Abstract
The condensation of stiff, highly charged DNA molecules into compact structures by condensing agents ranging from multivalent ions1 to small cationic proteins2,3 is of major biological and therapeutic importance4,5, yet the underlying microscopic mechanism remains poorly understood1,6,7,8,9. It has been proposed7,10 that DNA condensation is a purely electrostatic phenomenon driven by the existence of a strongly correlated liquid (SCL) of counterions at the DNA surface. The same theoretical argument predicts that multivalent counterions overcompensate the DNA charge when present at high concentration11, in turn destabilizing the condensates12. Here, we demonstrate the occurrence of DNA charge inversion by multivalent ions through measurements of the electrophoretic mobility of condensed DNA. By observing the multivalent-ion-induced condensation of a single DNA molecule using magnetic tweezers, we further show that charge inversion influences condensation by modulating the barrier for condensate nucleation in a manner consistent with the SCL mechanism.
Similar content being viewed by others
Main
The role played by spatial correlations of screening ions in biological systems remains poorly understood. Definitive experimental evidence is particularly difficult to obtain because of the short length scales involved. The strongly correlated liquid (SCL) mechanism predicts that charge inversion necessarily accompanies and influences counter-ion-induced like-charge attraction12, thus providing a unique opportunity to test this mechanism. We concentrate on DNA because of its high charge density, the level of experimental control that it provides and the direct relevance of condensation to genome packaging.
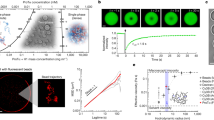
To verify the existence of DNA charge inversion, we measured the electrophoretic mobility, μ, of DNA condensates in solution using dynamic light scattering (DLS). The mobility, μ, reflects the bare charge of DNA plus that of counterions at its surface, and its sign is expected to reverse on charge inversion13,14,15. In DLS, the phase of laser light scattered from the condensates is monitored over time; condensates drifting at constant velocity in an electric field yield a phase that evolves linearly in time at a rate proportional to their mobility. Figure 1a shows the measured phase for concentrations c= 0.1 and 3 mM of the quadrivalent cation spermine ([C10N4H30]4+). The two signals have opposite slopes, indicating a sign reversal of μ (negative for c=0.1 mM and positive for c=3 mM). This is to our knowledge the first experimental report of DNA charge inversion induced solely by simple multivalent ions.
a, DLS measurements showing the applied electric field, E, and the measured phase as a function of time in solutions containing 5 ng μl−1 8 kbp DNA, 1 mM TRIS and 0.1 mM (red line) or 3 mM (blue line) spermine. The polarity of the applied voltage was periodically reversed to eliminate electro-osmotic flows. The opposite slopes for these two curves indicate that the electrophoretic mobility has reversed sign at the higher spermine concentration. b, Electrophoretic mobility, μ, of condensed DNA as a function of the spermine concentration in a buffer containing 1 mM TRIS (red circles), 10 mM TRIS (black squares) and 10 mM TRIS+50 mM KCl (blue triangles). Each data point is the average of three consecutive measurements with the corresponding standard deviation as the error.
Figure 1b shows the measured mobility of condensed DNA as a function of the concentration of spermine and buffer conditions. In 1 mM TRIS buffer, the mobility is positive for spermine concentrations greater than the charge-inversion concentration c0=0.5 mM. Increasing the TRIS buffer concentration to 10 mM hinders charge inversion, causing c0 to increase to 1 mM. Further adding 50 mM monovalent KCl salt causes charge inversion to disappear entirely for c≤3 mM, reminiscent of the disappearance of charge inversion at glass surfaces with increasing monovalent salt16. This is probably why DNA charge inversion by multivalent ions has not been reported earlier as most DNA studies are carried out at physiological (high) salt concentrations. Spermine concentrations above 3 mM were not accessible in our experiment owing to high electrochemical currents.
To ascertain whether this observed charge inversion inhibits DNA condensation12 we used magnetic tweezers, which allow monitoring the extension z of a single DNA molecule in time while applying a tunable pulling force, F (ref. 17). We previously showed18 using this technique that condensation causes a rapid step-like decrease in the DNA extension when F decreases below the condensation force, Fc (Fig. 2a), and that this first-order-like process is initiated by the spontaneous nucleation of a loop in the DNA (as shown in Fig. 2b). Here, we exploit this knowledge to elucidate the effect of the electrostatic environment and of charge inversion on DNA condensation.
