Abstract
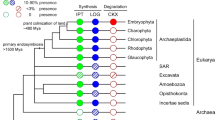
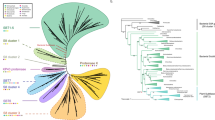
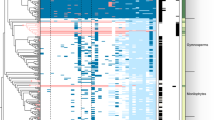
Land plants evolved more than 450 million years ago from a lineage of freshwater charophyte green algae1. The extent to which plant signalling systems existed before the evolutionary transition to land is unknown. Although charophytes occupy a key phylogenetic position for elucidating the origins of such signalling systems2–4, there is a paucity of sequence data for these organisms5,6. Here we carry out de novo transcriptomics of five representative charophyte species, and find putative homologues for the biosynthesis, transport, perception and signalling of major plant hormones. Focusing on the plant hormone ethylene, we provide evidence that the filamentous charophyte Spirogyra pratensis possesses an ethylene hormone system homologous to that in plants. Spirogyra produces ethylene and exhibits a cell elongation response to ethylene. Spirogyra ethylene-signalling homologues partially rescue mutants of the angiosperm Arabidopsis thaliana and respond post-translationally to ethylene when expressed in plant cells, indicative of unambiguously homologous ethylene-signalling pathways in Spirogyra and Arabidopsis. These findings imply that the common aquatic ancestor possessed this pathway prior to the colonization of land and that cell elongation was possibly an ancestral ethylene response. This highlights the importance of charophytes for investigating the origins of fundamental plant processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sanderson, M. J., Thorne, J. L., Wikström, N. & Bremer, K. Molecular evidence on plant divergence times. Am. J. Bot. 91, 1656–1665 (2004).
Karol, K. G., McCourt, R. M., Cimino, M. T. & Delwiche, C. F. The closest living relatives of land plants. Science 294, 2351–2353 (2001).
Turmel, M., Ehara, M., Otis, C. & Lemieux, C. Phylogenetic relationships among streptophytes as inferred from chloroplast small and large subunit rRNA gene sequences. J. Phycol. 38, 364–375 (2002).
Leliaert, F. et al. Phylogeny and molecular evolution of the green algae. Crit. Rev. Plant Sci. 31, 1–46 (2012).
Timme, R. E. & Delwiche, C. F. Uncovering the evolutionary origin of plant molecular processes: comparison of Coleochaete (Coleochaetales) and Spirogyra (Zygnematales) transcriptomes. BMC Plant Biol. 10, 96 (2010).
Wodniok, S. et al. Origin of land plants: Do conjugating green algae hold the key? BMC Evol. Biol. 11, 104 (2011).
McManus, M. T. Annual Plant Reviews, The Plant Hormone Ethylene (Wiley-Blackwell, 2012).
Jackson, M. B. Ethylene-promoted elongation: an adaptation to submergence stress. Ann. Bot. 101, 229–248 (2008).
Merchante, C., Alonso, J. M. & Stepanova, A. N. Ethylene signaling: simple ligand, complex regulation. Curr. Opin. Plant Biol. 16, 554–560 (2013).
Mount, S. M. & Chang, C. Evidence for a plastid origin of plant ethylene receptor genes. Plant Physiol. 130, 10–14 (2002).
Wang, W. et al. Identification of important regions for ethylene binding and signaling in the transmembrane domain of the ETR1 ethylene receptor of Arabidopsis. Plant Cell 18, 3429–3442 (2006).
Timme, R. E., Bachvaroff, T. R. & Delwiche, C. F. Broad phylogenomic sampling and the sister lineage of land plants. PLoS ONE 7, e29696. (2012).
Hori, K. et al. Klebsormidium flaccidum genome reveals primary factors for plant terrestrial adaptation. Nature Commun. 5, 3978. (2014).
Yang, S. F. & Hoffman, N. E. Ethylene biosynthesis and its regulation in higher plants. Ann. Rev. Plant Physiol. 35, 155–189 (1984).
Garcia-Jimenez, P. & Robaina, R. R. Effects of ethylene on tetrasporogenesis in Pterocladiella capillacea (Rhodophyta). J. Phycol. 48, 710–715 (2012).
Maillard, P., Thepenier, C. & Gudin, C. Determination of an ethylene biosynthesis pathway in the unicellular green alga, Haematococcus pluvialis. Relationship between growth and ethylene production. J. Appl. Phycol. 5, 93–98 (1993).
Huang, T.-C. & Chow, T.-J. Ethylene production by blue-green algae. Bot. Bull. Academia Sinica 25, 81–86 (1984).
Bleecker, A. B., Estelle, M. A., Somerville, C. & Kende, H. Insensitivity to ethylene conferred by a dominant mutation in Arabidopsis thaliana. Science 241, 1086–1089 (1988).
Blankenship, S. M. & Dole, J. M. 1-methylcyclopropene: a review. Postharv. Biol. Technol. 28, 1–25 (2003).
Yasumura, Y. et al. Studies of Physcomitrella patens reveal that ethylene-mediated submergence responses arose relatively early in land-plant evolution. Plant J. 72, 947–959 (2012).
Smalle, J., Haegman, M., Kurepa, J., Van Montagu, M. & Van Der Straeten, D. Ethylene can stimulate Arabidopsis hypocotyl elongation in the light. Proc. Natl Acad. Sci. USA 94, 2756–2761 (1997).
