Abstract
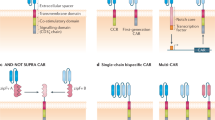
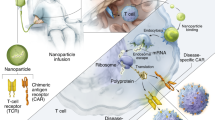
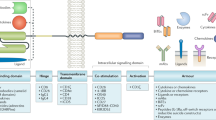
An emerging approach for treating cancer involves programming patient-derived T cells with genes encoding disease-specific chimeric antigen receptors (CARs), so that they can combat tumour cells once they are reinfused. Although trials of this therapy have produced impressive results, the in vitro methods they require to generate large numbers of tumour-specific T cells are too elaborate for widespread application to treat cancer patients. Here, we describe a method to quickly program circulating T cells with tumour-recognizing capabilities, thus avoiding these complications. Specifically, we demonstrate that DNA-carrying nanoparticles can efficiently introduce leukaemia-targeting CAR genes into T-cell nuclei, thereby bringing about long-term disease remission. These polymer nanoparticles are easy to manufacture in a stable form, which simplifies storage and reduces cost. Our technology may therefore provide a practical, broadly applicable treatment that can generate anti-tumour immunity ‘on demand’ for oncologists in a variety of settings.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rosenberg, S. A. & Restifo, N. P. Adoptive cell transfer as personalized immunotherapy for human cancer. Science 348, 62–68 (2015).
Plumridge, H. New costly cancer treatments face hurdles getting to patients. The Wall Street Journal (6 October 2014); https://www.wsj.com/articles/new-costly-cancer-treatments-face-hurdles-getting-to-patients-1412627150
Mangraviti, A. et al. Polymeric nanoparticles for nonviral gene therapy extend brain tumor survival in vivo. ACS Nano 9, 1236–1249 (2015).
Narayanan, K. et al. Mimicking cellular transport mechanism in stem cells through endosomal escape of new peptide-coated quantum dots. Sci. Rep. 3, 2184 (2013).
Maude, S. L. et al. Chimeric antigen receptor T cells for sustained remissions in leukemia. N. Engl. J. Med. 371, 1507–1517 (2014).
Davila, M. L., Kloss, C. C., Gunset, G. & Sadelain, M. CD19 CAR-targeted T cells induce long-term remission and B cell aplasia in an immunocompetent mouse model of B cell acute lymphoblastic leukemia. PLoS ONE 8, e61338 (2013).
Nakazawa, Y. et al. Evaluation of long-term transgene expression in piggyBac-modified human T lymphocytes. J. Immunother. 36, 3–10 (2013).
Burnight, E. R. et al. A hyperactive transposase promotes persistent gene transfer of a piggyBac DNA transposon. Mol. Ther. Nucleic Acids 1, e50 (2012).
Gade, T. P. et al. Targeted elimination of prostate cancer by genetically directed human T lymphocytes. Cancer Res. 65, 9080–9088 (2005).
Maude, S. L., Teachey, D. T., Porter, D. & Grupp, S. A. CD19-targeted chimeric antigen receptor T-cell therapy for acute lymphoblastic leukemia. Blood 125, 4017–4023 (2015).
Grupp, S. A. et al. Chimeric antigen receptor-modified T cells for acute lymphoid leukemia. N. Engl. J. Med. 368, 1509–1518 (2013).
Rosenberg, S. A. Cell transfer immunotherapy for metastatic solid cancer—what clinicians need to know. Nat. Rev. Clin. Oncol. 8, 577–585 (2011).
Wang, X. et al. Large-scale clinical-grade retroviral vector production in a fixed-bed bioreactor. J. Immunother. 38, 127–135 (2015).
Kuchenbaecker, K. B. et al. Identification of six new susceptibility loci for invasive epithelial ovarian cancer. Nat. Genet. 47, 164–171 (2015).
Johnson, L. A. et al. Rational development and characterization of humanized anti-EGFR variant III chimeric antigen receptor T cells for glioblastoma. Sci. Transl. Med. 7, 275ra222 (2015).
Magnani, C. F. et al. Immunotherapy of acute leukemia by chimeric antigen receptor-modified lymphocytes using an improved sleeping beauty transposon platform. Oncotarget 7, 51581–51597 (2016).
Abate-Daga, D. et al. A novel chimeric antigen receptor against prostate stem cell antigen mediates tumor destruction in a humanized mouse model of pancreatic cancer. Hum. Gene Ther. 25, 1003–1012 (2014).
Sadelain, M. CAR therapy: the CD19 paradigm. J. Clin. Invest. 125, 3392–3400 (2015).
Meacham, C. E. & Morrison, S. J. Tumour heterogeneity and cancer cell plasticity. Nature 501, 328–337 (2013).
