Abstract
As previous work has highlighted the significance of the cannabinoid receptor 1 (CNR1) and fatty acid amide hydrolase (FAAH) genes with respect to cannabis dependence (CD), this study sought to characterize the neural mechanisms that underlie these genetic effects. To this end, we collected DNA samples and fMRI data using a cue-elicited craving paradigm in thirty-seven 3-day-abstinent regular marijuana users. The participants were grouped according to their genotype on two single-nucleotide polymorphisms (SNPs) earlier associated with CD phenotypes: rs2023239 in CNR1 and rs324420 in FAAH. Between-group comparisons showed that carriers of the CNR1 rs2023239 G allele had significantly greater activity in reward-related areas of the brain, such as the orbitofrontal cortex (OFC), inferior frontal gyrus (IFG), and anterior cingulate gyrus (ACG), during exposure to marijuana cues, as compared with those with the A/A genotype for this SNP. The FAAH group contrasts showed that FAAH rs324420 C homozygotes also had greater activation in widespread areas within the reward circuit, specifically in the OFC, ACG, and nucleus accumbens (NAc), as compared with the FAAH A-allele carriers. Moreover, there was a positive correlation between neural response in OFC and NAc and the total number of risk alleles (cluster-corrected p<0.05). These findings are in accord with earlier reported associations between CNR1 and FAAH and CD intermediate phenotypes, and suggest that the underlying mechanism of these genetic effects may be enhanced neural response in reward areas of the brain in carriers of the CNR1 G allele and FAAH C/C genotype in response to marijuana cues.
Similar content being viewed by others
INTRODUCTION
The main psychoactive compound in marijuana, delta-9-tetrahydrocannabinol (Δ9-THC), binds to central cannabinoid, or CB1, receptors, in which it mimics the effects of endogenously produced cannabinoids. The administration of CB1 antagonists in mice results in a decrease in reward behavior in response to cannabinoids and other substances of abuse (Arnone et al, 1997; Berrendero et al, 2003; Castane et al, 2002; Ledent et al, 1999), and the administration of the antagonist SR141716A (Rimonabant) extinguishes reward-related behaviors such as conditioned place preference and self-administration suggesting that CB1 activation modulates these behaviors (Gardner et al, 2002). In the first study of cue- and drug-induced reinstatement of cannabinoid-seeking in non-human primates, it was found that continuous administration of rimonabant, but not naltrexone, decreased cue-induced drug seeking, THC-induced drug seeking, and the direct reinforcing effects of THC in squirrel monkeys (Justinova et al, 2003, 2008). Moreover, single-cell recordings in the ventral tegmental area (VTA), the origin of dopaminergic cell bodies, have shown that Δ9-THC increases neuronal firing rates in this area (Cheer et al, 2000). More interestingly, increased dopamine (DA) neuronal firing rates are coupled with increased DA neuronal bursts, and these effects are blocked by SR141716A (Diana et al, 1998; French et al, 1997). These findings suggest that cannabinoids increase DA activity in the NAc and prefrontal cortex (PFC) by activating CB1 receptors in the VTA, which increase DA neuronal firing and burst rates. In other words, CB1 receptors increase DA activity by local disinhibitory mechanisms. The gene that encodes for CB1, cannabinoid receptor 1 (CNR1), thus likely modulates endocannabinoid and DA-mediated reward signaling; consequently, it has attracted substantial attention in the search for genetic mediators of liability to substance use disorders (SUD).
Among human drug users, CNR1 variants have been associated with both SUD phenotypes generally (eg, Ballon et al, 2006; Comings et al, 1997; Covault et al, 2001; Herman et al, 2006; Racz et al, 2003; Schmidt et al, 2002; Zhang et al, 2004; Zuo et al, 2007, 2009) and cannabis dependence (CD) specifically (Agrawal and Lynskey, 2009; Hopfer et al, 2006), although some groups have also reported null findings for this gene (eg, Covault et al, 2001; Hartman et al, 2009; Li et al, 2000). A report by Zhang et al (2004) suggest that one variant, a G to A single-nucleotide polymorphism (SNP) in the third exon, may create an alternative CNR1 transcript in the brain, and this SNP is associated with general liability to substance abuse (Zhang et al, 2004). Our group has also reported an association between this SNP (rs2023239) and differences in two intermediate CD phenotypes: withdrawal and craving after marijuana abstinence (Haughey et al, 2008). Specifically, withdrawal and craving among 105 young adults who reported smoking marijuana daily were assessed at baseline and again after 5 days of abstinence. The G allele of rs2023239 showed a significant abstinence by genotype interaction on withdrawal, as well as a main effect on overall levels of craving. However, results for this SNP have not been uniformly consistent; using a case–control design, Hartman et al, 2009 did not find an association between rs2023239 variance and number of CD symptoms in an adolescent sample.
