced locomotion (Aracil-Fernández et al, 2012). In addition, systemic administration of the CB2R agonist β-caryophyllene reduced voluntary alcohol intake, alcohol-induced CPP, and locomotor sensitization in mice (Al Mansouri et al, 2014). In contrast to the above findings, CB2Rs appear to play an opposite role in mediating nicotine’s action. It was reported that genetic deletion of CB2Rs in mice attenuated nicotine self-administration (Navarrete et al, 2013) and abolished nicotine-induced CPP (Ignatowska-Jankowska et al, 2013). Congruently, pharmacological blockade of CB2Rs by SR 144528 attenuates nicotine-induced CPP in WT mice (Ignatowska-Jankowska et al, 2013).
In contrast to the above findings in mice, the results with CB2R ligands in rats appear controversial. Blanco-Calvo et al (2014) recently reported that pharmacological blockade of CB1Rs (by SR144176A) or CB2Rs (by AM630) prevents both cocaine-induced conditioned locomotion and cocaine-induced reduction of cell proliferation in the hippocampus of rats. Adamczyk et al (2012) reported that systemic administration of the CB2R antagonist SR144528 had no effect on i.v. cocaine self-administration, but attenuated cocaine-induced reinstatement of drug-seeking behavior in rats. Furthermore, Gamaleddin et al (2012b) reported that systemic administration of the CB2R agonist AM1241 or the antagonist AM630 does not alter nicotine self-administration or nicotine- or cue-induced reinstatement of nicotine-seeking behavior. The reasons underlying such conflicting findings are unclear. The simplest explanation may relate to different drugs of abuse (cocaine versus nicotine), different CB2R agonists (JWH133 versus AM1241 or O-1966) or antagonists (AM630 versus SR144528), different drug doses, different species of animals, and different measures in different animal models.
In this study, we explored whether the above-noted different effects in addiction-related models may be related to species differences in CB2R gene and receptor expression. To this end, we studied different effects of the CB2R agonist JWH133 on cocaine self-administration in rats and mice. We then carried out a series of experiments to study and compare CB2R gene structure and alternative splicing, CB2R mRNA expression, and CB2R amino-acid sequences and their 3D structures in mice and rats.
MATERIALS AND METHODS
Animal Subjects
Male Long–Evans rats (Charles River, Raleigh, NC) and WT and CB2-KO mice with C57BL/6J genetic backgrounds (Buckley et al, 2000) were used in cocaine self-administration experiments. WT and CB2-KO mice were bred within the Transgenic Animal Breeding Facility of the National Institute on Drug Abuse (NIDA). They were housed in a fully accredited animal facility and were maintained on a reversed 12 h light/dark cycle (lights on at 1900 h and lights off at 0700 h) with food and water available ad libitum in the home cage. The experimental procedures followed the Guide for the Care and Use of Laboratory Animals (1996) and were approved by the NIDA Animal Care and Use Committee.
Intravenous Cocaine Self-Administration
Surgery
Animals were prepared for i.v. cocaine self-administration by surgical catheterization of the right external jugular vein. The jugular catheters were constructed of microrenathane (Braintree Scientific, Braintree, MA), and catheterization was performed under sodium pentobarbital anesthesia using standard aseptic surgical techniques as described previously (Xi et al, 2011; Song et al, 2012; Xi et al, 2005). The self-administration cannulae were fixed to the skull with 4 stainless steel jeweler’s screws (Small Parts, Miami Lakes, FL) and dental acrylic cement. During experimental sessions, each catheter was connected to an injection pump via tubing encased in a protective metal spring from the head-mounted connector to the top of the experimental chamber. To help prevent clogging, the catheters were flushed daily with a gentamicin–heparin saline solution (0.1 mg/ml gentamicin and 30 IU/ml heparin; ICN Biochemicals, Cleveland, OH).
Self-administration apparatus
Intravenous self-administration experiments were conducted in operant response test chambers (Model ENV-008CT for rats; Model ENV-307A for mice) from Med Associates (Georgia, VT). Each test chamber had two levers: one active and one inactive. Depression of the active lever activated the infusion pump; depression of the inactive lever was counted but had no consequence. A cue light and a speaker were located 12 cm above the active lever. The house light was turned on at the start of each 3 h test session. Scheduling of experimental events and data collection was accomplished using Med Associates software.
Self-administration procedure
After recovery from surgery, each rat or mouse was placed into a test chamber (day time—dark phase) and allowed to lever-press for i.v. cocaine (1 mg/kg/infusion) delivered in 0.08 ml or 0.01 ml over 4.6 s in rats and mice, respectively, on a fixed ratio 1 (FR1) reinforcement schedule. Each cocaine infusion was associated with presentation of a stimulus light and tone. During the 4.6 s infusion time, additional responses on the active lever were recorded but did not lead to additional infusions. Each session lasted 3 h. FR1 reinforcement was used for 3–5 days until stable cocaine self-administration was established: a minimum of 20 presses on the active lever per test session and stability criteria of <15% variability in interresponse interval, <15% variability in number of infusions taken, and <15% variability in number of presses on the active lever for at least 3 consecutive days. Subjects were then allowed to continue cocaine self-administration (0.5 mg/kg/infusion) under FR2 reinforcement. This dose of cocaine was chosen based on previous studies showing that it lies within the middle range of the descending limb of the cocaine dose–-response self-administration curve, where reliable dose-dependent effects are observed (Xi et al, 2005). To avoid cocaine overdose, each animal was limited to a maximum of 50 cocaine injections per 3 h session.
Effects of JWH133 on cocaine self-administration
We first evaluated the effects of systemic administration of JWH133 (10 and 20 mg/kg, i.p., 30 min before testing) or vehicle (Tocrisolve-100) on FR2 cocaine self-administration in one group of rats and one group of mice (within-subjects design, n=6 rats/group and n=9 mice/group). The doses of JWH133 were based on our previous work (Xi et al, 2011).
Cocaine self-administration under progressive ratio (PR) reinforcement
Initial cocaine self-administration under FR1 and FR2 reinforcement was identical to that outlined above. After stable cocaine self-administration under FR2 reinforcement was established, the subjects were switched to cocaine (0.5 mg/kg/injection) self-administration under a PR schedule, during which the work requirement (lever presses) needed to receive a single i.v. cocaine infusion was progressively raised within each test session (Xi et al, 2005) according to the following PR series: 1, 2, 4, 6, 9, 12, 15, 20, 25, 32, 40, 50, 62, 77, 95, 118, 145, 178, 219, 268, 328, 402, 492, and 603 until breakpoint (BP) was reached. BP was defined as the maximal work load (lever presses) completed for a cocaine infusion before a 1-h period during which no infusion was obtained by the animal. The PR schedule is computer programmed to progress to a maximum of 603 and the average BP was ∼150. Animals were allowed to continue daily sessions of cocaine self-administration under PR reinforcement conditions until day-to-day variability in BP was within 1–2 ratio increments for three consecutive days. Once a stable BP was established, subjects were assigned to three subgroups to determine the effects of three different doses of JWH133 (0, 10, and 20 mg/kg, i.p.) on PR BP for cocaine self-administration. Additional groups of animals were used to observe the effects of intranasal administration of JWH133 (0, 10, 25, or 50 μg/side) on PR cocaine self-administration. Intranasal drug administration was performed under inhalant isoflurane anesthesia using the Fluovac System (Harvard Apparatus, Holliston, MA). We chose between-subjects design for this experiment because it is relatively difficult to re-establish stable BP cocaine self-administration after each test under PR reinforcement as compared with cocaine self-administration under FR2 conditions.