a, Measurement of DNA extension z while gradually lowering the applied force, F, on an 8 kbp nicked DNA molecule in 10 mM TRIS buffer with 1 mM spermine. The sudden drop at 105 s corresponds to condensation of the DNA molecule. b, Schematic representation of the energy barrier G‡ for the nucleation of single-molecule DNA condensation under tension. c–e, Condensation force, Fc, for an 8 kbp nicked DNA molecule as a function of spermine concentration in 1 mM TRIS (c), 10 mM TRIS (d) and in a buffer containing 10 mM TRIS and 50 mM KCl (e). The error bars are calculated as per Fig. 1. The different symbols in c are for two different molecules. The arrows indicate interpolated and extrapolated values of c0 from DLS measurements: 0.5, 1 and 6 mM for c, d and e, respectively. The lines are fits to equation (1) using measured values of c0, p=50 nm and a=1.5 nm. The black lines only use the Debye length of the monovalent buffer, whereas the blue lines include the multivalent ions in λD with a lower limit of 1 nm (size of the multivalent ions).
Figure 2c–e shows measurements of the condensation force, Fc, as a function of the spermine concentration in different monovalent electrolytes. Each curve exhibits an increase in Fc with increasing c up to a maximum at c=10−3–10−2 M, followed by a gradual decrease in Fc and a disappearance of condensation for c≳0.5 M. The gradual decrease at high concentration represents a more subtle form of re-entrant condensation19 at the single-molecule level20.
We repeated both electrophoresis and tweezers experiments with the trivalent ions cobalt sepulchrate ([CoC12H30N8]3+, cosep), cobalt hexamine ([Co(NH3)6]3+, cohex) and spermidine ([C7N3H22]3+). As shown in Fig. 3a,b, the mobility of DNA became less negative with increasing trivalent-ion concentration, c, and decreasing monovalent-buffer concentration. For cohex and spermidine, charge inversion had not yet occurred at the highest concentration that was experimentally accessible (6 mM). For cosep in 1 mM TRIS, μ became slightly positive (within one standard deviation of μ=0) at 6 mM. Figure 3c–e shows corresponding magnetic-tweezers measurements for the trivalent cations in 10 mM TRIS. The general trends are similar to those for spermine and Fc is again approximately parabolic in ln(c).
a,b, Electrophoretic mobility, μ, of condensed DNA as a function of multivalent-cation concentration, c, in 1 mM (a) and 10 mM (b) TRIS for spermine (black squares), cosep (red circles), cohex (blue triangles) and spermidine (green triangles). The DNA concentration and length are 5 ng μl−1 and 8 kbp, respectively. c–e, Condensation force, Fc, for an 8 kbp nicked DNA molecule as a function of multivalent-ion concentration, c, for the ions cosep (c), cohex (d) and spermidine (e) in a 10 mM TRIS buffer. Each concentration series was obtained on the same molecule. In c and e, data for two different molecules are shown as black squares and red open circles. The arrows indicate extrapolated values of c0 from DLS measurements (9, 14 and 26 mM for cosep, cohex and spermidine, respectively). Lines are fits of the force data to equation (1). f, Schematic representation of the geometry assumed in our model. All error bars are calculated as per Fig. 1.
The measured and extrapolated values of the charge-inversion concentration, c0 (green arrows in Figs 2 and 3), agree well with the peak in Fc(c) (except for the large value of Fc at 10 mM spermine in 1 mM TRIS). The high degree of coincidence between these two independently measured concentrations strongly suggests that charge inversion is responsible for the peak in Fc(c), consistent with the SCL theory result that the condensate free energy is minimal at c0 (ref. 12). The parabolic form of Fc(c) further agrees with predictions of the SCL model assuming a continuous, reversible condensation transition21. Because nucleation dynamics are observed, however, these predictions cannot be directly applied to our experimental situation. We therefore introduce a modified model that takes into account the barrier to nucleation.
Our model concentrates on the initial condensation event, before which the DNA molecule is in an extended conformation. Nucleation requires the formation of a loop, bringing two parts of the DNA into close contact and allowing the short-range attraction responsible for condensation to act18. Forming such a loop requires overcoming the electrostatic repulsion between two parts of the molecule with a range given by the Debye screening length, λD. We estimate the electrostatic energy required, Ge, by considering the approach of two like-charged cylinders of length l and radius a, as shown in Fig. 3f, and using the DNA effective potential φ=(k T/Z e)ln(c/c0) derived from the SCL model12. Here, k T is the thermal energy, Z is the multivalent-ion valence and −e is the electron charge. Ge∼φ2 is always positive except at the charge-inversion concentration, c0, where it vanishes; further details can be found in the Supplementary Information.