Although significant effort has been devoted to analyzing CNR1 associations with CD, another critical gene in the endocannabinoid system, FAAH, has also shown associations with CD phenotypes. FAAH encodes for fatty acid amide hydrolase (FAAH), an enzyme expressed in the brain and liver that inactivates N-arachidonoyl-ethanolamine (anandamide), an endogenous CB1 agonist (Cravatt and Lichtmann, 2003). FAAH knockout mice (FAAH (−/−)) show exaggerated responses to anandamide, suggesting that FAAH is the primary regulator of anandamide signaling (Cravatt et al, 2001). Decreases in FAAH efficacy may increase sensitivity to anandamide (Cravatt et al, 2001), thereby increasing CB1 binding and affecting DA activity along reward pathways. Alterations in endocannabinoid signaling have been associated with a variety of SUD phenotypes in animal models (Wiskerke et al, 2008). Although no common human mutation causes FAAH deletion, a missense SNP, C385A (rs324420), results in a mutant form of FAAH with reduced expression and cellular stability (Chiang et al, 2004). Among human marijuana users, the C allele of this SNP has been associated with increased risk for progression to CD (Tyndale et al, 2007). Our study of young marijuana users also reported an association between the C allele and craving after marijuana abstinence (Haughey et al, 2008). In a follow-up study, we administered marijuana to 40 young adult daily users after 24 h of abstinence, and found that the C allele was associated with increased severity of withdrawal symptoms after abstinence, and increased happiness after smoking marijuana (Schacht et al, 2009).
In sum, the existing literature provides strong evidence for the relevance of CNR1 and FAAH with respect to SUD and CD phenotypes. However, much of this work has been conducted with dichotomous diagnostic phenotypes. This study aimed to use an imaging genomics approach (eg, Hariri and Weinberger, 2003) to examine the effects of variance in these genes on marijuana cue-elicited brain activation, a finer-grained phenotype that is arguably more proximal to the downstream effects of CNR1 and FAAH variance. As we have reported earlier that cue reactivity in chronic heavy marijuana users is associated with greater blood oxygenated level dependent (BOLD) response in reward-related areas of the brain (Filbey et al, 2009) and that among heavy drinkers, genetic variants, including rs2023239 in CNR1, modulate reactivity to the taste of alcohol in the form of increased activation (Filbey et al, 2008; Hutchison et al, 2008), we hypothesized that genetic risk, specifically, the G allele of rs2023239 in CNR1 and the C allele of rs324420 in FAAH, would be associated with an increased BOLD response to marijuana-related cues in reward-associated brain areas.
MATERIALS AND METHODS
Participants
For this study, we re-analyzed earlier reported data by genotype (Filbey et al, 2009). The earlier described participants were 43 self-reported regular marijuana users who were recruited through flyers and media advertisement in the Albuquerque, NM metro area and provided informed consent to participate in this study. Of these, six had movement >2 mm and were excluded from the analyses. All participants were right handed and free of MRI contraindications (ie, no metallic implants, claustrophobia, pregnancy, and so on). The participants included in this study were required to speak English, to report regular marijuana use at least four times per week over the previous 6 months, and to be willing to abstain from marijuana use for 3 days. Participants were compensated for their participation.
DNA was collected from buccal cells and extracted according to modified standard procedures (Walker et al, 1999). Samples were genotyped on the CNR1 and FAAH polymorphisms (CNR1/rs2023239 and FAAH/rs324420) using Taqman PCR technology. Participants were grouped according to their CNR1 and FAAH genotypes separately. As there were insufficient numbers of individuals who were homozygous for the rare allele for both SNPs, individuals who were homozygous for the rare allele were combined with the heterozygotes for the statistical analyses. For CNR1, because there were no homozygotes for the G allele, the heterozygotes (n=10) were compared with the A/A individuals (n=24); for FAAH, A/A individuals (n=5) were combined with heterozygotes (n=15), and compared with C/C individuals (n=17). Table 1 describes the characteristics of the 37 participants included in this study.
Procedure
The study took place in two sessions. During the first session, participants provided a saliva sample for DNA analysis and a urine sample for toxicological analysis. Participants were then scheduled for a second session, and were instructed to abstain from marijuana use for 72 h before this session. They were informed that a second urine drug screen administered at this session would confirm their abstinence. Although toxicological analysis was not sufficiently sensitive to detect abstinence-induced changes in urine levels of THC metabolites, bogus pipeline procedures have shown efficacy in increasing the accuracy of self-reports of drug use (eg, (Lowe et al, 1986)). During the second session, participants also completed a battery of neuropsychological tests (not reported here) and self-report measures of mood and craving. Participants were then placed in the MRI scanner. After collecting a high-resolution anatomical scan for registration and localization of the fMRI data, participants completed stop-signal and monetary incentive delay tasks (not reported here). Participants were then administered a cue-elicited craving paradigm, described below. The cue paradigm was the last task completed during a 105-m scanning session.