Oral Sucrose Self-Administration
The procedures for oral sucrose self-administration testing were identical to the procedures used for cocaine self-administration, except for the following: (1) no surgery was carried out in this experiment; and (2) active lever presses led to delivery of 0.1 ml of 5% sucrose solution into a liquid food tray on the operant chamber wall. The effects of JWH133 (same doses as in previous experiments) on oral sucrose self-administration maintained under FR2 reinforcement were evaluated.
RT-PCR Analysis of CB2R Isoforms
Male Sprague–Dawley rats and WT and CB2-KO mice on C57BL/6J genetic background (Charles River) were decapitated, and the brains were rapidly removed and frozen in −50 °C isopentane solution, and then stored at −80 °C freezer until for assays. Punches (12 gauge needle) of brain regions were taken from 1 mm coronal sections cut using a cryostat at −20 °C. Total rodent RNAs were isolated using the TRIzol Reagent. RNA integrity numbers (RINs) were >8 measured by Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA) for rat and mouse RNAs. Single-strand cDNAs were synthesized using the Superscript III first-strand cDNA synthesis kit according to the manufacturer’s protocols (Invitrogen, Life Technologies, Carlsbad, CA). TaqMan probes (Figure 1 and Table 1) were designed using Primer Express 3.0 (Applied Biosystems, Life Technologies, Carlsbad, CA) at the splicing junctions of the different rat and mouse CB2R isoforms. The endogenous control TaqMan probes were rat VIC-labeled GAPDH (ABI Cat. no. 4352338E) and mouse rat VIC-labeled β-actin (ABI Cat. no. 4352341E). Duplex PCR assays containing both the target and endogenous control TaqMan probes were carried out with Advanced TaqMan Fast Universal PCR Master Mix in a 7500 Fast TaqMan instrument using a default thermocycling program. For rCB2D isoform expression analysis, SYBR green assay was carried out with primers designed across the intraexonic splicing site that produce PCR fragments with 223 bp representing spliced rCB2D isoform. Quantification of real-time PCR was carried out as described previously with duplexed PCR and technical duplicates (Liu et al, 2009). Regular PCR was carried out by 9700 ThermoCycler and the amplification products were analyzed in 1% and 3% agarose gel stained with ethidium bromide. Various tissues of WT and a C-terminal-deleted partial knockout strain (Buckley et al, 2000) were used for PCR amplification. The amplified PCR bands were cut from the agarose gel and DNA extracted by Qiagen gel extraction kit. The purified PCR fragments were cloned into TA cloning vector (PCR4 Topo, Invitrogen Life Technology) and sequenced by Sanger sequencing method (SeqWright Genomic Services, Houston, TX). RNA-Seq data in human, rhesus macaque, mouse, and rat were downloaded and processed to verify the splice structure of CB2R isoforms, according to standardized pipelines as reported previously (Zhang et al, 2014a).
Differential effects of JWH133 on cocaine self-administration in rats and mice. (a) Systemic administration (i.p.) of JWH133 significantly decreased cocaine self-administration under fixed ratio (FR2) reinforcement in mice, but not in rats, as assessed by mean numbers of cocaine infusions per hour. (b) Systemic administration of JWH133 significantly increased interinfusion intervals in mice, but not in rats. (c) Systemic administration of JWH133 significantly increased PR BP for cocaine self-administration in rats, but decreased PR BP in mice. (d) Intranasal (i.n.) administration of JWH133 (10, 25, or 50 μg/10 μl/side) produced biphasic effects—low doses increased whereas high doses decreased PR BP in rats. (e) Systemic administration of JWH133 significantly inhibited oral sucrose self-administration in WT mice, but not in rats or CB2-KO mice. *P<0.05, compared with vehicle control group.
RNAscope In Situ Hybridization (ISH)
Rat and mouse CB2R RNAscope probes in C1 channel were custom made by Advanced Cell Diagnostics (ACD, Hayward, CA). Customer probes using 20 ZZ pairs for each target were synthesized according to 3′−untranslated regions (UTRs) of rat rCB2R (NM_001164143.2; 1935–2843) or mouse mCB2R (NM_009924.3; 1877–2820). Catalog probes of mouse tyrosine hydroxylase (TH) and rat dopamine transporter (DAT) were ordered from ACD.
Rat or mouse brains were rapidly frozen in 100 ml −50 °C isopentane. The frozen brains were sealed in zipper plastic bag, wrapped in labeled aluminum foil, and stored at −80 °C. Before ISH, brains were placed in a cryostat (CM 3050S) at −20 °C for 2 h to allow for temperature equilibration and then sliced and trimmed to the desired coronal plane. The coronal sections were cut at 12 μm thickness using anti-rolling plate to flatten the section. The objective temperature was set to −18 °C and the chamber temperature to −19 °C. Supper Frost Plus slides (Fisher, Cat. no. 12-550-15) were used to pick up each thin section by flipping the slide upside down and allowing each brain section to stick to the slides. The sections were spread out evenly using gloved finger under each section. The slides were left at −20 °C for 10 min and transferred to −80 °C for storage. The fixation, protease pretreatment, probe hybridization, preamplification, amplification, and fluorescent labeling steps were carried out according to User Manual for Fresh Frozen Tissue (ACD).
Wide-field fluorescent images of ventral tegmental area (VTA), nucleus accumbens (NAC), prefrontal cortex (PFC), and dorsal striatum (DST) were captured using a QimagingExi Aqua camera (Biovision) attached to a Zeiss AXIO Imager M2 microscope using a × 40 objective (Zeiss PLAN-APOCHROMAT, NA=1.3) with oil immersion. Images were deconvoluted with Huygens software (v3.7, Scientific Volume Imaging), pseudo-colored (CB2R green, TH and DAT red, and DAPI blue), and color joined using iVision software (Liu et al, 2014). Image Processing and Analysis by Java (ImageJ, NIH) software was used to quantify mRNA signals in each individual cell.
Bioinformatics
Rat and mouse CB2R genomic structures were analyzed by NCBI Map Viewer (http://www.ncbi.nlm.nih.gov/mapview/) and UCSC Genome Browser (http://genome.ucsc.edu/). The exon and intron junctions were defined by alignment of exons with genomic sequences using Sequencher 5.0 software (Gene Codes Corporation, Ann Arbor, MI). The NCBI Blastn suite was used to align nucleotide sequences of rat and mouse CB2R EST and PCR fragment sequences. CB2R peptide sequences were aligned using CLUSTALW software (http://www.ebi.ac.uk/Tools/msa/clustalw2/). The CB2R functional sites were predicted by The Eukaryotic Linear Motif database (http://elm.eu.org/), N-glycosylation was predicted by NetNGlyc 1.0 (http://www.cbs.dtu.dk/services/NetNGlyc/), and phosphorylation sites were predicted by ExPaSyNetPhos 2.0 (http://www.cbs.dtu.dk/services/NetPhos/).
The CB2R peptide sequences of rat and mouse were submitted to the automated comparative protein modeling server Swiss-Model (Arnold et al, 2006; Guex and Peitsch, 1997; Schwede et al, 2003) (http://swissmodel.expasy.org/) to build up three-dimensional (3D) structures using the crystal structure of turkey β-adrenergic receptor (PDB: 4ajm) as a modeling template. The 3D figures were created with PyMOL (http://www.pymol.org/).
GenBank accession numbers were submitted as follows: JN420349 for rat CB2C and JX494784 for rat CB2D.
Data Analysis
Data are presented as means±SEM of cocaine infusions per hour, cocaine infusion intervals, and BPs in the absence or presence of JWH133 pretreatment. Two-way ANOVA (Graphpad Software v6.0) for repeated measures over JWH133 doses was used to determine the significance of JWH133 effects. Nonrepeated measures ANOVA over rats and mice were used to determine significances of the JWH133 effects on cocaine or sucrose self-administration.
