Abstract
Evidence suggests that depression is a risk factor for dementia; however, the relationship between the two conditions is not fully understood. A novel gene (TOMM40) has been consistently associated with Alzheimer’s disease (AD), but has received no attention in depression. We conducted a three-level cross-sectional study to investigate the association of the TOMM40 rs2075650 SNP with depression. We recruited a community sample of 1220 participants (571 controls, 649 lifetime depression) to complete a psychiatric background questionnaire, the Brief Symptom Inventory, and Big Five Inventory at Level-1, 243 (102 controls, 97 remitted, 44 currently depressed) to complete a face-to-face clinical interview and neuropsychological testing at Level-2 and 58 (33 controls, 25 remitted) to complete an emotional face-processing task during fMRI at Level-3. Our results indicated that the TOMM40 rs2075650 G allele was a significant risk factor for lifetime depression (p=0.00006) and, in depressed subjects, was a significant predictor of low extraversion (p=0.009). Currently depressed risk allele carriers showed subtle executive dysfunction (p=0.004) and decreased positive memory bias (p=0.021) together with reduced activity in the posterior (p(FWE)=0.045) and anterior (p(FWE)=0.041) cingulate during sad face emotion processing. Our results suggest that TOMM40 rs2075650 may be a risk factor for the development of depression characterized by reduced extraversion, impaired executive function, and decreased positive emotional recall, and reduced top-down cortical control during sad emotion processing.
Similar content being viewed by others
INTRODUCTION
There is a near two-fold increased risk of developing dementia, particularly Alzheimer’s disease (AD), after a diagnosis of depression (Green et al, 2003; Jorm, 2001; Ownby et al, 2006; Saczynski et al, 2010). This risk may be partially explained by depression forming a dementia prodrome, given the higher risk of dementia in late-life depression compared with earlier onset depression (Barnes et al, 2012). However, this cannot explain the greater risk of developing dementia in those who only suffer from depression earlier in their lives (Green et al, 2003), suggesting common biology linking the conditions. If true, then genetic factors associated with an increased risk of dementia may also contribute to an increased risk of depression.
Genetic susceptibility to developing AD is well recognized. Both the apolipoprotein E (APOE) gene and the TOMM40-APOE locus, tagged by rs2075650 in the translocase of outer mitochondrial membrane 40 (TOMM40) gene (see Supplementary Information), have been implicated (Hong et al, 2010; Li et al, 2008; Schiepers et al, 2011). Investigations of depression have also indicated that the APOE ɛ4 risk allele influences both the onset and risk for late-life depression (Butters et al, 2003; Yen et al, 2007) and has been associated with gray matter reductions (Cherbuin et al, 2007), reduced white matter integrity (Heise et al, 2011) and cognitive impairment (Greenwood et al, 2000) in healthy adults, further suggesting that genetic mechanisms associated with dementia may contribute to the development of mood disorder.
In comparison to APOE, the independent role of TOMM40 in AD and depression has been little studied. In large genome-wide association studies of AD, TOMM40 rs2075650 (intron 2, chromosome 19q13.32) has been one of the most significantly associated single-nucleotide polymorphisms (SNPs) (Harold et al, 2009; Naj et al, 2010; Seshadri et al, 2010). TOMM40 encodes a protein on the outer mitochondrial membrane essential for cell viability, thus variation in this gene could lead to mitochondrial dysfunction (Ferencz et al, 2012; Pfanner et al, 1997), of which dementia and depression are symptoms (Anglin et al, 2012; Fattal et al, 2007). TOMM40 rs2075650 is in linkage disequilibrium (LD) with the APOE ɛ4 rs429358 SNP (Harold et al, 2009; Potkin et al, 2009). However, evidence of an extended TOMM40-APOE haplotype that influences the regional effects and onset age of AD (Bekris et al, 2010; Roses, 2010) suggests that TOMM40 may have APOE-independent effects (Carrasquillo and Morgan, 2012). Even in relation to AD the risk associated with TOMM40 cannot be fully explained by LD, and it has been suggested that other SNPs must contribute (Bekris et al, 2012). Despite TOMM40 rs2075650 being a good candidate for major depressive disorder (MDD), its possible effect on depression-related phenotypes has not yet been investigated.
Pathophysiological similarities between MDD and AD provide candidate intermediate phenotypes that may identify shared genetic factors. Alterations in premorbid measures of personality factors have been demonstrated in both depression (Kendler et al, 2006a) and AD (Robins Wahlin and Byrne, 2011). Cognitive deficits are well-reported features of both MDD (Millan et al, 2012) and AD (Christensen et al, 1997) that share similar structural and functional brain abnormalities in regions such as hippocampus (Arnone et al, 2012a; Potkin et al, 2009; Shen et al, 2010), amygdala (Drevets et al, 2008; Shen et al, 2010), cingulate (Johnson et al, 2011; Pizzagalli, 2011; Ries et al, 2009), and dorsolateral prefrontal cortex (Morbelli et al, 2012; Pizzagalli, 2011). TOMM40 itself has also been associated with hippocampal atrophy (Potkin et al, 2009; Shen et al, 2010) in AD patients and reduced gray matter in the cingulate of healthy controls (Johnson et al, 2011). Variations in TOMM40 may therefore contribute to a biological vulnerability independent of APOE that underpins aspects of both AD and MDD.