MRI images were collected using a 3 Tesla Siemens Trio. fMRI scans were collected using a gradient echo, echoplanar sequence with ramp sampling correction using the intercommissural line (anterior commissure/posterior commissure) as a reference (repetition time: 2.0 s, echo time: 27 ms, α: 70°, matrix size: 64 × 64, 32 slices, voxel size: 3 × 3 × 4 mm3). A tilting acquisition was applied during the echoplanar imaging (EPI) sequence to compensate for the problems of B0 field spatial distortion in the orbitofrontal cortex (OFC). Slices were acquired higher than the anterior commissure/posterior commissure, approximately perpendicular to the sinuses (Deichmann et al, 2003; Weiskopf et al, 2007). The high-resolution anatomical MRI scan was collected with a multi-echo magnetization prepared rapid acquisition gradient echo (MPRAGE) sequence with the following parameters: repetition time/echo time/inversion recovery time=2300/2.74/900 ms, flip angle=8°, field of view=256 × 256 mm, slab thickness=176 mm, matrix=256 × 256 × 176, voxel size=1 × 1 × 1 mm, number of echos=4, pixel bandwidth=650 Hz, total scan time=6 min.
We used a previously described marijuana tactile cue-exposure paradigm (Filbey et al, 2009). Briefly, the paradigm was presented in two separate EPI runs of 12 pseudorandom tactile presentations of a marijuana pipe (marijuana cue × six trials) and a pencil (control cue × six trials). Each trial consisted of a 20-s cue exposure period, followed by a single 5-s urge question, and ended with a 20-s washout period to allow the hemodynamic response to return to baseline before the next trial. The total number of repetitions per run was 288 and the total task duration was 19 min and 12 s. The task was presented using a front projection to a mirror system mounted on the head coil. Responses were recorded using a fiber-optic pad. Stimulus presentations were delivered using E-Prime (Psychology Software Tools). The timing of the stimulus presentation was synchronized with trigger pulses from the magnet to ensure precise temporal integration of stimulus presentation and fMRI data acquisition. The University of New Mexico Human Research Review Committee approved all procedures used.
Analyses
Pre-processing of fMRI data followed a standard procedure. The images were realigned using INRIalign, a motion correction algorithm unbiased by local signal changes (Freire and Mangin, 2001; Freire et al, 2002). Next, using FEAT (fMRI Expert Analysis Tool) Version 5.98, part of FSL (fMRIB's Software Library, http://www.fmrib.ox.ac.uk/fsl/), the following pre-statistics processing was performed: non-brain tissue/skull removal using Brain Extraction Tool (BET); spatial smoothing using a Gaussian kernel of FWHM 8 mm3; mean-based intensity normalization of all volumes by the same factor; and high-pass temporal filtering (Gaussian-weighted least-squares straight line fitting, with sigma=50.0 s). Time-series statistical analysis was carried out using FILM (FMRIB's Improved Linear Model) with local autocorrelation correction. The first seven volumes of all EPI runs were discarded to allow the MR signal to reach steady state.
Explanatory variables were created by convolving the stimulus timing files with a double gamma hemodynamic response function in FEAT. A multiple linear regression analysis was performed to estimate the hemodynamic parameters for different explanatory variables (ie, active condition for marijuana cues, active condition for control cues) and a corresponding t-statistic indicates the significance of the activation of the stimulus. Contrast maps were created by contrasting marijuana active conditions vs control active conditions. These maps were then registered to a high-resolution image using FLIRT (FMRIB's Linear Image Registration Tool) (Jenkinson and Smith, 2001; Jenkinson et al, 2002). Group analyses were carried out using FLAME (FMRIB's Local Analysis of Mixed Effects) (Beckmann et al, 2003; Woolrich et al, 2004). Statistical maps were then registered to the Montreal Neurological Institute (MNI) template with a two-step process. First, EPI images were registered to the high-resolution MPRAGE image, which was subsequently registered to the 152 brain average MNI template. These registration steps were performed using FLIRT. After transformation of the masks into MNI space, higher-level analysis was carried out using FLAME. We set our threshold and multiple comparison correction using FEAT's cluster- thresholding method, which first defines contiguous clusters using a Z statistic maximum height threshold. Then, each cluster's estimated significance level (from Gaussian random field theory) is compared with the cluster probability threshold. Only clusters that meet these two levels of threshold are considered significantly active. Group analyses, using the genotype groups defined above, were carried out using FLAME (Beckmann et al, 2003; Woolrich et al, 2004). In addition, to determine possible additive effects of the CNR1 and FAAH, we correlated number of risk alleles (ie, CNR1 G and FAAH C) with BOLD response to the marijuana vs the control cue. Only participants with available genotypes for both CNR1 and FAAH were included in these analyses (N=34). Of these, 6 had 3 risk alleles, 12 with 2 risk alleles, 13 with 1 risk allele, and 3 with no risk allele. There were no participants with four risk alleles.