Two brain sections of different brain regions from each of two mice or two rats were used for quantification of CB2R mRNA in 6–8 cells per region. For VTA sections, mTH- or rDAT-positive and -negative cells were counted separately in order to quantify mCB2R and rCB2R in mice and rats, respectively. Pixels were used to quantify CB2R mRNA in each individual cell and intensities were used to quantify CB2R in dopamine and nondopamine cells of VTA. Regular two-way ANOVA was used for statistical analysis of species differences and mixed two-way ANOVA was used for analysis using dopamine and nondopamine cells as the within-subjects variable and mice and rats as the between-subjects variable. Multiple comparisons were corrected by the Sidak method included in the software. The corrected significance level was p<0.05.
RESULTS
Differential Effects of JWH133 on Cocaine Self-Administration in Mice and Rats
Figure 1 shows the effects of JWH133 on cocaine and sucrose self-administration under fixed ratio 2 (FR2) or PR reinforcement in both rats and mice. Systemic administration of JWH133 (10 and 20 mg/kg) significantly inhibited i.v. cocaine self-administration under FR2 reinforcement, as indicated by decreased number of cocaine infusions per hour and increased intercocaine infusion intervals, in mice, but not in rats. Although two-way ANOVA for repeated measures for the cocaine infusion rate data (Figure 1a) did not reveal a statistically significant JWH133 treatment main effect (F2, 26=5.53, p>0.05), a statistically significant species main effect (F1, 13=3.58, p<0.05) and a JWH133 × species interaction (F2, 26=5.48, p=0.01) were revealed. Two-way ANOVA for the cocaine infusion interval data (Figure 1b) revealed a significant JWH133 treatment main effect (F2, 26=5.89, p<0.01), species main effect (F1, 13=14.91, p<0.01), and a JWH133 × species interaction (F2, 26=5.47, p=0.01). Figure 1c shows the effects of JWH133 on BP for i.v. cocaine self-administration in rats and mice, illustrating that JWH133, at the same doses (10 and 20 mg/kg, i.p.), significantly increased PR BP for cocaine self-administration in rats, but decreased it in mice. Again, although two-way ANOVA did not reveal a statistically significant JWH133 treatment main effect (F2, 52=1.15, p>0.05), a statistically significant species main effect (F1, 52=43.49, p<0.001) and a significant JWH133 × species interaction (F2, 52=11.6, p<0.01) were revealed.
To determine the mechanisms by which JWH133 produced opposite effects on PR cocaine self-administration in rats and mice, we hypothesized that an increase in PR BP for cocaine self-administration in rats may be a compensatory response to a partial reduction in cocaine’s rewarding efficacy and, therefore, a high dose of JWH133 may be required to produce an inhibitory effect similar to that seen in mice. However, testing higher systemic doses of JWH133 (eg, >20 mg/kg) is problematic because of unwanted side effects, particularly locomotor inhibition. Therefore, we used intranasal drug delivery route by which many drugs may enter brain directly (Costantino et al, 2007; Illum et al, 2002). Figure 1d shows that intranasal administration of JWH133 produced biphasic effects—lower doses (10 and 25 μg/side) increased whereas a higher dose (50 μg/side) decreased BP for cocaine self-administration in rats (F3, 18=8.89, p<0.001). However, in WT mice, intranasal administration of JWH133 (10 and 20 μg/side) produced a monophasic inhibition of cocaine self-administration (F2, 11=4.59, p<0.05).
To determine whether systemic administration of JWH133 also alters other reward-reinforced behavior, we observed the effects of JWH133 on oral sucrose self-administration in rats and mice. We found that JWH133 significantly inhibited sucrose self-administration in WT mice (Figure 1, F2, 18=13.09, p<0.001), but not in rats or in CB2-KO mice (Figure 1e).
Different CB2R mRNA Isoforms Found in Rats and Mice
Figure 2 shows rat CB2 (rCB2R) and mouse CB2 (mCB2R) receptor gene and transcript (mRNA) structures, illustrating 3 exons spanning ∼25 kb, multiple CB2R mRNA isoforms, and the splicing junction locations of probes that were used to detect different CB2 mRNA isoforms. Using RT-PCR and DNA fragment agarose gel electrophoresis analysis, we detected CB2A and CB2B isoforms in both rats and mice, and CB2C and CB2D only in rats, but not in mice (Figure 3a–c). In this study, we used primers that targeted the conjunction region of exons 1–3 (94 bp) or exons 2 and 3 (85 bp) to detect CB2A and CB2B isoforms, respectively, in rat and mouse brain (Figure 3a and b). Unexpectedly, the CB2A primers also detected an additional 189 bp RT-PCR fragment in rats, not in mice, suggesting a novel rCB2C-specific isoform (Figure 3a). The sequence analysis of the PCR fragment of rCB2C indicated that it was generated by exons 1–3 via interexonic tripartite exon splicing events. The rCB2D was represented by a 223-bp RT-PCR fragment that was detected in amygdala, spleen, liver, and ovary tissues (Figure 3c). Sequence analysis of the PCR fragment of rCB2D indicated that it was generated by intraexonic splicing of the third exon in the 3′-UTR (FJ496961: 1465–2252) with 786 bp deletion. Mouse brain and peripheral tissues neither expressed the corresponding 189 bp rCB2C nor 233 bp rCB2D PCR fragments (Figure 3a and c). Analysis of the RNA-Seq data in spleen also suggested the presence of rCB2C and the absence of the corresponding mouse isoform (Zhang et al, 2013). Figure 3d shows that mCB2 mRNA signal was detectable only in WT mice, but not in CB2-KO mice (Figure 3dB and C), when using a probe that targeted the gene-deleted region in CB2-KO mice (Figure 3dA). However, the probe targeted at CB2A or CB2B showed enhanced expression in various tissues of CB2-KO mice, possibly because of compensatory effects after the partial C-terminal deletion in the CB2-KO mouse strain (Buckley et al, 2000; Liu et al, 2009).
(a) Mouse CB2 (mCB2R, Cnr2, 4D3) and (b) rat CB2 (rCB2R, Cnr2, 5q36) genomic structures, alternatively spliced transcripts, and the locations that each probe targeted to detect brain CB2 mRNA. Gene: open boxes represent exons, horizontal lines introns and black box mini-intron within the last exon. Transcripts: spliced exon numbers are indicated in the exons. The TaqMan probes were designed to fit the junctions of the spliced exons and are represented by horizontal black bars. The RNAscope probes hybridize 3′-UTR regions marked by gray boxes.
Agarose gel analysis results of PCR fragments, illustrating mCB2A, rCB2A, and rCB2C (a), mCB2B and rCB2B (b), and rCB2D (c) isoforms (ie, shown in Figure 2). When using a probe that targeted the gene-deleted region in CB2-KO mice, mCB2 mRNA signal was detectable only in the striatum and spleen of WT mice, but not in CB2-KO mice (d). MW, molecular weight marker; SPL, spleen; ko-CB2, knockout mice; wt, wild type; MID, midbrain; CTX, cortex; CER, cerebellum; HIP, hippocampus; PFC, prefrontal cortex; MID, midbrain; DST, dorsal striatum; NAC, nucleus accumbens; AMG, amygdala; LIV, liver; TES, testis; OVA, ovary; Prefix: m, mouse and r, rat.
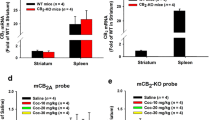
Quantitative PCR assay was carried out to compare expression of CB2R isoforms in brain and peripheral tissues of rats and mice. We used cortex CB2R mRNA level as a reference for brain expression and testis CB2 mRNA level as a reference for peripheral expression because cortical or testis CB2 mRNA levels are relatively the same in rats and mice. We found that the levels of CB2A and CB2B were higher in spleen (∼100–600-fold) than in brain and testis (Figure 4a and b). However, the rat-specific isoform rCB2C mRNA levels are relatively lower in spleen (approximately six fold; Figure 4c) whereas rCB2D (Figure 4d) is predominantly expressed in spleen (∼2000-fold) compared with rCB2A and rCB2B expression.