For condensation of a torsionally unconstrained molecule, the barrier takes the form G‡=Uloop+Ge+Gadd*. Here, Gadd* is an unknown constant, whereas  is the mechanical energy required for bending the DNA into a loop including the work against the force and p is the persistence length17. A positive electrostatic energy, Ge, must be compensated by a corresponding decrease in the loop energy, Uloop(F), yielding a simple expression for the condensation force as a function of multivalent-ion concentration
is the mechanical energy required for bending the DNA into a loop including the work against the force and p is the persistence length17. A positive electrostatic energy, Ge, must be compensated by a corresponding decrease in the loop energy, Uloop(F), yielding a simple expression for the condensation force as a function of multivalent-ion concentration

The parameter b is given by

where ε is the permittivity of water. b has a value in the range 0.5–3 μm in our experiments. For c near c0, equation (1) predicts a parabolic dependence of Fc on ln(c), with the maximum in Fc occurring at the charge-inversion concentration, c0. A more detailed geometry of the transition state yields a modified expression for b but preserves the dependence on c.
Figure 2c–e shows fits of the spermine data to equation (1). All fits use the same values of the fitting parameters l=40 nm and Fc(c0)=2.26 pN, whereas the values of c0 are set to the values measured by electrophoresis. A further complication is that the contribution of multivalent ions to λD, which becomes significant for c>c0, is not well established12. We therefore show two limiting cases, neglecting and fully including the contribution of the multivalent ions in the standard expression for λD. Figure 3c,d shows similar fits for the trivalent ions cosep and cohex. These fits again use l=40 nm, whereas c0 and Fc(c0) were fitted for each ion (c0=2 and 6 mM for cosep and cohex, respectively).
Our simple physical model correctly captures both the general form and the dependence of the concavity of Fc(c) on Z and λD. Furthermore, it correctly predicts the correspondence between c0 and the concentration where Fc(c) has its maximum. The fitted value of l=40 nm is of the expected order of magnitude, being comparable to the arc length of a half loop (17–32 nm for the force range F=3.5–1 pN). Some ion specificity is nonetheless evident in Fig. 3, particularly the data on spermidine which cannot be fitted to our model using reasonable small values of l (see Supplementary Information for further discussion).
It was suggested that re-entrant condensation in bulk experiments is due to incomplete dissociation of the multivalent ions22. This is not inconsistent with our observations: although incomplete dissociation is not significant at the concentrations where we observe charge inversion and the maximum in Fc(c), it may become relevant for c≫c0 where re-dissolution of bulk condensates is observed. In addition, for the trivalent ions in 10 mM TRIS our data cannot exclude the possibility that μ(c) remains negative (no charge inversion) but exhibits a maximum near 10 mM (ref. 15). In that case, the model predicts a qualitatively similar form for Fc(c).
To investigate the biological relevance of charge inversion and its relation to DNA condensation, we repeated our measurements using salmon protamine at physiological salt concentrations. Protamines are small basic proteins that condense DNA in the nucleus of spermatozoa, resulting in condensate morphologies comparable to those with multivalent ions2,3. It was recently shown using bulk measurements that short (150 bp) DNA fragments condensed by protamines in low salt (10 mM TRIS) exhibit charge inversion and de-condensation at high protamine concentrations23. Figure 4a shows that charge inversion of DNA condensates by protamines also occurs for long DNA fragments (8 kbp) at physiological salt concentrations (150 mM KCl). Furthermore, the condensation force measurements shown in Fig. 4b demonstrate that Fc decreases at high protamine concentration. This re-entrant condensation behaviour is shifted to higher protamine concentration with increasing salt, consistent with the DLS observations of charge inversion and with the behaviour with multivalent ions. Interestingly, DNA condensation by protamines is more robust to monovalent salt than that by multivalent ions24: as shown in Fig. 4c, Fc remains unaffected by monovalent salt up to 300 mM KCl with 1 ng μl−1 protamine in a manner consistent with its biological role.
a, Electrophoretic mobility, μ, of condensed DNA as a function of protamine concentration, cp, in 10 mM TRIS (black squares), 10 mM TRIS+50 mM KCl (red circles) and 10 mM TRIS+150 mM KCl (blue triangles). Charge inversion is observed in all three cases. The DNA concentration and length are 2 ng μl−1 and 8 kbp, respectively, except for the black open diamond where the DNA concentration is 4 ng μl−1 in 10 mM TRIS. b, Condensation force, Fc, for 8 kbp nicked DNA molecules as a function of protamine concentration in 10 mM TRIS (black squares and circles for two different molecules) and in 10 mM TRIS+150 mM KCl (red triangles). c, Condensation force as a function of KCl concentration in 10 mM TRIS with cp=1 ng μl−1 (different symbols correspond to different molecules). The condensation force is independent of salt concentration up to above typical physiological concentration, as shown by the arrow. Note that the protamine concentrations in the electrophoresis and tweezers measurements cannot be directly compared because, unlike the multivalent ions, the bulk protamine concentration is depleted by binding to DNA in the electrophoresis measurement (as shown by the 4 ng μl−1 DNA data in a). All error bars are calculated as per Fig. 1.