Equivalence tests revealed that the CNR1 and FAAH genotype groups were significantly different on gender. Thus, all CNR1 and FAAH analyses controlled for the effects of gender. There were no other significant differences between any of the groups.
RESULTS
Comparisons of the CNR1 groups showed that those with the G allele had significantly greater neural response to marijuana cues (compared with control cues) in a cluster of activation (volume=10 719 voxels) encompassing the OFC, IFG, insula and dorsal anterior cingulate gyrus (ACG) as compared with the CNR1 A/A group (cluster-corrected p<0.05, z>1.7). This cluster did not encompass other reward areas such as the striatum or the VTA. There was no significantly greater activation in the CNR1 A/A group as compared with the G/A group.
Comparisons of the FAAH groups showed that the FAAH C/C group had greater activation in a large cluster (volume=42 161 voxels) encompassing several areas of the reward system including the OFC, IFG, ACG, striatum, and VTA during marijuana vs control cues as compared with the A/A or A/C group (cluster-corrected p<0.05, z>1.9) (see list of local maxima in Table 2). There were no significantly greater areas of activation in the FAAH A/A or A/C group as compared with the C/C group.
In these analyses, there were five participants who had both high-risk genotypes (ie, CNR1 G/A and FAAH C/C). To account for this non-independence of the FAAH and CNR1 risk groups, the previous analyses were also carried out while controlling for the other risk genotype (eg, controlling for FAAH genotype in the CNR1 analyses). These analyses found no significant difference between the groups.
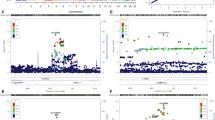
Finally, correlation analyses indicated that there was a significant positive correlation between number of risk alleles (ie, CNR1 G, FAAH C) and response to marijuana cues, such that the greater the number of risk alleles, the greater the BOLD response to marijuana cues (vs control cues) in one cluster (volume=33 842 voxels) encompassing reward areas, such as the OFC and striatum in addition to cingulate, occipital, and cerebellar regions (see Table 3, Figures 1 and 2).
Positive correlations between number of risk alleles and BOLD response in reward-associated areas. Analyses showed that the greater number of risk alleles (ie, CNR1 G, FAAH C, the greater the BOLD response to cues in several important areas related to reward processes such as the NAc, thalamus, ACG, OFC, and IFG (cluster-corrected p<0.05, z>1.9). Scatterplots illustrate the maximum z-scores for the OFC (r=0.43, p=0.01) and striatum (r=0.43, p=0.01) (y axis) against number of risk alleles (x axis).
DISCUSSION
The overarching goal of this study was to characterize the mechanisms that underlie earlier described associations between SNPs in CNR1 and FAAH and cue reactivity in heavy marijuana users (Haughey et al, 2008; Schacht et al, 2009). Our results show that rs2023239 in CNR1 and rs324420 in FAAH are associated with differential neural response to marijuana cues, such that carriers of the CNR1 G allele and FAAH C homozygotes had greater neural response in structures along the reward pathways. Further, our results show that this pattern of heightened response to cues increases as the number of risk alleles increases. These findings provide further evidence for the relevance of CNR1 and FAAH to intermediate phenotypes of CD, and suggest that a dysregulation in the reward system may mediate these genetic effects.
Although candidate genetic studies of CD to date are relatively sparse, these findings are congruent with the emergent literature (for a review, see (Agrawal and Lynskey, 2009)). Specifically, these findings support previous reports that CNR1 variants are associated with CD (Hartman et al, 2009; Agrawal et al, 2008), and that FAAH C allele carriers are more susceptible to marijuana dependence than FAAH A allele carriers (Haughey et al, 2008; Schacht et al, 2009; Tyndale et al, 2007). It is also noteworthy that the genetic effects were greater for the FAAH gene than for the CNR1 gene in terms of activation cluster size (ie, 42 161 vs 10 719 voxels) and maximum z-scores (4.3 vs 3.3). This is of particular interest given the recent reports from animal studies suggesting that FAAH blockade has complex effects that are similar to blocking CB1 receptors (Le Foll and Goldberg, 2004). The mechanism for this effect has been suggested to be not only increased anandamide levels but also elevated levels of other ethanolamides such as oleoylethanolamide (OEA) and palmitoylethanolamide (PEA) (Solinas et al, 2006; Wise et al, 2008). Emergent evidence for this effect particularly in response to nicotine has also been reported (Forget et al, 2009; Scherma et al, 2008). Unfortunately, this study does not allow for a direct investigation of the possible joint effects of cannabis and nicotine because there were only 12 smokers in our total sample.