In this study, we took an exploratory multiple-level hypothesis-testing approach, using intermediate phenotypes, to investigate the role of the minor (G) allele of TOMM40 rs2075650 in vulnerability to MDD. We tested the association between rs2075650, lifetime depression, and depressive symptoms in a large cohort and assessed whether personality traits are overexpressed in minor allele carriers. We then assessed whether the minor allele was associated with poorer cognitive function than the AA genotype using tests of memory, executive control, and affective bias. Finally, we investigated the effects of this SNP on brain function using a functional magnetic resonance imaging (fMRI) face emotion processing task as a putative neurobiological marker for depression (Juhasz et al, 2011), hypothesizing that risk allele carriers would show similar patterns of brain activations to those seen in currently depressed individuals.
MATERIALS AND METHODS
Participants
Participants aged 18–60 years, predominantly from Greater Manchester, were recruited primarily through NHS general practices and the study website. Detailed description of the recruitment methodology has been published previously (Juhasz et al, 2009; Juhasz et al, 2011). In summary, self-reported diagnostic and demographic data were collected from the Level-1 community sample, where a final cohort of 1220 participants was selected for the current investigation. All included participants were of Caucasian origin, provided DNA, and successfully genotyped for TOMM40 rs2075650, with no self-reported history of manic or hypomanic episodes, psychotic symptoms, or obsessive-compulsive disorder. From this initial pool, 129 participants completed a face-to-face clinical interview and an additional neuropsychological testing session. This sample was further enhanced with 114 participants recruited through advertisements, providing a total of 243 participants in the Level-2 cohort. Finally, a subset of 58 participants from Level-2 completed an fMRI scan at Level-3. Further details of the recruitment can be found in the Supplementary Information. All participants provided written informed consent, and the study was approved by the local ethics committees and carried out in accordance with the Declaration of Helsinki.
Level-1 Assessment
The Background Questionnaire included in the Level-1 questionnaire booklet has been detailed previously (Juhasz et al, 2009; Juhasz et al, 2011). Briefly, this questionnaire included items designed to probe personal psychiatric history, allowing for the coding of participants as either healthy controls or those who reported suffering from depression during their lifetime (see Supplementary Information). Self-reported lifetime depression was used to analyze genetic associations. The Brief Symptom Inventory was used to assess current psychiatric symptoms. Personality was measured using the Big Five Inventory (BFI-44) (John et al, 1991), a 44-item instrument that assesses traits of neuroticism, extraversion, openness to experience, agreeableness, and conscientiousness.
Level-2 Assessment
In addition to the information provided in the Level-1 background questionnaire, clinical diagnoses were established by trained researchers using the Structured Clinical Interview for DSM-IV (SCID-I/NP; First et al, 2002). Participants were grouped into controls (no lifetime psychiatric disorders), remitted depressed (past MDD currently in full remission), and currently depressed (current MDD or MDD in partial remission). Current depressive symptoms were measured using the Montgomery–Åsberg Depression Rating Scale (MADRS; Montgomery and Åsberg, 1979). Data from the diagnosed participants were used to validate the Level-1 questionnaire (see Juhasz et al, 2011).
The neuropsychological assessment included tasks known to be sensitive to depression (Elliott et al, 2011; Millan et al, 2012). The n-back task (Owen et al, 2005) was used as a measure of working memory, as previous work has demonstrated that this paradigm is sensitive to deficits in both AD and depression (Harvey et al, 2005; Waltz et al, 2004). The CANTAB Stockings of Cambridge (SoC; http://www.camcog.com/) task was used as an index of planning and executive function, as previous studies using the SoC have demonstrated executive deficits in depression (Beats et al, 1996; Elliott et al, 1996). Finally, an emotional word memory task, based on the work of Harmer et al (2009), was used as a measure sensitive to changes in affective bias and memory performance in depression. See Supplementary Information for details.
Level-3 Assessment
To assess whether changes in affective processing were associated with carrying the rs2075650 risk allele as distinct from current depressive mood, only healthy controls and remitted patients (with a MADRS score of⩽10) underwent fMRI scanning.
fMRI Image Acquisition
Participants were scanned using a 1.5 T Philips Intera while performing an emotional face-processing paradigm (Thomas et al, 2011). During this task, participants were asked to identify the gender of faces selected from the Ekman and Friesen (1976) stimuli displaying either a neutral expression, happiness, sadness, or fear (see Supplementary Information for details). Data were acquired using a T2*-weighted gradient echo-planar sequence with a repetition time (TR) of 2.1 s and an echo time (TE) of 40 ms. Each volume consisted of 29 contiguous axial slices (thickness 4.5 mm, inter-slice gap 0.5 mm). Voxel size was 3.5 × 3.5 × 5 mm3. A T1-weighted structural scan was acquired for preprocessing.
Genotyping
The TOMM40 rs2075650 SNP was genotyped using the Sequenom MassARRAY technology (Sequenom, San Diego). See Supplementary Information for information on genotyping.
Statistical Analysis
Level-1 genetic association analysis
PLINK v1.07 (http://pngu.mgh.harvard.edu/purcell/plink/) was used for testing Hardy–Weinberg equilibrium and associations of different genetic models (dominant (DOM), recessive (REC) and additive (ADD)), using linear and logistic regression between TOMM40 rs2075650 and phenotypes of interest (for power calculations, see Supplementary Information). Age and sex were covariates in all the analyses. Main effects of genotype were investigated using lifetime depression and current depression. Personality factors as possible intermediate phenotypes were tested by the main effects of genotype and by the genotype × depression status (current or lifetime) interaction. The positive false discovery rate (pFDR; q-value: http://genomics.princeton.edu/storeylab/qvalue/) was applied simultaneously across all models to maintain an α of 0.05 (Storey and Tibshirani, 2003).