RT-qPCR comparison of CB2R isoform tissue expression in mice and rats. The white bars represent rCB2 mRNA levels in rat tissues and the gray bars represent mCB2 mRNA levels in mouse tissues. The y axis is fold change in CB2 mRNA, in which rat cortical CB2 mRNA level was used as a reference to quantify the levels in other brain regions, whereas for rat testis CB2 level was used as a reference to quantify CB2 mRNA in peripheral tissues. For rat-specific rCB2C and rCB2D isoforms, the cortical CB2 mRNA level was used as a reference for both brain and peripheral tissues.
RNAscope ISH of CB2R in Rat and Mouse Brains
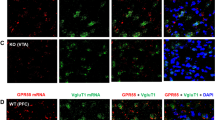
We then further used an ultrasensitive RNAscope ISH method to detect and quantify low densities of CB2R mRNA expression in brain slices of rats and mice, as the traditional ISH method may not be sensitive enough to detect the low levels of CB2R mRNA in rodent brains (Tubbs et al, 2013; Wang et al, 2012). Figure 5a shows mouse and rat coronal atlas, illustrating PFC, VTA, DST, and NAC. Figure 5b and c show the quantitative analysis results. We used ImageJ software to count the pixels of CB2R mRNA of 6–8 cells in 2 to 3 brain sections of each brain region and intensities of CB2R mRNA in VTA dopamine (DA+) and nondopamine (DA−) cells. We observed that CB2R mRNA pixels (2–25) in each cell were lowest in NAC and DST (2–5) and highest in VTA (20–25) in both rats and mice. Overall, more CB2R mRNAs were found in each brain region in mice than in rats (Figure 5b, F3, 8=46, p<0.001). There was less CB2R mRNA in DA neurons than those of non-DA neurons (Figure 5c, F1, 2=678, p<0.01) in rat VTA but no difference in mouse VTA. Figure 5d shows representative CB2R mRNA staining using the RNAscope probes that hybridize to the 3′-UTR of CB2R gene in mice (NM_009924, 1877–2820 bp, Figure 2a) and rats (NM_001164143, 1935–2843 bp, Figure 2b), illustrating CB2 mRNA expression in each brain region. In the VTA, CB2 mRNA was detectable in TH-positive or DAT-positive neurons, respectively, in mice and rats.
Brain CB2 mRNA expression by RNAscope in situ hybridization (ISH) assays. (a) Brain section diagrams, illustrating the anatomic locations and the coordinates of PFC, DST, NAC, and VTA for the ISH images. (b, c) Quantification of CB2R pixels or densities per cell in each brain region of mice and rats. (d) Representative RNAscope ISH microscope images of different brain regions. CB2 mRNA signals are green, TH and DAT signals are red, and nuclear signals are blue (DAPI). Calibration bar is 20 μm. *P<0.05, compared with rats.
Species Differences in CB2R Protein Structures
As there are no actual X-ray crystallographic structures of cannabinoid receptor CB1R and CB2R, we predicted 3D structures of rat and mouse CB2Rs based on the structure of turkey β-adrenergic receptor (residues 2–327; PDB ID code 4ajm, chain B) that shares amino-acid sequence identity of 23.5% (90/383) and similarity of 39.7% (152/383) with human hCB2R. Figure 6 shows the predicted 3D structures of mCB2R and rCB2R (residues 27–315) with 7 transmembrane domains (TMs) and 3 extra- and 3 intracellular loops (ECL 1–3 and ICL 1–3). We drew the N- and C-terminus portions in both 3D and 2D models in order to show the truncated sequence in mouse CB2R. The amino-acid sequences are highly conserved in the 7 TMs and less conserved in ECLs and ICLs. The acidic and basic amino-acid substitutions in ECLs and ICLs are marked by numbers and colored lines between rats and mice. The ligand-binding motifs of TM3-DRY (ionic lock) and TM7-NPXXY (water pocket) (Rosenbaum et al, 2009) and functional amino acids S112, D130, L201, Y207, and A244 (Feng and Song, 2003; Song and Feng, 2002; Tao et al, 1999) as well as important cysteine residues are completely conserved in primate and rodent CB2Rs (Mercier et al, 2010). The proline residues in the TM regions involved in π-helical conformation (bend helices) are also conserved (Filipek et al, 2003).
CB2R 3D and 2D structures in rat and mouse. (a, b) In silico models of mCB2R and rCB2R 3D structures. (c, d) The corresponding amino-acid sequences of mCB2R and rCB2R. Ribbons represent seven transmembrane domains and bended lines represent intracellular and extracellular domains. The ‘DRY’ motif is indicated by colored balls. The charged amino-acid exchanges in the extracellular and the intracellular loops between mCB2R and rCB2R are marked by colored narrow lines labeled with amino-acid codes and numbers in the 3D models. The truncated 13 amino acids of mCB2R are marked with single amino-acid code in the intracellular C-terminal domain of rCB2R. The autophosphorylation site S352 is marked in red. The charged amino acid substitutions and the C-terminal truncation of mCB2R are marked in the mCB2R with dark red coloration (c, d) and other amino acid substitutions between mCB2R and rCB2R are marked in light red coloration (d).
There are functional amino-acid substitutions between the mouse and rat receptors. The charged amino-acid substitutions between mCB2R and rCB2R are located in the proximity of the TMs (Figure 6c and d; IL1: R62Q, EL1: N103R, IL3: R218Q, and EL3: Q276K, the first letter for mouse and the second letter for rat) that might affect ligand binding and signal transduction. For example, Feng and Song (2003) have shown that the amino acids D3.49 and R3.50 in DRY motif of TM3 and A6.34 in TM6 of CB2R are involved in ligand binding and signal transduction. Furthermore, the positively charged amino-acid substitution of C136R between rodent and primate CB2R, respectively, is three amino acids downstream of the DRY domain and the substitution might change the ligand binding affinity between human and rat CB2Rs.
The most striking difference of CB2Rs between rat and mouse is that mCB2R lacks the intracellular C-terminal 13 amino acids TGPGSRTPGCSNC of rCB2R (348–360) as marked by single letters at the C-terminus of rCB2R (Figure 6b and d). We searched The Eukaryotic Linear Motif Database and NetPhosK 2.0 using the intracellular C-terminal sequence (59 amino acids) of rCB2R and found consensus motifs containing protein kinase C (PKC) phosphorylation site KSS (334–336, NetPhos score 0.995), G protein-coupled receptor kinase (GRK) phosphorylation site ETE (339–441, NetPhos score 0.663), and autophosphorylation site S352 (NetPhos score 0.962). The mCB2R truncated 13 amino-acid sequence is located in the intrinsically disordered protein (IDP) motif KTTTGPGSRTPGCS (344–358, ELM prediction) (Dinkel et al, 2012) that allows more interactions with protein partners and modification sites. Therefore, the rCB2R auophosphorylation site S352 (labeled in red in Figure 6b) and IDP motif do not exist in mCB2R C-terminal intracellular domain.
DISCUSSION
One of the major findings in this study is that systemic administration of JWH133, a highly selective CB2R agonist, produced different effects on i.v. cocaine self-administration in rats and mice. JWH133 dose-dependently inhibited cocaine self-administration under FR reinforcement in mice, but not in rats. Under PR reinforcement, JWH133 produced an increase in PR BP for cocaine self-administration in rats, but a decrease in PR BP for cocaine self-administration in mice. JWH133 may have relatively poor bioavailability because of rapid metabolism in liver after i.p. administration and/or rapid redistribution from plasma to fatty tissues after absorption. Therefore, higher drug doses of systemic JWH133 may be required to produce a similar inhibitory effect in rats as seen in mice. To explore this hypothesis, JWH133 was delivered intranasally—a route by which the drug can directly enter the brain and bypass peripheral tissues. We found that JWH133 produced biphasic effects: low doses increased whereas a high dose inhibited PR cocaine self-administration in rats, suggesting that the increase in PR BP could be a compensatory response to a reduction in cocaine’s rewarding efficacy after low doses of JWH133.