The current findings add to this literature by elucidating the underlying mechanisms related to this increased sensitivity. The existing functional imaging study of cue reactivity among heavy marijuana users (Filbey et al, 2009), in addition to those of alcohol (Filbey et al, 2008), cocaine (Goldstein et al, 2009), and nicotine users (Franklin et al, 2007) show that the mechanism of cue reactivity is associated neural response in the pathways that underlie the anticipation of reward. These findings also add to a growing body of evidence suggesting that measures of neuronal response to cues are sensitive to genetic variation in genes such as DRD4 (with smoking cues (McClernon et al, 2007); and alcohol cues (Filbey et al, 2008), and CNR1 (Hutchison et al, 2008), although replications of these results are needed. This dysregulation of neural response in areas related to reward anticipation may have a critical role in the etiology of CD (Filbey et al, 2009), and genetic variation that influences this dysregulation may exacerbate or mitigate the development of dependence.
Our findings also suggest the presence of additive genetic effects, such that a greater number of risk alleles across CNR1 or FAAH are associated with greater response in the reward system. A similar CNR1 × FAAH interaction was also noted by Haughey et al (2008), such that individuals with the G/A–A/C genotype for CNR1–FAAH showed more severe negative affect than other genotype groups after abstinence and again after cue exposure. Additive genetic influences have been reported earlier on subjective effects of marijuana. Specifically, additive genetic influences (rather than because of a single gene with large effects or environmental influences) were found to explain the variance in subjective response to marijuana (Lyons et al, 1997). Similar findings of additive genetic effects in CD have also been reported in adolescents, which were particularly evident in late adolescence (Kendler et al, 2008).
These findings have several important clinical implications. Pharmacological agents that target CB1 receptors and/or elevate brain levels of endocannabinoids might alleviate cannabis withdrawal and dependence differentially by genotype. For instance, the FAAH inhibitor URB597, which selectively increases anandamide levels in the brain of rodents and primates, has been suggested as a possible therapeutic avenue for the treatment of cannabis withdrawal by significantly attenuating rimonabant-precipitated withdrawal signs in THC-dependent mice although genetic mediators of this effect have not yet been analyzed (Schlosburg et al, 2009; Clapper et al, 2009). In addition, the determination of the genetic factors that are associated with CD holds promise for the future development of more targeted or personalized treatment. In light of the Tyndale et al (2007) finding that FAAH is associated with risk of progression to CD, as well as our own findings that both of the SNPs analyzed here show associations with intermediate CD phenotypes among fairly young users, determination of composite genetic risk for CD might allow intervention before progression to full-blown dependence.
Some caveats must be taken into consideration in the interpretation of these findings. First, the relatively small sample size does not allow for a three-group analysis (ie, with each allele combination for each SNP) or the ability to control for the effect of one gene on the other (ie, the same participants are used in both analyses and some overlap in the high-risk and low-risk groups of each set of analyses). Regarding the latter non-independence of the CNR1 and FAAH analyses, it is possible that the effects found in each are because of inclusion of persons with the alternative high-risk genotype being examined in the other set of analyses. The correlation analysis, to some extent, addresses the non-independence of these genotypes. Replication of these findings is necessary with a larger sample size that permits a more thorough interrogation of each allele contribution. There are also methodological issues that limit the interpretation of these findings. As we described in Filbey et al (2009), only 54% of the participants reported the marijuana pipe as their primary mode of use. However, keeping the cue consistent across participants controls for the effects of subject-specific cues. Another limitation is fatigue, which is a common caveat in fMRI studies. However, we believe that any effects are minimal because the task is not cognitively demanding.
To conclude, the present findings not only add to the growing literature on the effects of biologically relevant genes on the morbidity toward CD, but also suggest that these effects may be related to hyperactivity in reward pathways. Further, this study highlights the importance of analyzing families of genes (vs single genes) and their interactions to make better predictions of vulnerability to cannabis use disorders. Future studies should determine the mechanisms of gene × gene interactions, given that there is a biologically plausible pathway for their interaction.
References
Agrawal A, Lynskey MT (2009). Candidate genes for cannabis use disorders: findings, challenges and directions. Addiction 104: 518–532.
Agrawal A, Wetherill L, Dick DM, Xuei X, Hinrichs A, Hesselbrock V et al (2008). Evidence for association between polymorphisms in the cannabinoid receptor 1 (CNR1) gene and cannabis dependence. Am J Med Genet B Neuropsychiatr Genet 150B: 736–740.