Level-2 neuropsychological behavioral data analysis
The neuropsychological results were analyzed using SPSS 19 (http://www.ibm.com/software/analytics/spss/). Data were appropriately transformed to satisfy parametric assumptions. The 0-back condition of n-back was removed as performance was error-free. Age, sex, and current depression medication (as a binary indicator variable) were treated as covariates in all the models. To test the intermediate phenotype role of cognitive and affective domains, a single MANCOVA was performed for each outcome of each task, with the outcome vector comprising a linear combination of the task conditions. The multivariate main effects and interactions (using Pillai’s Trace) were used to infer effects of diagnosis, genotype, and diagnosis × genotype interactions. Significant diagnosis × genotype interactions were followed-up by assessing each of the individual task conditions as univariate between-subject ANOVA models in order to explore the direction of any interaction effects. Post-hoc testing was conducted on the univariate models using pairwise contrasts across the interaction term with the Šidák multiple-comparison correction. Due to the multiplicity inherent in testing multiple non-independent MANCOVA models, the same pFDR correction used in Level-1 was applied simultaneously across main effects and interactions. See Supplementary Information for details.
Level-3 imaging data analysis
The Level-3 imaging data were analyzed using Statistical Parametric Mapping (SPM8; http://www.fil.ion.ucl.ac.uk/spm/). Image preprocessing has been detailed previously (Thomas et al, 2011). At the first level, contrast maps of activation change were created by subtracting the neutral condition from each facial emotion of interest (happy, sad, and fear). For the second level analysis, each of the individual contrasts created in the first level were used as raw Y vector values in the linear model. The model was a standard factorial cell-means design constructed from genotype and diagnosis. Age, sex, and MADRS scores were entered as covariates, with the continuous measures mean-centered. Our initial analysis was also restricted to the sad-neutral contrast as the most reliable biomarker for MDD (Arnone et al, 2012b). Exploratory post-hoc analyses for happy-neutral and fear-neutral were also conducted.
Based on a priori hypotheses, region of interest (ROI) analyses of the areas common to MDD and AD discussed earlier (Arnone et al, 2012a; Drevets et al, 2008; Johnson et al, 2011; Macqueen and Frodl, 2011; Morbelli et al, 2012; Pizzagalli, 2011; Potkin et al, 2009; Ries et al, 2009; Shen et al, 2010) were conducted using the PickAtlas toolbox (http://fmri.wfubmc.edu/software/PickAtlas). All regions were included bilaterally in a single mask consisting of Brodmann areas (BA) 46, posterior cingulate cortex (PCC), anterior cingulate cortex (ACC), hippocampus, and amygdala. All results are reported at a ROI-corrected α of p(FWE)⩽0.05 using the MNI standard.
RESULTS
Demographic information for all three levels is presented in Table 1. Participants were similar in age and sex across all groups, with more females than males as expected. The distribution of the AA genotype and the G allele carriers (GA/GG) within each diagnostic group is also presented. TOMM40 rs2075650 SNP was in HWE in all levels and in all subgroups.
Level-1
Lifetime depression was significantly more frequent in G allele carriers in both additive and dominant models (Table 2). No relationship was shown for the recessive model. All further testing was conducted using a dominant model where GA and GG participants were collapsed into a single group due to the rarity (2% of the sample) of the GG genotype. For the remainder of the paper, our use of the G allele should be read as synonymous with GA and GG combined. G allele carriers did not have significantly greater current depression scores than non-carriers. Scores on the BFI-44-derived personality traits were not significantly affected by genotype. However, in those who suffered from current or past depression the possession of the risk G allele enhanced the low extraversion trait (significant genotype × lifetime depression/current depression scores interaction). There were no statistically significant interactions affecting other personality traits. A post-hoc analysis demonstrated that the association of lifetime depression and G allele carrier status was independent of any reported drug or alcohol use problems.
Level-2
Multivariate main effects, diagnosis × genotype interactions, and corrections for multiple testing (FDR q-values) are shown in Table 3. In summary, no significant multivariate main effects or diagnosis × genotype interactions were found for the n-back correct moves model or the SoC initial thinking time model. The SoC subsequent thinking time model showed no significant multivariate main effects but did indicate a significant multivariate diagnosis × genotype interaction. Follow-up univariate analysis of the individual task conditions indicated a significant interaction effect in the 2-move-solution condition (F(2, 202)=4.515, p=0.012) with the 4-move condition p-value only marginally<0.1 (F(2, 202)=2.599, p=0.077). However, none of the post-hoc contrasts across the interaction term survived correction.
The SoC average number of moves model also showed a significant multivariate diagnosis × genotype interaction with the follow-up univariate between-subject tests indicating significance in the 3-move-solution condition (F(2, 202)=5.630, p=0.004) and in the 4-move-solution condition (F(2, 202)=3.844, p=0.023). Šidák corrected post-hoc pairwise comparisons of the interaction term indicated significant differences between G allele carriers with current MDD and all other groups (all p⩽0.01) for the 3-move solution condition. This post-hoc univariate interaction result is shown in Figure 1.
