Abstract
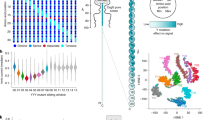
The simultaneous detection of a large number of different analytes is important in bionanotechnology research and in diagnostic applications. Nanopore sensing is an attractive method in this regard as the approach can be integrated into small, portable device architectures, and there is significant potential for detecting multiple sub-populations in a sample. Here, we show that highly multiplexed sensing of single molecules can be achieved with solid-state nanopores by using digitally encoded DNA nanostructures. Based on the principles of DNA origami, we designed a library of DNA nanostructures in which each member contains a unique barcode; each bit in the barcode is signalled by the presence or absence of multiple DNA dumbbell hairpins. We show that a 3-bit barcode can be assigned with 94% accuracy by electrophoretically driving the DNA structures through a solid-state nanopore. Select members of the library were then functionalized to detect a single, specific antibody through antigen presentation at designed positions on the DNA. This allows us to simultaneously detect four different antibodies of the same isotype at nanomolar concentration levels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Wanunu, M. Nanopores: A journey towards DNA sequencing. Phys. Life Rev. 9, 125–158 (2012).
Li, W. et al. Single protein molecule detection by glass nanopores. ACS Nano 7, 4129–4134 (2013).
Plesa, C. et al. Fast translocation of proteins through solid state nanopores. Nano Lett. 13, 658–663 (2013).
Firnkes, M., Pedone, D., Knezevic, J., Döblinger, M. & Rant, U. Electrically facilitated translocations of proteins through silicon nitride nanopores: conjoint and competitive action of diffusion, electrophoresis, and electroosmosis. Nano Lett. 10, 2162–2167 (2010).
Bayley, H. & Martin, C. R. Resistive-pulse sensing from microbes to molecules. Chem. Rev. 100, 2575–2594 (2000).
Bayley, H. & Cremer, P. S. Stochastic sensors inspired by biology. Nature 413, 226–230 (2001).
Braha, O. et al. Designed protein pores as components for biosensors. Chem. Biol. 4, 497–505 (1997).
Rotem, D., Jayasinghe, L., Salichou, M. & Bayley, H. Protein detection by nanopores equipped with aptamers. J. Am. Chem. Soc. 134, 2781–2787 (2012).
Movileanu, L., Howorka, S., Braha, O. & Bayley, H. Detecting protein analytes that modulate transmembrane movement of a polymer chain within a single protein pore. Nature Biotechnol. 18, 1091–1095 (2000).
Wei, R., Gatterdam, V., Wieneke, R., Tampé, R. & Rant, U. Stochastic sensing of proteins with receptor-modified solid-state nanopores. Nature Nanotech. 7, 257–263 (2012).
Harrington, L., Cheley, S., Alexander, L. T., Knapp, S. & Bayley, H. Stochastic detection of Pim protein kinases reveals electrostatically enhanced association of a peptide substrate. Proc. Natl Acad. Sci. USA 110, E4417–E4426 (2013).
Li, T., Liu, L., Li, Y., Xie, J. & Wu, H.-C. A universal strategy for aptamer-based nanopore sensing through host-guest interactions inside α-hemolysin. Angew. Chem. 127, 7678–7681 (2015).
Kawano, R. et al. Rapid detection of a cocaine-binding aptamer using biological nanopores on a chip. J. Am. Chem. Soc. 133, 8474–8477 (2011).
Kasianowicz, J. J., Henrickson, S. E., Weetall, H. H. & Robertson, B. Simultaneous multianalyte detection with a nanometer-scale pore. Anal. Chem. 73, 2268–2272 (2001).
Bell, N. A. W. & Keyser, U. F. Specific protein detection using designed DNA carriers and nanopores. J. Am. Chem. Soc. 137, 2035–2041 (2015).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Steinbock, L. J., Otto, O., Chimerel, C., Gornall, J. & Keyser, U. F. Detecting DNA folding with nanocapillaries. Nano Lett. 10, 2493–2497 (2010).
Bell, N. A. W., Muthukumar, M. & Keyser, U. F. Translocation frequency of double-stranded DNA through a solid-state nanopore. Phys. Rev. E 93, 022401 (2016).
Smeets, R. M. M., Keyser, U. F., Dekker, N. H. & Dekker, C. Noise in solid-state nanopores. Proc. Natl Acad. Sci. USA 105, 417–421 (2008).
Tabard-Cossa, V., Trivedi, D., Wiggin, M., Jetha, N. N. & Marziali, A. Noise analysis and reduction in solid-state nanopores. Nanotechnology 18, 305505 (2007).
Storm, A., Chen, J., Zandbergen, H. & Dekker, C. Translocation of double-strand DNA through a silicon oxide nanopore. Phys. Rev. E 71, 1–10 (2005).