Arnone M, Maruani J, Chaperon F, Thiebot MH, Poncelet M, Soubrie P et al (1997). Selective inhibition of sucrose and ethanol intake by SR 141716, an antagonist of central cannabinoid (CB1) receptors. Psychopharmacology (Berl) 132: 104–106.
Ballon N, Leroy S, Roy C, Bourdel MC, Charles-Nicolas A, Krebs MO et al (2006). (AAT)n repeat in the cannabinoid receptor gene (CNR1): association with cocaine addiction in an African-Caribbean population. Pharmacogenomics J 6: 126–130.
Beckmann CF, Jenkinson M, Smith SM (2003). General multilevel linear modeling for group analysis in FMRI. Neuroimage 20: 1052–1063.
Berrendero F, Mendizabal V, Murtra P, Kieffer BL, Maldonado R (2003). Cannabinoid receptor and WIN 55 212-2-stimulated [35S]-GTPgammaS binding in the brain of mu-, delta- and kappa-opioid receptor knockout mice. Eur J Neurosci 18: 2197–2202.
Castane A, Valjent E, Ledent C, Parmentier M, Maldonado R, Valverde O (2002). Lack of CB1 cannabinoid receptors modifies nicotine behavioural responses, but not nicotine abstinence. Neuropharmacology 43: 857–867.
Cheer JF, Marsden CA, Kendall DA, Mason R (2000). Lack of response suppression follows repeated ventral tegmental cannabinoid administration: an in vitro electrophysiological study. Neuroscience 99: 661–667.
Chiang KP, Gerber AL, Sipe JC, Cravatt BF (2004). Reduced cellular expression and activity of the P129T mutant of human fatty acid amide hydrolase: evidence for a link between defects in the endocannabinoid system and problem drug use. Hum Mol Genet 13: 2113–2119.
Clapper JR, Mangieri RA, Piomelli D (2009). The endocannabinoid system as a target for the treatment of cannabis dependence. Neuropharmacology 56 (Suppl 1): 235–243.
Comings DE, Muhleman D, Gade R, Johnson P, Verde R, Saucier G et al (1997). Cannabinoid receptor gene (CNR1): association with i.v. drug use. Mol Psychiatry 2: 161–168.
Covault J, Gelernter J, Kranzler H (2001). Association study of cannabinoid receptor gene (CNR1) alleles and drug dependence. Mol Psychiatry 6: 501–502.
Cravatt BF, Demarest K, Patricelli MP, Bracey MH, Giang DK, Martin BR et al (2001). Supersensitivity to anandamide and enhanced endogenous cannabinoid signaling in mice lacking fatty acid amide hydrolase. Proc Natl Acad Sci USA 98: 9371–9376.
Cravatt BF, Lichtman AH (2003). Fatty acid amide hydrolase: an emerging therapeutic target in the endocannabinoid system. Curr Opin Chem Biol 7: 469–475.
Deichmann R, Gottfried JA, Hutton C, Turner R (2003). Optimized EPI for fMRI studies of the orbitofrontal cortex. Neuroimage 19 (2 Pt 1): 430–441.
Diana M, Melis M, Muntoni AL, Gessa GL (1998). Mesolimbic dopaminergic decline after cannabinoid withdrawal. Proc Natl Acad Sci USA 95: 10269–10273.
Filbey FM, Ray L, Smolen A, Claus ED, Audette A, Hutchison KE (2008). Differential neural response to alcohol priming and alcohol taste cues is associated with DRD4 VNTR and OPRM1 genotypes. Alcohol Clin Exp Res 32: 1113–1123.
Filbey FM, Schacht JP, Myers US, Chavez RS, Hutchison KE (2009). Marijuana craving in the brain. Proc Natl Acad Sci USA 106: 13016–13021.
Forget B, Coen KM, Le Foll B (2009). Inhibition of fatty acid amide hydrolase reduces reinstatement of nicotine seeking but not break point for nicotine self-administration--comparison with CB(1) receptor blockade. Psychopharmacology (Berl) 205: 613–624.
Franklin TR, Wang Z, Wang J, Sciortino N, Harper D, Li Y et al (2007). Limbic activation to cigarette smoking cues independent of nicotine withdrawal: a perfusion fMRI study. Neuropsychopharmacology 32: 2301–2309.
Freire L, Mangin JF (2001). Motion correction algorithms may create spurious brain activations in the absence of subject motion. Neuroimage 14: 709–722.
Freire L, Roche A, Mangin JF (2002). What is the best similarity measure for motion correction in fMRI time series? IEEE Trans Med Imaging 21: 470–484.
French ED, Dillon K, Wu X (1997). Cannabinoids excite dopamine neurons in the ventral tegmentum and substantia nigra. Neuroreport 8: 649–652.