CB2R gene structure and mRNA splicing are different in different species (Liu et al, 2009). This raises the possibility that CB2Rs may have different regional and/or cellular distributions in brain between rats and mice and, therefore, may subserve different effects on motivation for drug taking and consumption. To explore this, we first carefully examined CB2R gene or transcript expression in rats and mice by quantitative RT-PCR. We found four different rCB2R mRNA isoforms. CB2A and CB2B are present in mice and rats, and CB2C and CB2D are present only in rats. It is unknown how these different CB2R mRNA isoforms alter CB2R expression in the brain. As the intact rCB2R protein-coding region exists in the CB2A, CB2B, CB2C, and CB2D isoforms, we suggest that these four isoforms may encode or translate the same CB2R in amino-acid sequence. The different sequences of 5′-UTR and 3′-UTR of the isoforms might alter brain regional and cell type-specific expression and mRNA stabilities. We note that the quantification of CB2R isoform using qRT-PCR and agarose fragment analysis might not be accurate because of different probe designs and PCR cycles, respectively. We therefore used the RNAscope ISH assays to observe CB2R mRNA expression patterns and quantify density in brain regions (VTA, NAC, PFC, and DST). We found that all of the examined mouse brain regions have more CB2R mRNA than rat brain regions. The higher levels of CB2R mRNA in mice might indicate a compensatory upregulation of mCB2R mRNA because of the C-terminal truncation. We also noted that CB2R mRNA levels in TH-positive DA neurons and TH-negative cells were very similar in mice, but were significantly lower in DAT-positive DA neurons than DAT-negative neurons in rats, suggesting species differences of CB2R mRNA expression in DA neurons. This might in part contribute to the different behavioral responses to the agonist JWH133 seen in rats and mice.
We also examined the rCB2R410 isoform (AF218846) that encodes 410 amino acids (Brown et al, 2002). The rCB2R410 isoform is produced by an intraexonic splicing site upstream of the stop codon (nucleotide positions at FJ694960: 1198–2141, deletion of 943 bp), creating a frameshift that changes the C-terminal amino-acid sequence (Brown et al, 2002). However, in this study, we could not detect this frameshift isoform expression in rat brain and peripheral tissues using TaqMan probe and primers across the rCB2R410 splicing site (data not shown). We detected the expected sizes of the rCB2R360 and rCB2D PCR fragments. Therefore, we concluded that the rCB2R360 is a predominant isoform in rat. The rCB2D isoform is caused by splicing site downstream of the stop codon (nucleotide positions at FJ694960: 1355–2141, deletion of 786 bp) that does not alter the C-terminal amino-acid sequence.
In addition to species differences in CB2R gene splicing, we also found that CB2Rs display significant species differences in amino acid sequences. There is 96% amino-acid homology between human and rhesus and 93% amino-acid homology between rat and mouse CB2 receptors. Human CB2R shares similar amino-acid homologies with mouse (82%) and rat (81%). However, the mCB2R truncates the intracellular C-terminal 13 amino acids because of a premature stop codon that reduces mCB2R to 347 amino acids instead of 360 amino acids (Liu et al, 2009). The truncated carboxyl-terminus of CB2R contains a functional autophosphorylation site (Ser352) that may induce CB2R internalization, an effect that is blocked by the CB2R antagonist SR144528 (Bouaboula et al, 1999). The truncated C-terminus potentially contains the PDZ (postsynaptic density protein structural domain)-binding motif PGCSNC (355–360) (Bar-Shira and Chechik, 2013) that might position the rCB2R in the postsynaptic density region as shown in hippocampus (Brusco et al, 2008a, 2008b). Therefore, the postsynaptic signaling mechanisms of the CB2R might differ from the presynaptic retrograde signaling of the CB1R that is induced by depolarization-suppressing glutamate excitatory and GABA inhibitory neurotransmissions (Benarroch, 2007). However, synergistic actions between CB1R and CB2R might be implied by pre- and postsynaptic localizations of CB1R in nucleus accumbens (Pickel et al, 2006). In comparison, rCB2R360 (FJ694960) retains 360 amino acids (Liu et al, 2009) and an intact C-terminal intracellular sequence, including the autophosphorylation Ser352 site and the putative PDZ (355–360) domain, similar to the situation in the hCB2R. The shortened C-terminus of mCB2R might also change G-protein coupling for ligand-induced intracellular signal transduction. These findings suggest that the intracellular signal pathways underlying CB2R signaling may be different in mouse and rat, and this may in part explain why JWH133 produces different effects on cocaine self-administration behavior in rats and mice. According to the similarity of the primary amino-acid sequences of the CB2R, rCB2Rs might be more relevant to hCB2Rs, whereas according to the similarity of gene splicing pattern, mCB2Rs might be more relevant to hCB2Rs.
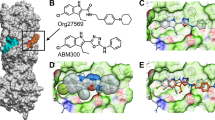
Importantly, CB2Rs in different species also display different functional responses to the same ligands. For example, JWH133 is reported to be relatively selective for hCB2Rs over rCB2Rs, whereas the selective antagonist AM630 is more potent at rCB2Rs than at hCB2Rs, and the affinity of JWH133 for rCB2Rs or mCB2Rs is unknown (Huffman et al, 1999; Marriott et al, 2006). However, another CB2R agonist, S-AM1241, shows clear differential affinity for rCB2R (893±58.5 nM) and mCB2R (577±58.4 nM), and regulation of cAMP levels for rCB2R (EC50: 785+564 nM) and mCB2R (EC50: 2000+475 nM) in rCB2R- and mCB2R-transfected CHO cells (Yao et al, 2006; Bingham et al, 2007), respectively. JWH133 might have differential affinities and cAMP effects at rCB2Rs and mCB2Rs as well. The endocannabinoids (anandamide and 2-arachidonyl glycerol) and WIN55212-2 are more hCB2R selective, whereas CP55940 has similar affinity for both hCB2Rs and rCB2Rs (Griffin et al, 2000; Mukherjee et al, 2004; Yao et al, 2006). For computer modeling, we docked JWH133 to the active site of hCB2R structure and predicted that JWH133 binding sites are located in TM2 (F87 and F91), TM3 (V113, W114, and F117), TM5 (L185, W194, and I198), TM6 (W258 and M265), and TM7 (S285). Although JWH133 binding sites are conserved in primates and rodents (Figure 7a and b), considerable differences in CB2R amino-acid sequences and tertiary (3D) structures may underlie different receptor responses to various cannabinoid ligands in different species. Taken together, all these findings suggest that species differences in CB2R gene and receptor expression in the brain may in part explain the different effects of JWH133 on cocaine self-administration described above.
Computer models of JWH133 binding site of CB2R. (a) JHW133 docking to the active site of hCB2R. Green rings and lines represent JHW133 structure. (b) Complex conformations between JHW133 and hCB2R. The JHW133 structure is represented by gray spheres and the amino-acid residues that contact with JHW133 are indicated by three-letter amino-acid symbols and numbers.