Li, X. et al. Nanoparticle-mediated transcriptional modification enhances neuronal differentiation of human neural stem cells following transplantation in rat brain. Biomaterials 84, 157–166 (2016).
Kim, J., Kang, Y., Tzeng, S. Y. & Green, J. J. Synthesis and application of poly(ethylene glycol)-co-poly(beta-amino ester) copolymers for small cell lung cancer gene therapy. Acta Biomater. 41, 293–301 (2016).
Zhang, X., Edwards, J. P. & Mosser, D. M. The expression of exogenous genes in macrophages: obstacles and opportunities. Methods Mol. Biol. 531, 123–143 (2009).
Marodon, G. et al. Specific transgene expression in human and mouse CD4+ cells using lentiviral vectors with regulatory sequences from the CD4 gene. Blood 101, 3416–3423 (2003).
Ellmeier, W., Sunshine, M. J., Losos, K., Hatam, F. & Littman, D. R. An enhancer that directs lineage-specific expression of CD8 in positively selected thymocytes and mature T cells. Immunity 7, 537–547 (1997).
Wu, C. Y., Rupp, L. J., Roybal, K. T. & Lim, W. A. Synthetic biology approaches to engineer T cells. Curr. Opin. Immunol. 35, 123–130 (2015).
Kebriaei, P. et al. Phase I trials using sleeping beauty to generate CD19-specific CAR T cells. J. Clin. Invest. 126, 3363–3376 (2016).
Monjezi, R. et al. Enhanced CAR T-cell engineering using non-viral sleeping beauty transposition from minicircle vectors. Leukemia 31, 186–194 (2017).
Brooks, P. J., Yang, N. N. & Austin, C. P. Gene therapy: the view from NCATS. Hum. Gene Ther. 27, 7–13 (2016).
Gaspar, V. et al. Minicircle DNA vectors for gene therapy: advances and applications. Expert Opin. Biol. Ther. 15, 353–379 (2015).
Dobrenkov, K. et al. Monitoring the efficacy of adoptively transferred prostate cancer-targeted human T lymphocytes with PET and bioluminescence imaging. J. Nucl. Med. 49, 1162–1170 (2008).
Fischer, K., Andreesen, R. & Mackensen, A. An improved flow cytometric assay for the determination of cytotoxic T lymphocyte activity. J. Immunol. Methods 259, 159–169 (2002).
May, C. et al. Therapeutic haemoglobin synthesis in β-thalassaemic mice expressing lentivirus-encoded human β-globin. Nature 406, 82–86 (2000).
Acknowledgements
We thank I. Stanishevskaya and D. Ehlert (cognitionstudio.com) for the design of the illustrations. We also thank M. Sadelain (Memorial Sloan-Kettering Cancer Center, New York, New York) for the Eμ-ALL01 cell line, and for the DNA construct that encodes an all-murine CD19-specific CAR. This work was supported in part by the Fred Hutchinson Cancer Research Center's Immunotherapy Initiative with funds provided by the Bezos Family Foundation, a New Idea Award from the Leukemia & Lymphoma Society, the Phi Beta Psi Sorority, the National Science Foundation (CAREER, award no. 1452492 and EAGER award no. 1644363), and the National Cancer Institute of the National Institutes of Health under award no. R01CA207407. M.T.S. was also supported by a Research Scholar Grant (RSG-16-110-01 – LIB) from the American Cancer Society.
Author information
Authors and Affiliations
Contributions
T.T.S., S.B.S. and H.F.M. designed and performed experiments and analysed and interpreted data. L.E.M. helped clone plasmid vectors, W.J. synthesized the PBAE polymer, and D.R. and E.B. helped with the large-scale purification of CAR-encoding plasmid DNA. M.E.W. performed the Southern blot analysis. S.P.S.P. performed and analysed in vivo safety/toxicity studies, and M.T.S. designed the study, performed experiments, analysed and interpreted data, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 2240 kb)
Rights and permissions
About this article
Cite this article
Smith, T., Stephan, S., Moffett, H. et al. In situ programming of leukaemia-specific T cells using synthetic DNA nanocarriers. Nature Nanotech 12, 813–820 (2017). https://doi.org/10.1038/nnano.2017.57
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2017.57
This article is cited by
-
Biomaterials to enhance adoptive cell therapy
Nature Reviews Bioengineering (2024)
-
In vivo human T cell engineering with enveloped delivery vehicles
Nature Biotechnology (2024)
-
Drug delivery methods for cancer immunotherapy
Drug Delivery and Translational Research (2024)
-
In vivo manufacture and manipulation of CAR-T cells for better druggability
Cancer and Metastasis Reviews (2024)
-
CAR-T cell therapy: a game-changer in cancer treatment and beyond
Clinical and Translational Oncology (2024)