Gardner B, Zu LX, Sharma S, Liu Q, Makriyannis A, Tashkin DP et al (2002). Autocrine and paracrine regulation of lymphocyte CB2 receptor expression by TGF-beta. Biochem Biophys Res Commun 290: 91–96.
Goldstein RZ, Tomasi D, Alia-Klein N, Honorio Carrillo J, Maloney T, Woicik PA et al (2009). Dopaminergic response to drug words in cocaine addiction. J Neurosci 29: 6001–6006.
Hariri AR, Weinberger DR (2003). Imaging genomics. Br Med Bull 65: 259–270.
Hartman CA, Hopfer CJ, Haberstick B, Rhee SH, Crowley TJ, Corley RP et al (2009). The association between cannabinoid receptor 1 gene (CNR1) and cannabis dependence symptoms in adolescents and young adults. Drug Alcohol Depend 104: 11–16.
Haughey HM, Marshall E, Schacht JP, Louis A, Hutchison KE (2008). Marijuana withdrawal and craving: influence of the cannabinoid receptor 1 (CNR1) and fatty acid amide hydrolase (FAAH) genes. Addiction 103: 1678–1686.
Herman AI, Kranzler HR, Cubells JF, Gelernter J, Covault J (2006). Association study of the CNR1 gene exon 3 alternative promoter region polymorphisms and substance dependence. Am J Med Genet B Neuropsychiatr Genet 141B: 499–503.
Hopfer CJ, Young SE, Purcell S, Crowley TJ, Stallings MC, Corley RP et al (2006). Cannabis receptor haplotype associated with fewer cannabis dependence symptoms in adolescents. Am J Med Genet B Neuropsychiatr Genet 141B: 895–901.
Hutchison KE, Haughey H, Niculescu M, Schacht J, Kaiser A, Stitzel J et al (2008). The incentive salience of alcohol: translating the effects of genetic variant in CNR1. Arch Gen Psychiatry 65: 841–850.
Jenkinson M, Bannister P, Brady M, Smith S (2002). Improved optimization for the robust and accurate linear registration and motion correction of brain images. Neuroimage 17: 825–841.
Jenkinson M, Smith S (2001). A global optimisation method for robust affine registration of brain images. Med Image Anal 5: 143–156.
Justinova Z, Munzar P, Panlilio LV, Yasar S, Redhi GH, Tanda G et al (2008). Blockade of THC-seeking behavior and relapse in monkeys by the cannabinoid CB(1)-receptor antagonist rimonabant. Neuropsychopharmacology 33: 2870–2877.
Justinova Z, Tanda G, Redhi GH, Goldberg SR (2003). Self-administration of delta9-tetrahydrocannabinol (THC) by drug naive squirrel monkeys. Psychopharmacology (Berl) 169: 135–140.
Kendler KS, Schmitt E, Aggen SH, Prescott CA (2008). Genetic and environmental influences on alcohol, caffeine, cannabis, and nicotine use from early adolescence to middle adulthood. Arch Gen Psychiatry 65: 674–682.
Le Foll B, Goldberg SR (2004). Rimonabant, a CB1 antagonist, blocks nicotine-conditioned place preferences. Neuroreport 15: 2139–2143.
Ledent C, Valverde O, Cossu G, Petitet F, Aubert JF, Beslot F et al (1999). Unresponsiveness to cannabinoids and reduced addictive effects of opiates in CB1 receptor knockout mice. Science 283: 401–404.
Li T, Liu X, Zhu ZH, Zhao J, Hu X, Ball DM et al (2000). No association between (AAT)n repeats in the cannabinoid receptor gene (CNR1) and heroin abuse in a Chinese population. Mol Psychiatry 5: 128–130.
Lowe JB, Windsor RA, Adams B, Morris J, Reese Y (1986). Use of a bogus pipeline method to increase accuracy of self-reported alcohol consumption among pregnant women. J Stud Alcohol 47: 173–175.
Lyons MJ, Toomey R, Meyer JM, Green AI, Eisen SA, Goldberg J et al (1997). How do genes influence marijuana use? The role of subjective effects. Addiction 92: 409–417.
McClernon FJ, Hutchison KE, Rose JE, Kozink RV (2007). DRD4 VNTR polymorphism is associated with transient fMRI-BOLD responses to smoking cues. Psychopharmacology (Berl) 194: 433–441.
Racz I, Bilkei-Gorzo A, Toth ZE, Michel K, Palkovits M, Zimmer A (2003). A critical role for the cannabinoid CB1 receptors in alcohol dependence and stress-stimulated ethanol drinking. J Neurosci 23: 2453–2458.
Schacht JP, Selling RE, Hutchison KE (2009). Intermediate cannabis dependence phenotypes and the FAAH C385A variant: an exploratory analysis. Psychopharmacology (Berl) 203: 511–517.