During speciation, CB1R evolved earlier than CB2R. Chemotaxonomic and phylogenomic studies (McPartland, 2004) revealed that the genomes of non-chordate invertebrates do not encode CB1R and CB2R genes that are present in the genomes of vertebrates. Urochordates, a relative of vertebrates, encode only CB1R and not CB2R (Elphick, 2007). In non-chordate invertebrates lacking CB1R and CB2R genes, endocannabinoids, that is, anandamide and 2-arachidonyl glycerol, are found. These might act on presynaptic transient receptor potential cation channel subfamily V member 1 (Trpv1) or on vanilloid receptor 1 (VR1) that evolved earlier than vertebrate cannabinoid receptors for regulation of invertebrate synaptic plasticity (Elphick, 2012). The enrichment of CB1R in neuron and CB2R in microglia might indicate commutative interactions of cannabinoids in neuron–glia circuitry (Cutando et al, 2013; Zhang et al, 2014b). The neuronal pre- and postsynaptic colocalization of CB1R and CB2R might imply cooperative signaling of cannabinoids (Brusco et al, 2008a; Reyes et al, 2009).
The existence of a premature stop codon might represent an effective regulation to change mCB2R function by truncation of the C-terminal 13 amino acids. A mechanism also occurs in transient receptor potential cation channel (TRPC2) in hominoids and old-world monkeys that alters pheromone perception (Zhang and Webb, 2003), and in the taste receptor TAS2R38 that changes the bitter taste perception of phenylthiocarbamide (PTC) in different mammalian species, including humans (Wooding, 2011). The CB2R gene exhibits tissue- and species-specific alternative splicing because of cis-directed evolutionary changes in the nucleotide sequences of splicing sites of inter- and intraexon junctions (Barbosa-Morais et al, 2012). Within exon 2 of the rCB2R gene, there is a potential 3′-splicing site (AG) with the consensus sequence of CAG/G that is changed to the nonconsensus sequence CAG/T in the corresponding mouse exon 2. Therefore, the rCB2R exons 1–3 could be spliced together to form a new rCB2C isoform. Tissue-specific gene expression is largely conserved in mammalian species but alternative splicing patterns are less conserved and often species specific (Merkin et al, 2012). The human hCB2R gene lost rodent exon 1 and gained two exons further upstream during ∼75 million years of evolutionary branching from rodent (Modrek and Lee, 2003). Thus, promoter sequences and epigenetic regulation (Onaivi et al, 2012) of human CB2R differ from rodents, as shown that hCB2A is predominantly expressed in human testis (Liu et al, 2009; Onaivi et al, 2012). The emergence of alternative splicing sites of rat exon 2 and exon 3 created more rat rCB2R transcript variants (4–5 isoforms) than those of mouse mCB2R (2 isoforms) (Jorda et al, 2003; Koren et al, 2007). Human-specific alternative splicing is also present in hCB1R (CNR1) gene in which intraexonic splicing of the coding exon produces different N-terminal domains that exhibit tissue-specific expression patterns (Shire et al, 1995). Therefore, alternative splicing plays a major role in mammalian evolution of cannabinoid system.
Conventional ISH is unable to detect CB2R expression in nonstimulated rodent brain neurons because of lower sensitivity and specificity of unamplified probes (Wang et al, 2012). The ultrasensitive RNAscope ISH with amplified fluorescent probes detected more CB2R mRNA signals in mouse brain than in rat brain (Figure 5), and rCB2R protein was observed in postsynaptic regions (Brusco et al, 2008a, 2008b). Differential brain CB2R expression may also in part explain different effects of JWH133 on cocaine self-administration in rats and mice. This gene–behavior correlation is also observed in Lewis and Fischer 344 rat strains. For example, Lewis rats display lower PR BPs for cocaine self-administration, but have higher CB2R expression in hippocampus than those of Fischer 344 rats who display higher PR BPs for cocaine self-administration (Rivera et al, 2013; Kosten et al, 2007), which agrees with our findings in mice and rats. Therefore, quantitative and qualitative changes in the mCB2R gene might contribute to differential receptor responses to cocaine self-administration.
In conclusion, evolutionary changes in CB2R including its coding sequences, splicing patterns, and gene regulatory elements might contribute in part to differential effects of CB2R ligands on cocaine self-administration (Gamaleddin et al, 2012a; Ignatowska-Jankowska et al, 2013; Xi et al, 2011). With growing evidence that CB2Rs play an important role in addiction, psychiatric, and neurological disorders, more studies are required to explore species differences in CB2R gene expression and signal transduction. Proper selection of animal models is important in the development of effective CB2R-based therapeutics for addiction and other psychiatric and neurological diseases in humans.
FUNDING AND DISCLOSURE
The authors declare no conflict of interest.
References
Adamczyk P, Miszkiel J, McCreary AC, Filip M, Papp M, Przegaliński E (2012). The effects of cannabinoid CB1, CB2 and vanilloid TRPV1 receptor antagonists on cocaine addictive behavior in rats. Brain Res 1444: 45–54.
Al Mansouri S, Ojha S, Al Maamari E, Al Ameri M, Nurulain SM, Bahi A (2014). The cannabinoid receptor 2 agonist, β-caryophyllene, reduced voluntary alcohol intake and attenuated ethanol-induced place preference and sensitivity in mice. Pharmacol Biochem Behav 124: 260–268.
Aracil-Fernández A, Trigo JM, García-Gutiérrez MS, Ortega-Álvaro A, Ternianov A, Navarro D et al (2012). Decreased cocaine motor sensitization and self-administration in mice overexpressing cannabinoid CB2 receptors. Neuropsychopharmacology 37: 1749–1763.
Arnold K, Bordoli L, Kopp J, Schwede T (2006). The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22: 195–201.
Atwood BK, Mackie K (2010). CB2: a cannabinoid receptor with an identity crisis. Br J Pharmacol 160: 467–479.
Bar-Shira O, Chechik G (2013). Predicting protein-protein interactions in the post synaptic density. Mol Cell Neurosci 56: 128–139.
Barbosa-Morais NL, Irimia M, Pan Q, Xiong HY, Gueroussov S, Lee LJ et al (2012). The evolutionary landscape of alternative splicing in vertebrate species. Science 338: 1587–1593.
Benarroch E (2007). Endocannabinoids in basal ganglia circuits: implications for Parkinson disease. Neurology 69: 306–309.
Bingham B, Jones PG, Uveges AJ, Kotnis S, Lu P, Smith VA et al (2007). Species-specific in vitro pharmacological effects of the cannabinoid receptor 2 (CB2) selective ligand AM1241 and its resolved enantiomers. Br J Pharmacol 151: 1061–1070.
Blanco-Calvo E, Rivera P, Arrabal S, Vargas A, Pavón FJ, Serrano A et al (2014). Pharmacological blockade of either cannabinoid CB1 or CB2 receptors prevents both cocaine-induced conditioned locomotion and cocaine-induced reduction of cell proliferation in the hippocampus of adult male rat. Front Integr Neurosci 7: 106.
Bouaboula M, Dussossoy D, Casellas P (1999). Regulation of peripheral cannabinoid receptor CB2 phosphorylation by the inverse agonist SR 144528. Implications for receptor biological responses. J Biol Chem 274: 20397–20405.
Brown SM, Wager-Miller J, Mackie K (2002). Cloning and molecular characterization of the rat CB2 cannabinoid receptor. Biochim Biophys Acta 1576: 255–264.
Brusco A, Tagliaferro P, Saez T, Onaivi ES (2008a). Postsynaptic localization of CB2 cannabinoid receptors in the rat hippocampus. Synapse 62: 944–949.
Brusco A, Tagliaferro PA, Saez T, Onaivi ES (2008b). Ultrastructural localization of neuronal brain CB2 cannabinoid receptors. Ann NY Acad Sci 1139: 450–457.
Buckley NE, McCoy KL, Mezey E, Bonner T, Zimmer A, Felder CC et al (2000). Immunomodulation by cannabinoids is absent in mice deficient for the cannabinoid CB2 receptor. Eur J Pharmacol 396: 141–149.
Callén L, Moreno E, Barroso-Chinea P, Moreno-Delgado D, Cortés A, Mallol J et al (2012). Cannabinoid receptors CB1 and CB2 form functional heteromers in brain. J Biol Chem 287: 20851–20865.