Univariate results from the 3-move-solution condition of the Stockings of Cambridge (SoC) task detailing the direction of interaction for the mean number of moves between the two genotype categories and the three diagnostic groups. Error bars represent SE. *p⩽0.01.
For the emotional word memory task, a significant multivariate main effect of diagnosis was shown for the number of correctly recalled immediate words and the number of correctly recalled delayed words. There was a significant multivariate diagnosis × genotype interaction for the number of delayed intrusions (not surviving FDR). The follow-up univariate between-subject tests showed a significant interaction for the number of positive intrusions only (F(2, 225)=6.143, p=0.003). Šidák corrected post-hoc pairwise contrasts for the interaction term in the positive word condition indicated that G allele carriers with current MDD had fewer positive intrusions on delayed recall than AA carriers with current MDD (p<0.001), remitted G allele carriers (p=0.002), and control G carriers (p=0.026). This post-hoc univariate interaction result is shown in Figure 2.
Univariate results for the mean number of intrusive positive words recalled from the delayed word memory task, detailing the direction of the interaction effect between the two genotypes and three diagnostic categories. Error bars represent SE. *p⩽0.05.
Level-3
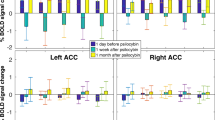
Analysis of covariance for the sad-neutral contrast indicated a significant ROI-corrected main effect of genotype in the left PCC (F(1,51)=17.34, p(FWE)=0.045) at −7 −46 20 and the left dorsal ACC (dACC; F(1,51)=17.62, p(FWE)=0.041) at −4 35 20. A corrected follow-up t-contrast indicated that greater responses were found for the AA genotype group compared with the G allele group in both regions (PCC: t(51)=4.20, p(FWE)=0.021; ACC: t(51)=4.16, p(FWE)=0.023) (Figure 3). No ROI-corrected clusters were found for the main effect of diagnosis. The genotype × diagnosis interaction also indicated no ROI-corrected clusters, suggesting that the effects were due to genotype alone (Figure 4). The cell sizes for the interaction term were: AA genotype+remitted MDD (n=18), AA genotype controls (n=23), G allele+remitted MDD (n=7), and G allele controls (n=10). Owing to the rarity of the minor allele, this result is tentative due to the small numbers in each group. Repeating the analysis using the happy-neutral contrast and the fear-neutral contrast revealed no significant ROI-corrected activations for either contrast and no diagnosis × genotype interaction. See Supplementary Information for details of medicated participants and power.
Peak voxel activation for the sad-neutral contrast of the emotional faces task in the AA and GA/GG genotype groups. Columns illustrate whole-brain mean percentage of signal change. Error bars represent 90% confidence intervals.
Peak voxel activation for the sad-neutral contrast of the emotional faces illustrating the interaction between diagnosis and rs2075650 genotype. Error bars represent 90% confidence intervals.
DISCUSSION
Our main finding is an association between the TOMM40 G allele and a history of depression. Using depression-related intermediate phenotypes, our results indicate that rs2075650 polymorphism does not increase the risk of depression via the expected neuroticism pathway but rather appears to interact with extraversion. This effect was not found globally at the behavioral level but only appeared within the context of depression, namely the risk allele carriers displayed decreased extraversion if they had lifetime or current depression. The TOMM40 risk allele also appeared to be associated with cognitive deficits during current depressive episodes, specifically mild executive dysfunction and a decrease in positive memory intrusions, and may therefore act by enhancing cognitive deficits within the most vulnerable group. Our imaging data identified functional differences between the protective and risk allele carriers that did not appear to be influenced by diagnosis. Instead of the predicted increase in activation in ROIs during sad emotion processing, we found a stable decrease of activity in the PCC and a state/medication-dependent deactivation in the dACC.
Decreased Extraversion and Depression
Both extraversion and neuroticism are key personality traits for affective processing (Canli, 2004). In fact, it has been claimed that 50% of the genetic vulnerability to depression is shared with genes that influence the expression of neuroticism (Juhasz et al, 2009), whereas extraversion is only weakly linked (Kendler et al, 2006b). Our results showed no interaction between TOMM40 and depression on neuroticism but a significant finding for decreased extraversion. TOMM40 may therefore act as a particular mediator of the extraversion trait alone, reducing expression in carriers of the risk allele with either a history of, or current, depressive episodes. This is compatible with a number of studies implicating low extraversion as a risk factor for developing depression (Cox et al, 2004; Farmer et al, 2002; Jylha and Isometsa, 2006; Jylha et al, 2009) although the association is not as strong as with neuroticism. It may be that even mildly decreased extraversion is important during depressive episodes (Jylha et al, 2009) where it might inhibit recovery. Further study will be needed to clarify this relationship. In addition, given the established relationship between TOMM40 and AD, it is interesting to note that a decrease in extraversion, as well as increases of neuroticism, have been found to precede and represent risk factors for AD and thus may be markers of early dementia onset (Robins Wahlin and Byrne, 2011).
Executive Dysfunction and a Reduced Positive Memory Bias
Poor performance during the SoC task has been previously demonstrated in depressed patients (Beats et al, 1996; Elliott et al, 1996), a finding often interpreted as a top-down, executive control deficit (Disner et al, 2011). Our finding of some influence of the TOMM40 risk allele on SoC performance in currently depressed, but not remitted depressed, individuals suggests that there may be a state-dependent effect of the risk allele on executive dysfunction. Although the effect was modest, and therefore caution is needed, it is of interest that impaired cognitive function, in addition to low extraversion, has been associated with poorer treatment outcome in depression (Clark et al, 2009), consistent with a potential role for the TOMM40 risk allele as a marker of worse illness course.
