Abstract
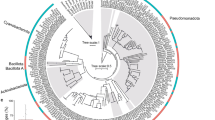
Light-harvesting complex (LHC) proteins in chloroplast thylakoid membranes not only transfer absorbed light energy to the two photosystems but also regulate the rate of energy transfer to avoid photodamage. Here we demonstrate that Lhcb9, a recently discovered LHC protein in the moss Physcomitrella patens, functions to connect LHC proteins with photosystem I (PSI), resulting in the formation of two different types of PSI supercomplexes in thylakoid membranes. We observed that the Lhcb9-containing PSI supercomplex is disassembled in response to excess light conditions. On the basis of our phylogenetic analysis, it appears that P. patens acquired Lhcb9 by horizontal gene transfer from the earlier green algal lineage, leading to the presence of both green alga-type and vascular plant-type PSI supercomplexes, which would have been crucial for conquering the dynamic environmental interface between aquatic and terrestrial conditions it faced during evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Nelson, N. & Yocum, C. F. Structure and function of photosystems I and II. Annu. Rev. Plant Biol. 57, 521–565 (2006).
Allen, J. F., Bennet, J., Steinback, K. E. & Arntzen, C. J. Chloroplast protein phosphorylation couples plastoquinone redox state to distribution of excitation energy between photosystems. Nature 291, 21–25 (1981).
Bellafiore, S., Barneche, F., Peltier, G. & Rochaix, J. D. State transitions and light adaptation require chloroplast thylakoid protein kinase STN7. Nature 433, 892–895 (2005).
Wollman, F. A. State transitions reveal the dynamics and flexibility of the photosynthetic apparatus. EMBO J. 20, 3623–3630 (2001).
Allen, J. F. & Forsberg, J. Molecular recognition in thylakoid structure and function. Trends Plant Sci. 6, 317–326 (2001).
Minagawa, J. State transitions—the molecular remodeling of photosynthetic supercomplexes that controls energy flow in the chloroplast. Biochim. Biophys. Acta 1807, 897–905 (2011).
Horton, P., Ruban, A. V. & Walters, R. G. Regulation of light harvesting in green plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 47, 655–684 (1996).
Niyogi, K. K. Photoprotection revisited: genetic and molecular approaches. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 333–359 (1999).
Peers, G. et al. An ancient light-harvesting protein is critical for the regulation of algal photosynthesis. Nature 462, 518–521 (2009).
Li, X. P. et al. A pigment-binding protein essential for regulation of photosynthetic light harvesting. Nature 403, 391–395 (2000).
Bonente, G. et al. Analysis of LhcSR3, a protein essential for feedback de-excitation in the green alga Chlamydomonas reinhardtii. PLoS Biol. 9, e1000577 (2011).
Tokutsu, R. & Minagawa, J. Energy-dissipative supercomplex of photosystem II associated with LHCSR3 in Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 110, 10016–10021 (2013).
Betterle, N. et al. Light-induced dissociation of an antenna hetero-oligomer is needed for non-photochemical quenching induction. J. Biol. Chem. 284, 15255–15266 (2009).
Goral, T. K. et al. Light-harvesting antenna composition controls the macrostructure and dynamics of thylakoid membranes in Arabidopsis. Plant J. 69, 289–301 (2012).
Niyogi, K. K. & Truong, T. B. Evolution of flexible non-photochemical quenching mechanisms that regulate light harvesting in oxygenic photosynthesis. Curr. Opin. Plant Biol. 16, 307–314 (2013).
Rensing, S. A. et al. The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science 319, 64–69 (2008).
Alboresi, A., Gerotto, C., Cazzaniga, S., Bassi, R. & Morosinotto, T. A red-shifted antenna protein associated with photosystem II in Physcomitrella patens. J. Biol. Chem. 286, 28978–28987 (2011).
Engelmann, E. et al. Influence of the photosystem I-light harvesting complex I antenna domains on fluorescence decay. Biochemistry 45, 6947–6955 (2006).