Costantino HR, Illum L, Brandt G, Johnson PH, Quay SC (2007). Intranasal delivery: physicochemical and therapeutic aspects. Int J Pharm 337: 1–24.
Cutando L, Busquets-Garcia A, Puighermanal E, Gomis-Gonzalez M, Delgado-Garcia JM, Gruart A et al (2013). Microglial activation underlies cerebellar deficits produced by repeated cannabis exposure. J Clin Invest 123: 2816–2831.
den Boon FS, Chameau P, Schaafsma-Zhao Q, van Aken W, Bari M, Oddi S et al (2012). Excitability of prefrontal cortical pyramidal neurons is modulated by activation of intracellular type-2 cannabinoid receptors. Proc Natl Acad Sci USA 109: 3534–3539.
Dinkel H, Michael S, Weatheritt RJ, Davey NE, Van Roey K, Altenberg B et al (2012). ELM—the database of eukaryotic linear motifs. Nucleic Acids Res 40(Database issue): D242–D251.
Elphick MR (2007). BfCBR: a cannabinoid receptor ortholog in the cephalochordate Branchiostoma floridae (Amphioxus). Gene 399: 65–71.
Elphick MR (2012). The evolution and comparative neurobiology of endocannabinoid signalling. Philos Trans R Soc Lond B Biol Sci 367: 3201–3215.
Feng W, Song ZH (2003). Effects of D3.49A, R3.50A, and A6.34E mutations on ligand binding and activation of the cannabinoid-2 (CB2) receptor. Biochem Pharmacol 65: 1077–1085.
Filipek S, Teller DC, Palczewski K, Stenkamp R (2003). The crystallographic model of rhodopsin and its use in studies of other G protein-coupled receptors. Annu Rev Biophys Biomol Struct 32: 375–397.
Gamaleddin I, Wertheim C, Zhu AZ, Coen KM, Vemuri K, Makryannis A et al (2012a). Cannabinoid receptor stimulation increases motivation for nicotine and nicotine seeking. Addict Biol 17: 47–61.
Gamaleddin I, Zvonok A, Makriyannis A, Goldberg SR, Le Foll B (2012b). Effects of a selective cannabinoid CB2 agonist and antagonist on intravenous nicotine self administration and reinstatement of nicotine seeking. PLoS One 7: e29900.
Garcia-Gutiérrez MS, Manzanares J (2011). Overexpression of CB2 cannabinoid receptors decreased vulnerability to anxiety and impaired anxiolytic action of alprazolam in mice. J Psychopharmacol 25: 111–120.
Gong J-P, Onaivi ES, Ishiguro H, Liu Q-R, Tagliaferro PA, Brusco A et al (2006). Cannabinoid CB2 receptors: immunohistochemical localization in rat brain. Brain Res 1071: 10–23.
Griffin G, Tao Q, Abood ME (2000). Cloning and pharmacological characterization of the rat CB(2) cannabinoid receptor. J Pharmacol Exp Ther 292: 886–894.
Guex N, Peitsch MC (1997). SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18: 2714–2723.
Gutiérrez MS, Ortega-Álvaro A, Busquets-García A, Pérez-Ortiz JM, Caltana L, Ricatti MJ et al (2013). Synaptic plasticity alterations associated with memory impairment induced by deletion of CB2 cannabinoid receptors. Neuropharmacology 73: 388–396.
Huffman JW, Liddle J, Yu S, Aung MM, Abood ME, Wiley JL et al (1999). 3-(1′,1′-Dimethylbutyl)-1-deoxy-delta8-THC and related compounds: synthesis of selective ligands for the CB2 receptor. Bioorg Med Chem 7: 2905–2914.
Ignatowska-Jankowska BM, Muldoon PP, Lichtman AH, Damaj MI (2013). The cannabinoid CB receptor is necessary for nicotine-conditioned place preference, but not other behavioral effects of nicotine in mice. Psychopharmacology (Berl) 229: 591–601.
Illum L, Watts P, Fisher AN, Hinchcliffe M, Norbury H, Jabbal-Gill I et al (2002). Intranasal delivery of morphine. J Pharmacol Exp Ther 301: 391–400.
Ishiguro H, Carpio O, Horiuchi Y, Shu A, Higuchi S, Schanz N et al (2010a). A nonsynonymous polymorphism in cannabinoid CB2 receptor gene is associated with eating disorders in humans and food intake is modified in mice by its ligands. Synapse 64: 92–96.
Ishiguro H, Horiuchi Y, Ishikawa M, Koga M, Imai K, Suzuki Y et al (2010b). Brain cannabinoid CB2 receptor in schizophrenia. Biol Psychiatry 67: 974–982.
Jorda MA, Rayman N, Valk P, De Wee E, Delwel R (2003). Identification, characterization, and function of a novel oncogene: the peripheral cannabinoid receptor Cb2. Ann NY Acad Sci 996: 10–16.
Koren E, Lev-Maor G, Ast G (2007). The emergence of alternative 3' and 5' splice site exons from constitutive exons. PLoS Comput Biol 3: e95.
Kosten TA, Zhang XY, Haile CN (2007). Strain differences in maintenance of cocaine self-administration and their relationship to novelty activity responses. Behav Neusci 121: 380–388.
Lanciego JL, Barroso-Chinea P, Rico AJ, Conte-Perales L, Callen L, Roda E et al (2011). Expression of the mRNA coding the cannabinoid receptor 2 in the pallidal complex of Macaca fascicularis. J Psychopharmacol 25: 97–104.
Liu Q-R, Pan CH, Hishimoto A, Li C-Y, Xi Z-X, Llorente-Berzal A et al (2009). Species differences in cannabinoid receptor 2 (CNR 2 gene): identification of novel human and rodent CB2 isoforms, differential tissue expression and regulation by cannabinoid receptor ligands. Genes Brain Behav 8: 519–530.
Liu Q-R, Rubio FJ, Bossert JM, Marchant NJ, Fanous S, Hou X et al (2014). Detection of molecular alterations in methamphetamine-activated Fos-expressing neurons from a single rat dorsal striatum using fluorescence-activated cell sorting (FACS). J Neurochem 128: 173–185.
Marriott KS, Huffman JW, Wiley JL, Martin BR (2006). Synthesis and pharmacology of 11-nor-1-methoxy-9-hydroxyhexahydrocannabinols and 11-nor-1-deoxy-9-hydroxyhexahydrocannabinols: new selective ligands for the cannabinoid CB2 receptor. Bioorg Med Chem 14: 2386–2397.
Matsuda LA (1997). Molecular aspects of cannabinoid receptors. Crit Rev Neurobiol 11: 143–166.
McPartland JM (2004). Phylogenomic and chemotaxonomic analysis of the endocannabinoid system. Brain Res Brain Res Rev 45: 18–29.
Mechoulam R, Parker LA (2013). The endocannabinoid system and the brain. Annu Rev Psychol 64: 21–47.
Mercier RW, Pei Y, Pandarinathan L, Janero DR, Zhang J, Makriyannis A (2010). hCB2 ligand-interaction landscape: cysteine residues critical to biarylpyrazole antagonist binding motif and receptor modulation. Chem Biol 17: 1132–1142.
Merkin J, Russell C, Chen P, Burge CB (2012). Evolutionary dynamics of gene and isoform regulation in Mammalian tissues. Science 338: 1593–1599.
Modrek B, Lee CJ (2003). Alternative splicing in the human, mouse and rat genomes is associated with an increased frequency of exon creation and/or loss. Nat Genet 34: 177–180.
Morgan NH, Stanford IM, Woodhall GL (2009). Functional CB2 type cannabinoid receptors at CNS synapses. Neuropharmacology 57: 356–368.