Negative bias is central to cognitive models of depression where positive bias associated with normal processing of affective stimuli is reduced (Elliott et al, 2011; Ellwart et al, 2003; Harmer et al, 2009). In our study, the bias towards the intrusive recall of positive words was seen in all the groups apart from currently depressed individuals carrying the risk allele. The risk allele may therefore play a part in the biasing of the retrieval of affective material from memory (Hamilton and Gotlib, 2008).
Neuronal Differences in Cingulate Activity
Using fMRI during face emotion processing, an imaging marker for depression (Scharinger et al, 2010), we found that possession of the risk allele, independent of a history of depression, was associated with altered function of the cingulate cortex during sad emotion processing. Given the finding was in both remitted depressed participants and controls, it appears that the risk allele is associated with a general alteration in affective processing. In depression, both morphological changes (Caetano et al, 2006) and dysfunction of the dACC (Fu et al, 2004) have been demonstrated and implicated as predictors of treatment response (Fu et al, 2008; Pizzagalli et al, 2001; Pizzagalli, 2011). Dysfunction of the dACC is consistent with theories of impaired top-down control in depressed individuals (Disner et al, 2011). The PCC is associated with affective self-referential processing (Johnson et al, 2009; Vogt et al, 2006), and abnormal responses in this region have been reported in depressed subjects (Berman et al, 2011; Drevets, 2000). One possible interpretation of our results is that the presence of a trait alteration in function in the cingulate cortex, through carrying the risk allele, may interact with those occurring during depression. In particular, the effect in the dACC may contribute to impaired recovery from depression. Of interest also, given the putative link between depression and AD, is that one of the earliest detectable hypo-metabolic neuronal regions in AD is the PCC (Minoshima et al, 1997), with the TOMM40 risk allele known to impact brain regions vulnerable to AD by downstream apoptotic processes (Ferencz et al, 2012). These findings suggest that a potential mechanism for depression as a precursor to AD may be via dysfunctional activity within the cingulate.
Limitations
The main limitation of the current study was our focus on a single SNP. Although significant associations were found, it is difficult to know how much of an effect this polymorphism has without further understanding the influence of other mutations within the TOMM40-APOE locus. Although the Level-1 cohort was a large sample, the nature of questionnaire data and subsequent response biases means that further replication will be required to support our findings, particularly as the diagnosis of depression was self-reported. In our Level-2 findings, the main effect of emotional word bias did not survive FDR correction, thus this result will need replication. In addition, the sample size for the Level-3 imaging study was modest by genetic standards and as such replication of this result will be required. Also, only remitted depressed and healthy individuals were imaged, and therefore we cannot generalize our findings to a currently depressed state.
Summary
The current study presents evidence of a possible new risk allele for the development of depression. Our results indicate that possession of the TOMM40 G allele is associated with lifetime risk of depression and with altered neural processing of sad faces shown by fMRI. In addition, current depressive states revealed further genotypic associations with impaired cognitive performance, changes in positively biased affective recall, and a decrease in extraverted personality traits. Extrapolation of these results beyond the current study remains speculative given that this is the first study to investigate the link between depression and TOMM40 rs2075650. It may be that the risk allele distorts the development of neural systems processing executive function and emotion and that therefore the effects of the TOMM40 G allele might be expected not only to increase risk of depression but also to prolong illness. This is currently unknown and would be worthy of further investigation. It is intriguing to consider that the same underlying alterations in neural system function may also contribute to vulnerability for the development of dementia, potentially explaining the enhanced risk of AD in individuals who have suffered depression in the past, and the reduced age of onset for AD in these vulnerable populations. Further investigations will be needed in order to fully understand the relationship between TOMM40, dementia, and depression.
FUNDING AND DISCLOSURE
The study was supported by the Sixth Framework Program of the European Union, NewMood, LSHM-CT-2004-503474, by the National Institute for Health Research Manchester Biomedical Research Centre, and by the TAMOP-4.2.1.B-09/1/KMR-2010-0001, Hungary. MMF was supported by an MRC Studentship. RE received consultancy fees from Cambridge Cognition and P1vital. JFWD variously performed consultancy, speaking engagements, and research for Bristol-Myers Squibb, AstraZeneca, Eli Lilly, Schering Plough, Janssen-Cilag, and Servier (all fees are paid to the University of Manchester to reimburse them for the time taken); he has share options in P1vital. IMA has received consultancy fees from Servier and Alkermes, an honorarium for speaking from Lundbeck, and grant support from Servier and AstraZeneca. All the other authors report no biomedical financial interests or potential conflicts of interest.
References
Anglin RE, Garside SL, Tarnopolsky MA, Mazurek MF, Rosebush PI (2012). The psychiatric manifestations of mitochondrial disorders: a case and review of the literature. J Clin Psychiatry 73: 506–512.
Arnone D, McKie S, Elliott R, Juhasz G, Thomas EJ, Downey D et al (2012a). State-dependent changes in hippocampal grey matter in depression. Mol Psychiatry 12: 1265–1272.
Arnone D, McKie S, Elliott R, Thomas EJ, Downey D, Juhasz G et al (2012b). Increased amygdala responses to sad but not fearful faces in major depression: relation to mood state and pharmacological treatment. Am J Psychiatry 169: 841–850.