In summary, we demonstrated for the first time the charge inversion of DNA by multivalent ions. We further showed that the charge-inversion concentration coincides with the concentration most conducive to condensation of a single DNA molecule, lending considerable support to post-mean-field theories of DNA condensation. Corresponding trends are observed for condensation by protamines, suggesting a similar underlying mechanism, but the protein-induced condensation remains robust under physiological salt conditions.
Methods
Electrophoretic-mobility measurements were carried out using a Malvern ZetasizerNano ZS instrument using the M3-PALS technique. Equal volumes (0.5 ml) of solutions containing 7,922 bp DNA fragments and multivalent ions or protamines were mixed to a final concentration of 5 ng μl−1 DNA and the specified concentration of multivalent ions or protamines. Measurements were carried out after a 10 min incubation period. The quoted values of the charge-inversion concentration were obtained by linear interpolation or extrapolation on a lin-log scale of the two measurements nearest to μ=0.
For magnetic-tweezers measurements17, 8 kbp DNA constructs were prepared by ligating biotin- and digoxigenin-labelled DNA fragments (∼500 bp) to a 7,922 bp fragment. The biotin-labelled fragments were dephosphorylated before ligation to create torsionally unconstrained molecules. The constructs were then tethered between a 2.8 mm streptavidin-coated paramagnetic bead (Dynabeads, M-280 Streptavidin) and an anti-digoxigenin-coated glass surface that formed the wall of a 50 μl liquid cell. A nearby magnet permitted applying a known force, F, to the bead. The height of the bead above this surface (DNA extension, z) was monitored optically. A 3.2 mm polystyrene bead (Bangs Laboratories) bound to the surface was used as a reference for position tracking. Single-molecule DNA condensation measurements were carried out by measuring z in time while lowering F in discrete steps of 7% every 4 s. When z decreased below ∼1 μm, the force was rapidly increased to prevent the bead from sticking to the glass surface.
Phosphate buffer saline (PBS) containing 10 mM phosphate, 137 mM NaCl and 2.7 mM KCl at pH 7.4 was used for attaching reference beads to the surface. Standard buffer containing 10 mM phosphate, 10 mM NaN3, 0.2 mg ml−1 bovine serum albumin (BSA) and 0.1% Tween at pH 7.5 was used for attaching the DNA molecules to the surface. Measurements were done in a Tris hydroxymethylaminoethane monovalent buffer at pH 7.5 (TRIS) with varying concentrations of KCl and multivalent ions, c. All ions and the salmon protamine (grade IV) were ordered from Sigma and used as received. Solutions were changed by flushing at least 1 ml of the new solution through the liquid cell. Before condensation measurements, the standard buffer was removed by rinsing with 2 ml PBS, 1 ml 0.5 M KCl in TRIS, and 3 ml TRIS. This was done because BSA influenced the condensation dynamics, presumably owing to BSA clustering and adhering to DNA in the presence of multivalent ions.
References
Bloomfield, V. A. DNA condensation by multivalent cations. Biopolymers 44, 269–282 (1997).
Hud, N. V., Allen, M. J., Downing, K. H., Lee, J. & Balhorn, R. Identification of the elemental packing unit of DNA in mammalian sperm cells by atomic force microscopy. Biochem. Biophys. Res. Commun. 193, 1347–1354 (1993).
Ward, W. S. & Coffey, D. S. DNA packing and organization in mammalian spermatozoa: comparison with somatic cells. Biol. Reprod. 44, 569 (1991).
Thomas, T. & Thomas, T. J. Polyamines in cell growth and cell death: molecular mechanisms and therapeutic applications. Cell. Mol. Life Sci. 58, 244–258 (2001).
Hougaard, D. Polyamine cytochemistry: localization and possible functions of polyamines. Int. Rev. Cytol. 138, 51–88 (1992).
Gelbart, W. M., Bruinsma, R. F., Pincus, P. A. & Parsegian, V. A. DNA-inspired electrostatics. Phys. Today 53, 38–44 (2000).