Mukherjee S, Adams M, Whiteaker K, Daza A, Kage K, Cassar S et al (2004). Species comparison and pharmacological characterization of rat and human CB2 cannabinoid receptors. Eur J Pharmacol 505: 1–9.
Navarrete F, Rodríguez-Arias M, Martín-García E, Navarro D, García-Gutiérrez MS, Aguilar MA et al (2013). Role of CB2 cannabinoid receptors in the rewarding, reinforcing, and physical effects of nicotine. Neuropsychopharmacology 38: 2515–2524.
Onaivi ES, Ishiguro H, Gong J-P, Patel S, Meozzi PA, Myers L et al (2008). Functional expression of brain neuronal CB2 cannabinoid receptors are involved in the effects of drugs of abuse and in depression. Ann NY Acad Sci 1139: 434–449.
Onaivi ES, Ishiguro H, Gu S, Liu Q-R (2012). CNS effects of CB2 cannabinoid receptors: beyond neuro-immuno-cannabinoid activity. J Psychopharmacol 26: 92–103.
Pickel VM, Chan J, Kearn CS, Mackie K (2006). Targeting dopamine D2 and cannabinoid-1 CB1 receptors in rat nucleus accumbens. J Comp Neurol 495: 299–313.
Reyes BA, Rosario JC, Piana PM, Van Bockstaele EJ (2009). Cannabinoid modulation of cortical adrenergic receptors and transporters. J Neurosci Res 87: 3671–3678.
Rivera P, Miguens M, Coria SM, Rubio L, Higuera-Matas A, Bermudez-Silva FJ et al (2013). Cocaine self-administration differentially modulates the expression of endogenous cannabinoid system-related proteins in the hippocampus of Lewis vs. Fischer 344 rats. Int J Neuropsychopharmacol 16: 1277–1293.
Rosenbaum DM, Rasmussen SG, Kobilka BK (2009). The structure and function of G-protein-coupled receptors. Nature 459: 356–363.
Schwede T, Kopp J, Guex N, Peitsch MC (2003). SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res 31: 3381–3385.
Shire D, Carillon C, Kaghad M, Calandra B, Rinaldi-Carmona M, Le Fur G et al (1995). An amino-terminal variant of the central cannabinoid receptor resulting from alternative splicing. J Biol Chem 270: 3726–3731.
Song R, Zhang HY, Li X, Bi GH, Gardner EL, Xi ZX (2012). Increased vulnerability to cocaine in mice lacking dopamine D3 receptors. Proc Natl Acad Sci USA 109: 17675–17680.
Song ZH, Feng W (2002). Absence of a conserved proline and presence of a conserved tyrosine in the CB2 cannabinoid receptor are crucial for its function. FEBS Lett 531: 290–294.
Suárez J, Llorente R, Romero-Zerbo SY, Mateos B, Bermúdez-Silva FJ, de Fonseca FR et al (2009). Early maternal deprivation induces gender-dependent changes on the expression of hippocampal CB1 and CB2 cannabinoid receptors of neonatal rats. Hippocampus 19: 623–632.
Tao Q, McAllister SD, Andreassi J, Nowell KW, Cabral GA, Hurst DP et al (1999). Role of a conserved lysine residue in the peripheral cannabinoid receptor CB2: evidence for subtype specificity. Mol Pharmacol 55: 605–613.
Tubbs RR, Wang H, Wang Z, Minca EC, Portier BP, Gruver AM et al (2013). Ultrasensitive RNA in situ hybridization for detection of restricted clonal expression of low-abundance immunoglobulin light chain mRNA in B-cell lymphoproliferative disorders. Am J Clin Pathol 140: 736–746.
Van Sickle MD, Duncan M, Kingsley PJ, Mouihate A, Urbani P, Mackie K et al (2005). Identification and functional characterization of brainstem cannabinoid CB2 receptors. Science 310: 329–332.
Vinckenbosch N, Dupanloup I, Kaessmann H (2006). Evolutionary fate of retroposed gene copies in the human genome. Proc Natl Acad Sci USA 103: 3220–3225.
Wang F, Flanagan J, Su N, Wang L-C, Bui S, Nielson A et al (2012). RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J Mol Diagn 14: 22–29.
Wooding S (2011). Signatures of natural selection in a primate bitter taste receptor. J Mol Evol 73: 257–265.
Xi Z-X, Gilbert JG, Pak AC, Ashby CR Jr., Heidbreder CA, Gardner EL (2005). Selective dopamine D3 receptor antagonism by SB-277011A attenuates cocaine reinforcement as assessed by progressive-ratio and variable-cost-variable-payoff fixed-ratio cocaine self-administration in rats. Eur J Neurosci 21: 3427–3438.
Xi Z-X, Peng X-Q, Li X, Song R, Zhang H-Y, Liu Q-R et al (2011). Brain cannabinoid CB receptors modulate cocaine's actions in mice. Nat Neurosci 14: 1160–1166.
Yao BB, Mukherjee S, Fan Y, Garrison TR, Daza AV, Grayson GK et al (2006). In vitro pharmacological characterization of AM1241: a protean agonist at the cannabinoid CB2 receptor? Br J Pharmacol 149: 145–154.
Zhang J, Webb DM (2003). Evolutionary deterioration of the vomeronasal pheromone transduction pathway in catarrhine primates. Proc Natl Acad Sci USA 100: 8337–8341.
Zhang S-J, Liu C-J, Shi M, Kong L, Chen J-Y, Zhou W-Z et al (2013). RhesusBase: a knowledgebase for the monkey research community. Nucleic Acids Res 41(Database issue): D892–D905.
Zhang S-J, Liu C-J, Yu P, Zhong X, Chen J-Y, Yang X et al (2014a). Evolutionary interrogation of human biology in well-annotated genomic framework of rhesus macaque. Mol Biol Evol 31: 1309–1324.
Zhang Y, Chen K, Sloan SA, Bennett ML, Scholze AR, O'Keeffe S et al (2014b). An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J Neurosci 34: 11929–11947.
Acknowledgements
We gratefully acknowledge Dr Yavin Shaham, chief of the Behavioral Neuroscience Research Branch, Intramural Research Program, NIDA-NIH, for invaluable advice on statistical analysis. H-YZ, G-HB, XL, JL, Z-XX, ELG, and Q-RL are supported by the Intramural Research Program of the National Institute on Drug Abuse, National Institute of Health (NIDA-NIH). HQ is supported by the National Natural Science Foundation of China (31171270), and C-YL and S-JZ are supported by the National Key Basic Research Program of China (2012CB518004 and 2013CB531202). ESO acknowledges Guest Researcher support at NIDA-NIH and NIH grant DA032890.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhang, HY., Bi, GH., Li, X. et al. Species Differences in Cannabinoid Receptor 2 and Receptor Responses to Cocaine Self-Administration in Mice and Rats. Neuropsychopharmacol 40, 1037–1051 (2015). https://doi.org/10.1038/npp.2014.297
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2014.297
This article is cited by
-
Exploring the therapeutic potential of natural compounds modulating the endocannabinoid system in various diseases and disorders: review
Pharmacological Reports (2023)
-
Repeated cocaine administration upregulates CB2 receptor expression in striatal medium-spiny neurons that express dopamine D1 receptors in mice
Acta Pharmacologica Sinica (2022)
-
Involvement of the ghrelin system in the maintenance and reinstatement of cocaine-motivated behaviors: a role of adrenergic action at peripheral β1 receptors
Neuropsychopharmacology (2022)
-
Cannabinoid receptor 2 deletion influences social memory and synaptic architecture in the hippocampus
Scientific Reports (2021)
-
Beta-caryophyllene inhibits cocaine addiction-related behavior by activation of PPARα and PPARγ: repurposing a FDA-approved food additive for cocaine use disorder
Neuropsychopharmacology (2021)