Chen, Y. F., Randlett, M. D., Findell, J. L. & Schaller, G. E. Localization of the ethylene receptor ETR1 to the endoplasmic reticulum of Arabidopsis. J. Biol. Chem. 277, 19861–19866 (2002).
Grefen, C. et al. Subcellular localization and in vivo interaction of the Arabidopsis thaliana ethylene receptor family members. Molecular Plant 1, 308–320 (2008).
Clark, K. L., Larsen, P. B., Wang, X. & Chang, C. Association of the Arabidopsis CTR1 Raf-like kinase with the ETR1 and ERS ethylene receptors. Proc. Natl Acad. Sci. USA 95, 5401–5406 (1998).
Gao, Z et al. Localization of the Raf-like kinase CTR1 to the endoplasmic reticulum of Arabidopsis through participation in ethylene receptor signaling complexes. J. Biol. Chem. 278, 34725–34732 (2003).
Ju, C. et al. CTR1 phosphorylates the central regulator EIN2 to control ethylene hormone signaling from the ER membrane to the nucleus in Arabidopsis. Proc. Natl Acad. Sci. USA 109, 19486–19491 (2012).
Alonso, J. M. et al. EIN2, a bifunctional transducer of ethylene and stress responses in Arabidopsis. Science 284, 2148–2152 (1999).
Qiao, H. et al. Processing and subcellular trafficking of ER-tethered EIN2 control response to ethylene gas. Science 338, 390–393 (2012).
Guo, H. & Ecker, J. R. Plant responses to ethylene gas are mediated by SCFEBF1/EBF2-dependent proteolysis of EIN3 transcription factor. Cell 115, 667–677 (2003).
Chao, Q. et al. Activation of the ethylene gas response pathway in Arabidopsis by the nuclear protein ETHYLENE-INSENSITIVE3 and related proteins. Cell 89, 1133–1144 (1997).
Solano, R., Stepanova, A., Chao, Q. & Ecker, J. R. Nuclear events in ethylene signaling: a transcriptional cascade mediated by ETHYLENE-INSENSITIVE3 and ETHYLENE-RESPONSE-FACTOR1. Genes Dev. 12, 3703–3714 (1998).
Hua, J. & Meyerowitz, E. M. Ethylene responses are negatively regulated by a receptor gene family in Arabidopsis thaliana. Cell 94, 261–271 (1998).
Nakano, T., Suzuki, K., Fujimura, T. & Shinshi, H. Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol. 140, 411–432 (2006).
Sakayama, H., Hara,Y. & Nozaki, H. Taxonomic re-examination of six species of Nitella (Charales, Charophyceae) from Asia, and phylogenetic relationships within the genus based on rbcL and atpB gene sequences. Phycologia 43, 91–104 (2004).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nature Biotechnol. 29, 644–652 (2011).
Voinnet, O., Rivas, S., Mestre, P. & Baulcombe, D. An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J. 33, 949–956 (2003).
Acknowledgements
We thank Hidetoshi Sakayama (Kobe University) for help with the Nitella transcriptome, Mark Tucker (USDA-ARS, Beltsville) for 1-MCP, Brad Binder (University of Tennessee, Knoxville) for preliminary ethylene measurements, Jocelyn Rose (Cornell University), and B. Binder and Chang lab members for comments on the manuscript. We thank the Imaging Core Facility, as well as the Institute for Bioscience and Biotechnology Research at University of Maryland. This work was supported in part by NSF grants EF0523719 (Microbial Genome Sequencing) and DEB-1036506 (Assembling the Tree of Life) to C.F.D., NSF grant MCB-0923796 to C.C., a Belgian American Educational Foundation Fellowship to B.V.d.P. and the HHMI Undergraduate Research Fellowship (from UMD) and ASPB Summer Undergraduate Fellowship to J.H.T. C.C. and C.F.D. are supported in part by the Maryland Agricultural Experiment Station.
Author information
Authors and Affiliations
Contributions
C.J. designed and carried out experiments in Arabidopsis, tobacco and yeast. B.V.d.P. designed and carried out experiments in Spirogyra and onion cells. E.D.C. designed and carried out bioinformatic analyses and transcriptome assembly. J.H.T. initiated the project and designed some of the experiments. T.R.G. performed initial transcriptome assembly and assisted with bioinformatic analyses. C.F.D. conceived and co-directed the project. C.C. co-directed the project and wrote the manuscript with assistance from the co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Ju, C., Van de Poel, B., Cooper, E. et al. Conservation of ethylene as a plant hormone over 450 million years of evolution. Nature Plants 1, 14004 (2015). https://doi.org/10.1038/nplants.2014.4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2014.4
This article is cited by
-
Cross-stress gene expression atlas of Marchantia polymorpha reveals the hierarchy and regulatory principles of abiotic stress responses
Nature Communications (2023)
-
The origin, evolution and functional divergence of HOOKLESS1 in plants
Communications Biology (2023)
-
The Destruction of Endogenous ABA Balance Caused by Fruit-specific SlNCED1-RNAi Affects Tomato Fruit Set
Journal of Plant Growth Regulation (2023)
-
Characterization of the transcriptional responses of Armillaria gallica 012m to GA3
Archives of Microbiology (2023)
-
The origin of a land flora
Nature Plants (2022)