Barnes DE, Yaffe K, Byers AL, McCormick M, Schaefer C, Whitmer RA (2012). Midlife vs late-life depressive symptoms and risk of dementia. Arch Gen Psychiatry 69: 493–498.
Beats BC, Sahakian BJ, Levy R (1996). Cognitive performance in tests sensitive to frontal lobe dysfunction in the elderly depressed. Psychol Med 26: 591–603.
Bekris LM, Galloway NM, Montine TJ, Schellenberg GD, Yu CE (2010). APOE mRNA and protein expression in postmortem brain are modulated by an extended haplotype structure. Am J Med Genet B Neuropsychiatr Genet 153B: 409–417.
Bekris LM, Lutz F, Yu CE (2012). Functional analysis of APOE locus genetic variation implicates regional enhancers in the regulation of both TOMM40 and APOE. J Hum Genet 57: 18–25.
Berman MG, Peltier S, Nee DE, Kross E, Deldin PJ, Jonides J (2011). Depression, rumination and the default network. Soc Cogn Affect Neurosci 6: 548–555.
Butters MA, Sweet RA, Mulsant BH, Ilyas Kamboh M, Pollock BG, Begley AE et al (2003). APOE is associated with age-of-onset, but not cognitive functioning, in late-life depression. Int J Geriatr Psychiatry 18: 1075–1081.
Caetano SC, Kaur S, Brambilla P, Nicoletti M, Hatch JP, Sassi RB et al (2006). Smaller cingulate volumes in unipolar depressed patients. Biol Psychiatry 59: 702–706.
Canli T (2004). Functional brain mapping of extraversion and neuroticism: learning from individual differences in emotion processing. J Pers 72: 1105–1132.
Carrasquillo MM, Morgan K (2012). Commentary on ‘Functional analysis of APOE locus genetic variation implicates regional enhancers in the regulation of both TOMM40 and APOE’. J Hum Genet 57: 3–4.
Cherbuin N, Leach LS, Christensen H, Anstey KJ (2007). Neuroimaging and APOE genotype: a systematic qualitative review. Dement Geriatr Cogn Disord 24: 348–362.
Christensen H, Griffiths K, MacKinnon A, Jacomb P (1997). A quantitative review of cognitive deficits in depression and Alzheimer-type dementia. J Int Neuropsychol Soc 3: 631–651.
Clark L, Chamberlain SR, Sahakian BJ (2009). Neurocognitive mechanisms in depression: implications for treatment. Annu Rev Neurosci 32: 57–74.
Cox BJ, McWilliams LA, Enns MW, Clara IP (2004). Broad and specific personality dimensions associated with major depression in a nationally representative sample. Compr Psychiatry 45: 246–253.
Disner SG, Beevers CG, Haigh EA, Beck AT (2011). Neural mechanisms of the cognitive model of depression. Nat Rev Neurosci 12: 467–477.
Drevets WC (2000). Neuroimaging studies of mood disorders. Biol Psychiatry 48: 813–829.
Drevets WC, Price JL, Furey ML (2008). Brain structural and functional abnormalities in mood disorders: implications for neurocircuitry models of depression. Brain Struct Funct 213: 93–118.
Ekman P, Friesen WV (1976) Pictures of Facial Affect. Consulting Psychologists Press: Palo Alto, CA, USA.
Elliott R, Sahakian BJ, McKay AP, Herrod JJ, Robbins TW, Paykel ES (1996). Neuropsychological impairments in unipolar depression: the influence of perceived failure on subsequent performance. Psychol Med 26: 975–989.
Elliott R, Zahn R, Deakin JF, Anderson IM (2011). Affective cognition and its disruption in mood disorders. Neuropsychopharmacol 36: 153–182.
Ellwart T, Rinck M, Becker ES (2003). Selective memory and memory deficits in depressed inpatients. Depress Anxiety 17: 197–206.
Farmer AE, Redman K, Harris T, Mahmood A, Sadler S, Pickering A et al (2002). Neuroticism, extraversion, life events and depression. Br J Psychiatry 181: 118–122.
Fattal O, Link J, Quinn K, Cohen BH, Franco K (2007). Psychiatric comorbidity in 36 adults with mitochondrial cytopathies. CNS Spectr 12: 429–438.
Ferencz B, Karlsson S, Kalpouzos G (2012). Promising genetic biomarkers of preclinical Alzheimer’s disease: the influence of APOE and TOMM40 on brain integrity. Int J Geriatr Psychiatry 2012: 421452.
First MB, Spitzer RL, Gibbon M, Williams JBW (2002) Structured Clinical Interview for DSM-IV-TR Axis I Disorders, Research Version (SCID-I). State Psychiatric Institute: New York, NY, USA.
Fu CH, Williams SC, Cleare AJ, Scott J, Mitterschiffthaler MT, Walsh ND et al (2008). Neural responses to sad facial expressions in major depression following cognitive behavioral therapy. Biol Psychiatry 64: 505–512.
Fu CHY, Williams SCR, Cleare AJ, Brammer MJ, Walsh ND, Kim J et al (2004). Attenuation of the neural response to sad faces in major depression by antidepressant treatment. Arch Gen Psychiatry 61: 877–889.