Rouzina, I. & Bloomfield, V. A. Macroion attraction due to electrostatic correlation between screening counterions. 1. Mobile surface-adsorbed ions and diffuse ion cloud. J. Phys. Chem. 100, 9977–9989 (1996).
Oosawa, F. Interaction between parallel rodlike macroions. Biopolymers 6, 1633 (1968).
Kornyshev, A. A. & Leikin, S. Electrostatic zipper motif for DNA aggregation. Phys. Rev. Lett. 82, 4138–4141 (1999).
Shklovskii, B. I. Wigner crystal model of counterion induced bundle formation of rodlike polyelectrolytes. Phys. Rev. Lett. 82, 3268–3271 (1999).
Shklovskii, B. I. Screening of a macroion by multivalent ions: Correlation-induced inversion of charge. Phys. Rev. E 60, 5802–5811 (1999).
Nguyen, T. T., Rouzina, I. & Shklovskii, B. I. Reentrant condensation of DNA induced by multivalent counterions. J. Chem. Phys. 112, 2562–2568 (2000).
Tanaka, M. Electrophoresis of a rod macroion under polyelectrolyte salt: Is mobility reversed for DNA? J. Phys. Condens. Matter 16, S2127–S2134 (2004).
Tanaka, M. & Grosberg, A. Y. Electrophoresis of a charge-inverted macroion complex: molecular-dynamics study. Eur. Phys. J. E 7, 371–379 (2002).
Hsiao, P.-Y. & Luijten, E. Salt-induced collapse and reexpansion of highly charged flexible polyelectrolytes. Phys. Rev. Lett. 97, 148301 (2006).
van der Heyden, F. H. J., Stein, D., Besteman, K., Lemay, S. G. & Dekker, C. Charge inversion at high ionic strength studied by streaming currents. Phys. Rev. Lett. 96, 224502 (2006).
Strick, T. R., Allemand, J.-F., Croquette, V. & Bensimon, D. Twisting and stretching single DNA molecules. Prog. Biophys. Mol. Biol. 74, 115–140 (2000).
Besteman, K., Hage, S., Dekker, N. H. & Lemay, S. G. Role of tension and twist in single-molecule DNA condensation. Phys. Rev. Lett. 98, 058103 (2007).
Pelta, J., Livolant, F. & Sikorav, J.-L. DNA aggregation induced by polyamines and cobalthexamine. J. Biol. Chem. 271, 5656–5662 (1996).
Murayama, Y., Sakamaki, Y. & Sano, M. Elastic response of single DNA molecules exhibits a reentrant collapsing transition. Phys. Rev. Lett. 90, 018102 (2003).
Zhang, R. & Shklovskii, B. I. The pulling force of a single DNA molecule condensed by spermidine. Physica A 349, 563–570 (2005).
Yang, J. & Rau, D. C. Incomplete ion dissociation underlies the weakened attraction between DNA helices at high spermidine concentrations. Biophys. J. 89, 1932–1940 (2005).
Raspaud, E., Pelta, J., de Frutos, M. & Livolant, F. Solubility and charge inversion of complexes of DNA and basic proteins. Phys. Rev. Lett. 97, 068103 (2006).
Raspaud, E., Olvera de la Cruz, M., Sikorav, J.-L. & Livolant, F. Precipitation of DNA by polyamines: A polyelectrolyte behavior. Biophys. J. 74, 381 (1998).
Acknowledgements
We thank B. I. Shklovskii for useful discussions and C. Dekker for general support. This work was funded by FOM and NWO.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Besteman, K., Van Eijk, K. & Lemay, S. Charge inversion accompanies DNA condensation by multivalent ions. Nature Phys 3, 641–644 (2007). https://doi.org/10.1038/nphys697
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nphys697
This article is cited by
-
Recent Developments on Ionic Liquids and Deep Eutectic Solvents for Drug Delivery Applications
Pharmaceutical Research (2022)
-
Direct visualization of local activities of long DNA strands via image–time correlation
European Biophysics Journal (2021)
-
Self-assembly of highly ordered DNA origami lattices at solid-liquid interfaces by controlling cation binding and exchange
Nano Research (2020)
-
Gold-Hybridized Zinc Oxide Nanorods as Real-Time Low-Cost NanoBiosensors for Detection of virulent DNA signature of HPV-16 in Cervical Carcinoma
Scientific Reports (2019)
-
Direct evidence for sequence-dependent attraction between double-stranded DNA controlled by methylation
Nature Communications (2016)