Wientjes, E., van Stokkum, I. H., van Amerongen, H. & Croce, R. The role of the individual Lhcas in photosystem I excitation energy trapping. Biophys. J. 101, 745–754 (2011).
Mimuro, M., Yokono, M. & Akimoto, S. Variations in photosystem I properties in the primordial cyanobacterium Gloeobacter violaceus PCC 7421. Photochem. Photobiol. 86, 62–69 (2010).
Galka, P. et al. Functional analyses of the plant photosystem I-light-harvesting complex II supercomplex reveal that light-harvesting complex II loosely bound to photosystem II is a very efficient antenna for photosystem I in state II. Plant Cell 24, 2963–2978 (2012).
Stauber, E. J. et al. Proteomics of Chlamydomonas reinhardtii light-harvesting proteins. Eukaryot. Cell 2, 978–994 (2003).
Takahashi, H., Iwai, M., Takahashi, Y. & Minagawa, J. Identification of the mobile light-harvesting complex II polypeptides for state transitions in Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 103, 477–482 (2006).
Rochaix, J. D. Role of thylakoid protein kinases in photosynthetic acclimation. FEBS Lett. 581, 2768–2775 (2007).
Sonoike, K. Photoinhibition of photosystem I. Physiol. Plant. 142, 56–64 (2011).
Joliot, P. & Johnson, G. N. Regulation of cyclic and linear electron flow in higher plants. Proc. Natl Acad. Sci. USA 108, 13317–13322 (2011).
Busch, A. et al. Composition and structure of photosystem I in the moss Physcomitrella patens. J. Exp. Bot. 64, 2689–2699 (2013).
Wientjes, E., van Amerongen, H. & Croce, R. LHCII is an antenna of both photosystems after long-term acclimation. Biochim. Biophys. Acta 1827, 420–426 (2013).
Amunts, A., Drory, O. & Nelson, N. The structure of a plant photosystem I supercomplex at 3.4Å resolution. Nature 447, 58–63 (2007).
Stauber, E. J., Busch, A., Naumann, B., Svatos, A. & Hippler, M. Proteotypic profiling of LHCI from Chlamydomonas reinhardtii provides new insights into structure and function of the complex. Proteomics 9, 398–408 (2009).
Drop, B. et al. Photosystem I of Chlamydomonas reinhardtii contains nine light-harvesting complexes (Lhca) located on one side of the core. J. Biol. Chem. 286, 44878–44887 (2011).
Swingley, W. D. et al. Characterization of photosystem I antenna proteins in the prasinophyte Ostreococcus tauri. Biochim. Biophys. Acta 1797, 1458–1464 (2010).
Yue, J., Hu, X., Sun, H., Yang, Y. & Huang, J. Widespread impact of horizontal gene transfer on plant colonization of land. Nature Commun. 3, 1152 (2012).
Nishiyama, T., Hiwatashi, Y., Sakakibara, I., Kato, M. & Hasebe, M. Tagged mutagenesis and gene-trap in the moss, Physcomitrella patens by shuttle mutagenesis. DNA Res. 7, 9–17 (2000).
Berthold, D. A., Babcock, G. T. & Yocum, C. F. A highly resolved, oxygen-evolving photosystem-II preparation from spinach thylakoid membranes—EP and electron-transport properties. FEBS Lett. 134, 231–234 (1981).
Iwai, M., Takahashi, Y. & Minagawa, J. Molecular remodeling of photosystem II during state transitions in Chlamydomonas reinhardtii. Plant Cell 20, 2177–2189 (2008).
Anderson, J. M. & Melis, A. Localization of different photosystems in separate regions of chloroplast membranes. Proc. Natl Acad. Sci. USA 80, 745–749 (1983).
Ghirardi, M. L. & Melis, A. Photosystem electron-transport capacity and light-harvesting antenna size in maize chloroplasts. Plant Physiol. 74, 993–998 (1984).