Green RC, Cupples A, Kurz A, Auerbach S, Go R, Sadovnick D et al (2003). Depression as a risk factor for Alzheimer disease: the MIRAGE study. Arch Neurol 60: 753–759.
Greenwood PM, Sunderland T, Friz JL, Parasuraman R (2000). Genetics and visual attention: Selective deficits in healthy adult carriers of the E4 allele of the apolipoprotein E gene. Proc Natl Acad Sci USA 97: 11661–11666.
Hamilton JP, Gotlib IH (2008). Neural substrates of increased memory sensitivity for negative stimuli in major depression. Biol Psychiatry 63: 1155–1162.
Harmer CJ, O’Sullivan U, Favaron E, Massey-Chase R, Ayres R, Reinecke A et al (2009). Effect of acute antidepressant administration on negative affective bias in depressed patients. Am J Psychiatry 166: 1178–1184.
Harold D, Abraham R, Hollingworth P, Sims R, Gerrish A, Hamshere ML et al (2009). Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat Genetics 41: 1088–1093.
Harvey PO, Fossati P, Pochon JB, Levy R, Lebastard G, Lehericy S et al (2005). Cognitive control and brain resources in major depression: an fMRI study using the n-back task. Neuroimage 26: 860–869.
Heise V, Filippini N, Ebmeier KP, Mackay CE (2011). The APOE ɛ4 allele modulates brain white matter integrity in healthy adults. Mol Psychiatry 16: 908–916.
Hong MG, Alexeyenko A, Lambert JC, Amouyel P, Prince JA (2010). Genome-wide pathway analysis implicates intracellular transmembrane protein transport in Alzheimer disease. J Hum Genet 55: 707–709.
John OP, Donahue EM, Kentle RL (1991) The Big Five Inventory: Versions 4a and 54. Technical report. University of California, Insititute of Personality and Social Research: Berkeley, CA, USA.
Johnson MK, Nolen-Hoeksema S, Mitchell KJ, Levin Y (2009). Medial cortex activity, self-reflection and depression. Soc Cogn Affect Neurosci 4: 313–327.
Johnson SC, La Rue A, Hermann BP, Xu G, Koscik RL, Jonaitis EM et al (2011). The effect of TOMM40 poly-T length on gray matter volume and cognition in middle-aged persons with APOE epsilon3/epsilon3 genotype. Alzheimers Dement 7: 456–465.
Jorm AF (2001). History of depression as a risk factor for dementia: an updated review. Aust N Z J Psychiatry 35: 776–781.
Juhasz G, Chase D, Pegg E, Downey D, Toth ZG, Stones K et al (2009). CNR1 gene is associated with high neuroticism and low agreeableness and interacts with recent negative life events to predict current depressive symptoms. Neuropsychopharmacol 34: 2019–2027.
Juhasz G, Dunham JS, McKie S, Thomas E, Downey D, Chase D et al (2011). The CREB1-BDNF-NTRK2 pathway in depression: multiple gene-cognition-environment interactions. Biol Psychiatry 69: 762–771.
Jylha P, Isometsa E (2006). The relationship of neuroticism and extraversion to symptoms of anxiety and depression in the general population. Depress Anxiety 23: 281–289.
Jylha P, Melartin T, Rytsala H, Isometsa E (2009). Neuroticism, introversion, and major depressive disorder—traits, states, or scars? Depress Anxiety 26: 325–334.
Kendler KS, Gatz M, Gardner CO, Pedersen NL (2006a). Personality and major depression. Arch Gen Psychiatry 63: 1113–1120.
Kendler KS, Gatz M, Gardner CO, Pedersen NL (2006b). Personality and major depression: a Swedish Longitudinal, Population-Based Twin Study. Arch Gen Psychiatry 63: 1113–1120.
Li H, Wetten S, Li L, St. Jean PL, Upmanyu R, Surh L et al (2008). Candidate single-nucleotide polymorphisms from a genomewide association study of Alzheimer disease. Arch Neurol 65: 45–53.
Macqueen G, Frodl T (2011). The hippocampus in major depression: evidence for the convergence of the bench and bedside in psychiatric research? Mol Psychiatry 16: 252–264.
Millan MJ, Agid Y, Brune M, Bullmore ET, Carter CS, Clayton NS et al (2012). Cognitive dysfunction in psychiatric disorders: characteristics, causes and the quest for improved therapy. Nat Rev Drug Discov 11: 141–168.
Minoshima S, Giordani B, Berent S, Frey KA, Foster NL, Kuhl DE (1997). Metabolic reduction in the posterior cingulate cortex in very early Alzheimer’s disease. Ann Neurol 42: 85–94.
Montgomery SA, Åsberg M (1979). A new depression scale designed to be sensitive to change. Br J Psychiatry 134: 382–389.
Morbelli S, Drzezga A, Perneczky R, Frisoni GB, Caroli A, van Berckel BN et al (2012). Resting metabolic connectivity in prodromal Alzheimer's disease. A European Alzheimer Disease Consortium (EADC) project. Neurobiol Aging 33: 2533–2550.
Naj AC, Beecham GW, Martin ER, Gallins PJ, Powell EH, Konidari I et al (2010). Dementia revealed: novel chromosome 6 locus for late-onset Alzheimer disease provides genetic evidence for folate-pathway abnormalities. PloS Genet 6: 1–10.
Owen AM, McMillan KM, Laird AR, Bullmore E (2005). N-back working memory paradigm: a meta-analysis of normative functional neuroimaging studies. Hum Brain Mapp 25: 46–59.