Muthukumar, M. Mechanism of DNA transport through pores. Annu. Rev. Biophys. Biomol. Struct. 36, 435–450 (2007).
Chen, P. et al. Probing single DNA molecule transport using fabricated nanopores. Nano Lett. 4, 2293–2298 (2004).
Lu, B., Albertorio, F., Hoogerheide, D. P. & Golovchenko, J. A. Origins and consequences of velocity fluctuations during DNA passage through a nanopore. Biophys. J. 101, 70–79 (2011).
Saphire, E. O. et al. Contrasting IgG structures reveal extreme asymmetry and flexibility. J. Mol. Biol. 319, 9–18 (2002).
Preiner, J. et al. IgGs are made for walking on bacterial and viral surfaces. Nature Commun. 5, 1–8 (2014).
Mammen, M., Choi, S.-K. & Whitesides, G. M. Polyvalent interactions in biological systems: implications for design and use of multivalent ligands and inhibitors. Angew. Chem. Int. Ed. 37, 2754–2794 (1998).
Kauert, D. J., Kurth, T., Liedl, T. & Seidel, R. Direct mechanical measurements reveal the material properties of 3D DNA-origami. Nano Lett. 11, 5558–5563 (2011).
Castro, C. E., Su, H.-J., Marras, A. E., Zhou, L. & Johnson, J. Mechanical design of DNA nanostructures. Nanoscale 7, 5913–5921 (2015).
McMullen, A., de Haan, H. W., Tang, J. X. & Stein, D. Stiff filamentous virus translocations through solid-state nanopores. Nature Commun. 5, 4171 (2014).
Marchi, A. N., Saaem, I., Vogen, B. N., Brown, S. & LaBean, T. H. Towards larger DNA origami. Nano Lett 14, 5740–5747 (2014).
Rosenstein, J. K., Wanunu, M., Merchant, C. A., Drndic, M. & Shepard, K. L. Integrated nanopore sensing platform with sub-microsecond temporal resolution. Nature Methods 9, 487–492 (2012).
Rodriguez-Larrea, D. & Bayley, H. Multistep protein unfolding during nanopore translocation. Nature Nanotech. 8, 288–295 (2013).
Rosen, C. B., Rodriguez-Larrea, D. & Bayley, H. Single-molecule site-specific detection of protein phosphorylation with a nanopore. Nature Biotechnol. 32, 179–181 (2014).
Nivala, J., Marks, D. B. & Akeson, M. Unfoldase-mediated protein translocation through an α-hemolysin nanopore. Nature Biotechnol. 31, 247–250 (2013).
Nivala, J., Mulroney, L., Li, G., Schreiber, J. & Akeson, M. Discrimination among protein variants using an unfoldase-coupled nanopore. ACS Nano 8, 12365–12375 (2014).
Fu, J., Liu, M., Liu, Y. & Yan, H. Spatially-interactive biomolecular networks organized by nucleic acid nanostructures. Acc. Chem. Res. 45, 1215–1226 (2012).
Pippig, D. A., Baumann, F., Strackharn, M., Aschenbrenner, D. & Gaub, H. E. Protein-DNA chimeras for nano assembly. ACS Nano 8, 6551–6555 (2014).
Bell, N. A. W. et al. Multiplexed ionic current sensing with glass nanopores. Lab Chip 13, 1859–1862 (2013).
Acknowledgements
The authors thank J. Kong and V. Thacker for useful discussions. N.A.W.B. and U.F.K. acknowledge funding from an ERC starting grant (Passmembrane 261101) and an ERC consolidator grant (Designerpores 647144). N.A.W.B. also acknowledges funding from an EPSRC doctoral prize award.
Author information
Authors and Affiliations
Contributions
N.A.W.B. conceived the idea, N.A.W.B. and U.F.K. designed the experiments. N.A.W.B. performed the experiments and analysed the data, and N.A.W.B. and U.F.K. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors have made a patent application related to this work.
Supplementary information
Supplementary information
Supplementary information (PDF 2345 kb)
Rights and permissions
About this article
Cite this article
Bell, N., Keyser, U. Digitally encoded DNA nanostructures for multiplexed, single-molecule protein sensing with nanopores. Nature Nanotech 11, 645–651 (2016). https://doi.org/10.1038/nnano.2016.50
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2016.50
This article is cited by
-
Simultaneous identification of viruses and viral variants with programmable DNA nanobait
Nature Nanotechnology (2023)
-
Probing when dCas9 tolerates DNA mismatches
Nature Biomedical Engineering (2023)
-
Discriminating protein tags on a dsDNA construct using a Dual Nanopore Device
Scientific Reports (2022)
-
Functionalizing DNA origami to investigate and interact with biological systems
Nature Reviews Materials (2022)
-
Localised solid-state nanopore fabrication via controlled breakdown using on-chip electrodes
Nano Research (2022)