Abstract
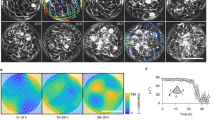
The shape and differentiated state of many cell types are highly sensitive to the rigidity of the microenvironment. The physical mechanisms involved, however, are unknown. Here, we present a theoretical model and experiments demonstrating that the alignment of stress fibres within stem cells is a non-monotonic function of matrix rigidity. We treat the cell as an active elastic inclusion in a surrounding matrix, allowing the actomyosin forces to polarize in response to elastic stresses developed in the cell. The theory correctly predicts the monotonic increase of the cellular forces with the matrix rigidity and the alignment of stress fibres parallel to the long axis of cells. We show that the anisotropy of this alignment depends non-monotonically on matrix rigidity and demonstrate it experimentally by quantifying the orientational distribution of stress fibres in stem cells. These findings offer physical insight into the sensitivity of stem-cell differentiation to tissue elasticity and, more generally, introduce a cell-type-specific parameter for actomyosin polarizability.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
O’Neill, C., Jordan, P. & Ireland, G. Evidence for two distinct mechanisms of anchorage stimulation in freshly explanted and 3t3 swiss mouse fibroblasts. Cell 44, 489–496 (1986).
Chen, C. S., Mrksich, M., Huang, S., Whitesides, G. M. & Ingber, D. E. Geometric control of cell life and death. Science 276, 1425–1428 (1997).
McBeath, R., Pirone, D. M., Nelson, C. M., Bhadriraju, K. & Chen, C. S. Cell shape, cytoskeleton tension, and RohA regulate stem cell lineage commitment. Dev. Cell 6, 483–495 (2004).
Engler, A. J. et al. Myotubes differentiate optimally on substrates with tissue-like stiffness: Pathological implications for soft or stiff microenvironments. J. Cell Biol. 166, 877–887 (2004).
Yeung, T. et al. Effects of substrate stiffness on cell morphology, cytoskeletal structure, and adhesion. Cell Motil. Cytoskeleton 60, 24–34 (2005).
Discher, D. E., Janmey, P. & Wang, Y. Tissue cells feel and respond to the stiffness of their substrate. Science 310, 1139–1143 (2005).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Georges, P. C., Miller, W. J., Meaney, D. F., Sawyer, E. S. & Janmey, P. A. Matrices with compliance comparable to that of brain tissue select neuronal over glial growth in mixed cortical cultures. Biophys. J. 90, 3012–3018 (2006).
Cai, Y. et al. Cytoskeletal coherence requires myosin-IIA contractility. J. Cell Sci. 123, 413–423 (2010).
Bhattacharya, D., Talwar, S., Mazumder, A. & Shivashankar, G. V. Spatio-temporal plasticity in chromatin organization in mouse cell differentiation and during drosophila embryogenesis. Biophys. J. 96, 3832–3839 (2009).
Wang, N., Tytell, J. D. & Ingber, D. E. Mechanotransduction at a distance: Mechanically coupling the extracellular matrix with the nucleus. Nat. Rev. Mol. Cell Biol. 10, 75–82 (2009).
Bershadsky, A., Kozlov, M. & Geiger, B. Adhesion-mediated mechanosensitivity: A time to experiment, and a time to theorize. Curr. Opin. Cell Biol. 18, 472–481 (2006).
Iba, T. & Sumpio, B. Morphological response of human endothelial cells subjected to cyclic strain in vitro. Microvasc. Res. 42, 245–254 (1991).
Rodriguez, J. P., Gonzalez, M., Rios, S. & Cambiazo, V. Cytoskeletal organization of human mesenchymal stem cells (msc) changes during their osteogenic differentiation. J. Cell. Biochem. 93, 721–731 (2004).
Curtis, A., Aitchison, G. & Tsapikouni, T. Orthogonal (transverse) arrangements of actin in endothelia and fibroblasts. J. R. Soc. Interface 3, 753–756 (2006).
Ghibaudo, M. et al. Traction forces and rigidity sensing regulate cell functions. Soft Matter 4, 1836–1843 (2008).
Kumar, S. et al. Viscoelastic retraction of single living stress fibres and its impact on cell shape, cytoskeletal organization, and extracellular matrix mechanics. Biophys. J. 90, 3762–3773 (2006).
Dubin-Thaler, B. J. et al. Quantification of cell edge velocities and traction forces reveals distinct motility modules during cell spreading. PLOS One 3, e3735 (2008).
Griffin, M. A. et al. Patterning, prestress, and peeling dynamics of myocytes. Biophys. J. 86, 1209–1222 (2004).
Wang, N. et al. Cell prestress. i. Stiffness and prestress are closely associated in adherent contractile cells. Am. J. Physiol. Cell Physiol. 282, C606–C616 (2002).