Ownby RL, Crocco E, Acevedo A, John V, Loewenstein D (2006). Depression and risk for Alzheimer disease. Arch Gen Psychiatry 63: 530–538.
Pfanner N, Craig EA, Hönlinger A (1997). Mitochondrial preprotein translocase. Annu Rev Cell Dev Biol 13: 25–51.
Pizzagalli D, Pascual-Marqui RD, Nitschke JB, Oakes TR, Larson CL, Abercrombie HC et al (2001). Anterior cingulate activity as a predictor of degree of treatment response in major depression: evidence from brain electrical tomography analysis. Am J Psychiatry 158: 405–415.
Pizzagalli DA (2011). Frontocingulate dysfunction in depression: toward biomarkers of treatment response. Neuropsychopharmacol 36: 183–206.
Potkin SG, Guffanti G, Lakatos A, Turner JA, Kruggel F, Fallon JH et al (2009). Hippocampal atrophy as a quantitative trait in a genome-wide association study identifying novel susceptibility genes for Alzheimer's disease. PLoS One 4: e6501.
Ries ML, Wichmann A, Bendlin BB, Johnson SC (2009). Posterior cingulate and lateral parietal gray matter volume in older adults with depressive symptoms. Brain Imaging Behav 3: 233–239.
Robins Wahlin TB, Byrne GJ (2011). Personality changes in Alzheimer’s disease: a systematic review. Int J Geriatr Psychiatry 26: 1019–1029.
Roses AD (2010). An inherited variable poly-T repeat genotype in TOMM40 in Alzheimer disease. Arch Neurol 67: 536–541.
Saczynski JS, Beiser A, Seshadri S, Auerbach S, Wolf PA, Au R (2010). Depressive symptoms and risk of dementia: the Framingham heart study. Neurology 75: 35–41.
Scharinger C, Rabl U, Sitte HH, Pezawas L (2010). Imaging genetics of mood disorders. Neuroimage 53: 810–821.
Schiepers OJ, Harris SE, Gow AJ, Pattie A, Brett CE, Starr JM et al (2011). APOE E4 status predicts age-related cognitive decline in the ninth decade: longitudinal follow-up of the Lothian Birth Cohort 1921. Mol Psychiatry 17: 315–324.
Seshadri S, Fitzpatrick AL, Ikram MA, DeStefano AL, Gudnason V, Boada M et al (2010). Genome-wide analysis of genetic loci associated with Alzheimer disease. JAMA 303: 1832–1840.
Shen L, Kim S, Risacher SL, Nho K, Swaminathan S, West JD et al (2010). Whole genome association study of brain-wide imaging phenotypes for identifying quantitative trait loci in MCI and AD: a study of the ADNI cohort. Neuroimage 53: 1051–1063.
Storey JD, Tibshirani R (2003). Statistical significance for genomewide studies. Proc Natl Acad Sci USA 100: 9440–9445.
Thomas EJ, Elliott R, McKie S, Arnone D, Downey D, Juhasz G et al (2011). Interaction between a history of depression and rumination on neural response to emotional faces. Psychol Med 41: 1845–1855.
Vogt BA, Vogt L, Laureys S (2006). Cytology and functionally correlated circuits of human posterior cingulate areas. Neuroimage 29: 452–466.
Waltz JA, Knowlton BJ, Holyoak KJ, Boone KB, Back-Madruga C, McPherson S et al (2004). Relational integration and executive function in Alzheimer's disease. Neuropsychol 18: 296–305.
Yen Y-C, Rebok GW, Gallo JJ, Yang M-J, Lung F-W, Shih C-H (2007). ApoE4 allele is associated with late-life depression: a population-based study. Am J Geriatr Psychiatry 15: 858–868.
Acknowledgements
The sponsors funded the work but had no further role in the design or conduct of the study or in the preparation, review, or approval of the manuscript. Diana Chase, PhD, and Kathryn Lloyd-Williams, BSc, assisted with recruitment and data acquisition. We are grateful to Heaton Mersey Medical Practice and Cheadle Medical Practice for their assistance in the recruitment. For all their assistance, we thank the staff of the Wellcome Trust Clinical Research Facility, Manchester, where the fMRI research was performed.
Author contributions
GJ and MMF had full access to all the data in the study, performed the statistical analysis, and GJ takes responsibility for the integrity of the data and the accuracy of the data analysis. All other authors contributed to the wording, content, and construction of the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Neuropsychopharmacology website
Supplementary information
Rights and permissions
About this article
Cite this article
McFarquhar, M., Elliott, R., McKie, S. et al. TOMM40 rs2075650 May Represent a New Candidate Gene for Vulnerability to Major Depressive Disorder. Neuropsychopharmacol 39, 1743–1753 (2014). https://doi.org/10.1038/npp.2014.22
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2014.22
Keywords
This article is cited by
-
Neuroimaging Genomics a Predictor of Major Depressive Disorder (MDD)
Molecular Neurobiology (2023)
-
TOMM40 and APOE variants synergistically increase the risk of Alzheimer’s disease in a Chinese population
Aging Clinical and Experimental Research (2021)
-
Distinct effects of folate pathway genes MTHFR and MTHFD1L on ruminative response style: a potential risk mechanism for depression
Translational Psychiatry (2016)
-
A mitochondrial bioenergetic basis of depression
Journal of Bioenergetics and Biomembranes (2015)