Asada, K., Heber, U. & Schreiber, U. Pool size of electrons that can be donated to P700+, as determined in intact leaves—donation to P700+ from stromal components via the intersystem chain. Plant Cell Physiol. 33, 927–932 (1992).
Haldrup, A., Jensen, P. E., Lunde, C. & Scheller, H. V. Balance of power: a view of the mechanism of photosynthetic state transitions. Trends Plant Sci. 6, 301–305 (2001).
Yokono, M., Akimoto, S., Koyama, K., Tsuchiya, T. & Mimuro, M. Energy transfer processes in Gloeobacter violaceus PCC 7421 that possesses phycobilisomes with a unique morphology. Biochim. Biophys. Acta 1777, 55–65 (2008).
Mimuro, M. et al. Delayed fluorescence observed in the nanosecond time region at 77 K originates directly from the photosystem II reaction center. Biochim. Biophys. Acta 1767, 327–334 (2007).
Yokono, M. et al. Alterations in photosynthetic pigments and amino acid composition of D1 protein change energy distribution in photosystem II. Biochim. Biophys. Acta 1817, 754–759 (2012).
Hippler, M., Klein, J., Fink, A., Allinger, T. & Hoerth, P. Towards functional proteomics of membrane protein complexes: analysis of thylakoid membranes from Chlamydomonas reinhardtii. Plant J. 28, 595–606 (2001).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Ito, H., Yokono, M., Tanaka, R. & Tanaka, A. Identification of a novel vinyl reductase gene essential for the biosynthesis of monovinyl chlorophyll in Synechocystis sp. PCC6803. J. Biol. Chem. 283, 9002–9011 (2008).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Felsenstein, J. PHYLIP (Phylogeny Inference Package) v.3.69. (Dept. Genome Sci., Univ. Washington, Seattle, 2005).
Acknowledgements
We are grateful to Dr Ichiro Terashima for fruitful discussion of the manuscript; Drs Mitsuyasu Hasebe and Yuji Hiwatashi for providing the P. patens WT strain and extensive technical advice on using the moss; Dr Hiroshi Abe for technical advice regarding cloning and gene construction; the Support Unit for Bio-Material Analysis, RIKEN BSI Research Resources Center, with special thanks to Kaori Otsuki, Masaya Usui and Aya Abe for MS; and Kaoru Kotoshiba and Hiroe Watanabe for technical support. This work was supported by JST PRESTO, JSPS KAKENHI Grant Number 23687008, and grants from RIKEN Center for Advanced Photonics, Extreme Photonics Research Project.
Author information
Authors and Affiliations
Contributions
M.I. designed the study, constructed the transgenic plants, performed the genetic, biochemical and PAM experiments, analysed the data and wrote the paper; M.Y. and S.A. performed the time-resolved fluorescence spectroscopy experiments and analysed the data; M.Y. performed the phylogenetic analysis; M.K. and K.N. provided essential assistance for PAM experiments; and A.N. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Iwai, M., Yokono, M., Kono, M. et al. Light-harvesting complex Lhcb9 confers a green alga-type photosystem I supercomplex to the moss Physcomitrella patens. Nature Plants 1, 14008 (2015). https://doi.org/10.1038/nplants.2014.8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2014.8
This article is cited by
-
Revealing the significance of chlorophyll b in the moss Physcomitrium patens by knocking out two functional chlorophyllide a oxygenase
Photosynthesis Research (2023)
-
The role of mixed vibronic Qy-Qx states in green light absorption of light-harvesting complex II
Nature Communications (2020)
-
An atypical short-chain dehydrogenase–reductase functions in the relaxation of photoprotective qH in Arabidopsis
Nature Plants (2020)
-
Vibronic mixing enables ultrafast energy flow in light-harvesting complex II
Nature Communications (2020)
-
Formation of a PSI–PSII megacomplex containing LHCSR and PsbS in the moss Physcomitrella patens
Journal of Plant Research (2019)