Scherma M, Panlilio LV, Fadda P, Fattore L, Gamaleddin I, Le Foll B et al (2008). Inhibition of anandamide hydrolysis by cyclohexyl carbamic acid 3′-carbamoyl-3-yl ester (URB597) reverses abuse-related behavioral and neurochemical effects of nicotine in rats. J Pharmacol Exp Ther 327: 482–490.
Schlosburg JE, Carlson BL, Ramesh D, Abdullah RA, Long JZ, Cravatt BF et al (2009). Inhibitors of endocannabinoid-metabolizing enzymes reduce precipitated withdrawal responses in THC-dependent mice. AAPS J 11: 342–352.
Schmidt LG, Samochowiec J, Finckh U, Fiszer-Piosik E, Horodnicki J, Wendel B et al (2002). Association of a CB1 cannabinoid receptor gene (CNR1) polymorphism with severe alcohol dependence. Drug Alcohol Depend 65: 221–224.
Solinas M, Justinova Z, Goldberg SR, Tanda G (2006). Anandamide administration alone and after inhibition of fatty acid amide hydrolase (FAAH) increases dopamine levels in the nucleus accumbens shell in rats. J Neurochem 98: 408–419.
Tyndale RF, Payne JI, Gerber AL, Sipe JC (2007). The fatty acid amide hydrolase C385A (P129T) missense variant in cannabis users: studies of drug use and dependence in Caucasians. Am J Med Genet B Neuropsychiatr Genet 144B: 660–666.
Walker AH, Najarian D, White DL, Jaffe JF, Kanetsky PA, Rebbeck TR (1999). Collection of genomic DNA by buccal swabs for polymerase chain reaction-based biomarker assays. Environ Health Perspect 107: 517–520.
Weiskopf N, Hutton C, Josephs O, Turner R, Deichmann R (2007). Optimized EPI for fMRI studies of the orbitofrontal cortex: compensation of susceptibility-induced gradients in the readout direction. Magma 20: 39–49.
Wise LE, Cannavacciulo R, Cravatt BF, Martin BF, Lichtman AH (2008). Evaluation of fatty acid amides in the carrageenan-induced paw edema model. Neuropharmacology 54: 181–188.
Wiskerke J, Pattij T, Schoffelmeer AN, De Vries TJ (2008). The role of CB1 receptors in psychostimulant addiction. Addict Biol 13: 225–238.
Woolrich MW, Behrens TE, Beckmann CF, Jenkinson M, Smith SM (2004). Multilevel linear modelling for FMRI group analysis using Bayesian inference. Neuroimage 21: 1732–1747.
Zhang PW, Ishiguro H, Ohtsuki T, Hess J, Carillo F, Walther D et al (2004). Human cannabinoid receptor 1: 5′ exons, candidate regulatory regions, polymorphisms, haplotypes and association with polysubstance abuse. Mol Psychiatry 9: 916–931.
Zuo L, Kranzler HR, Luo X, Covault J, Gelernter J (2007). CNR1 variation modulates risk for drug and alcohol dependence. Biol Psychiatry 62: 616–626.
Zuo L, Kranzler HR, Luo X, Yang BZ, Weiss R, Brady K et al (2009). Interaction between two independent CNR1 variants increases risk for cocaine dependence in European Americans: a replication study in family-based sample and population-based sample. Neuropsychopharmacology 34: 1504–1513.
Acknowledgements
This research was funded by grant KO1 5 KO1 DA021632-02 to Dr Filbey from the National Institute on Drug Abuse/National Institutes of Health. Dr Schacht was supported by grant F31 DA021496 from the National Institute on Drug Abuse/National Institutes of Health. We would like to thank Marilee Morgan and Melynda Byers for the genetic analyses of the participants.
Author information
Authors and Affiliations
Corresponding author
Additional information
DISCLOSURE
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Filbey, F., Schacht, J., Myers, U. et al. Individual and Additive Effects of the CNR1 and FAAH Genes on Brain Response to Marijuana Cues. Neuropsychopharmacol 35, 967–975 (2010). https://doi.org/10.1038/npp.2009.200
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2009.200
Keywords
This article is cited by
-
The cannabinoid receptor-1 gene interacts with stressful life events to increase the risk for problematic alcohol use
Scientific Reports (2022)
-
Systematic review of structural and functional neuroimaging studies of cannabis use in adolescence and emerging adulthood: evidence from 90 studies and 9441 participants
Neuropsychopharmacology (2022)
-
The endocannabinoid system in humans: significant associations between anandamide, brain function during reward feedback and a personality measure of reward dependence
Neuropsychopharmacology (2021)
-
Deconstructing the neurobiology of cannabis use disorder
Nature Neuroscience (2020)
-
Effects of the cannabinoid receptor agonist CP-55,940 on incentive salience attribution
Psychopharmacology (2020)