Wang, N., Ostuni, E., Whitesides, G. M. & Ingber, D. E. Micropatterning tractional forces in living cells. Cell Motil. Cyto. 52, 97–106 (2002).
Chicurel, M. E., Chen, C. S. & Ingber, D. E. Cellular control lies in the balance of forces. Curr. Opin. Cell Biol. 10, 232–239 (1998).
Balaban, N. Q. et al. Force and focal adhesion assembly: A close relationship studied using elastic micropatterned substrates. Nature Cell Biol. 3, 466–472 (2001).
Schwarz, U. S., Erdmann, T. & Bischofs, I. B. Focal adhesions as mechanosensors: The two-spring model. Biosystems 83, 225–232 (2006).
Grinnell, F. Fibroblast-collagen-matrix contraction: Growth-factor signalling and mechanical loading. Trends Cell Biol. 10, 362–365 (2000).
Eshelby, J. D. The determination of elastic field of an ellipsoidal inclusion, and related problems. Proc. R. Soc. A 241, 376–396 (1957).
Mura, T. Micromechanics of Defects in Solids (Kluwer Academic, 1991).
Landau, L. D. & Lifshitz, E. M. Theory of Elasticity 3rd edn (Course of Theoretical Physics, Vol. 7, Reed Educational and Professional Publishing, 1986).
Jaswon, M. A. & Bhargava, R. D. Two-dimensional elastic inclusion problems. Proc. Camb. Phil. Soc. 57, 669–680 (1961).
Siems, R. Mechanical interactions of point defects. Phys. Status Solidi 30, 645–658 (1968).
Schwarz, U. S. & Safran, S. A. Elastic interactions of cells. Phys. Rev. Lett. 88, 048102 (2002).
Carlsson, A. E. Contractile stress generation by actomyosin gels. Phys. Rev. E 74, 051912 (2006).
Riveline, D. et al. Focal contacts as mechanosensors: Externally applied local mechanical force induces growth of focal contacts by an mdia1-dependent and rock-independent mechanism. J. Cell Biol. 153, 1175–1185 (2001).
Schwarz, U. S. & Bischofs, I. B. Physical determinants of cell organization in soft media. Med. Eng. Phys. 27, 763–772 (2005).
Pelham, R. J. & Wang, Y. L. Cell locomotion and focal adhesions are regulated by substrate flexibility. Proc. Natl Acad. Sci. USA 94, 13661–13665 (1997).
Engler, A. J., Rehfeldt, F., Sen, S. & Discher, D. E. Microtissue Elasticity: Measurements by Atomic Force Microscopy and its Influence on Cell Differentiation Vol. 83, 521–545 (Academic, 2007).
Rasband, W. S. ImageJ US National Institute of Health, Bethesda, Maryland, USA, <http://rsb.info.nih.gov/ij/>, 1997–2007.
Haralick, R. & Shapiro, L. Computer and Robot Vision Vol. 1 (Addison-Wesley, 1992).
Otsu, N. Threshold selection method from grey-level histograms. IEEE Trans. Syst. Man Cybernetics 9, 62–66 (1979).
Acknowledgements
We thank R. De, R. Paul and N. Gov for many useful discussions. We are grateful to the Israel Science Foundation, the Clore Center for Biological Physics, the Schmidt Minerva Center and an EU Network grant for their support. F.R. gratefully acknowledges financial support through the Feodor Lynen fellowship of the Alexander von Humboldt Foundation. D.E.D. thanks NFS and NIH. A.E.X.B. was supported by a scholarship from the Natural Sciences and Engineering Research Council of Canada.
Author information
Authors and Affiliations
Contributions
A.Z. and S.A.S. developed the theory. F.R., A.E.X.B. and D.E.D. designed the experiments; F.R. carried out the experiments; A.E.X.B. wrote the image analysis algorithm. All authors analysed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 453 kb)
Rights and permissions
About this article
Cite this article
Zemel, A., Rehfeldt, F., Brown, A. et al. Optimal matrix rigidity for stress-fibre polarization in stem cells. Nature Phys 6, 468–473 (2010). https://doi.org/10.1038/nphys1613
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nphys1613
This article is cited by
-
Combined computational modeling and experimental study of the biomechanical mechanisms of platelet-driven contraction of fibrin clots
Communications Biology (2023)
-
Investigating the nature of active forces in tissues reveals how contractile cells can form extensile monolayers
Nature Materials (2021)
-
What factors determine the number of nonmuscle myosin II in the sarcomeric unit of stress fibers?
Biomechanics and Modeling in Mechanobiology (2021)
-
Stimuli-responsive hydrogels as a model of the dynamic cellular microenvironment
Polymer Journal (2020)
-
Encapsulated piezoelectric nanoparticle–hydrogel smart material to remotely regulate cell differentiation and proliferation: a finite element model
Computational Mechanics (2019)