Abstract
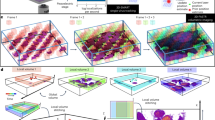
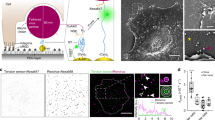
Viral infection is initiated when a virus binds to cell surface receptors. Because the cell membrane is dynamic and heterogeneous, imaging living cells and simultaneously quantifying the first viral binding events is difficult. Here, we show an atomic force and confocal microscopy set-up that allows the surface receptor landscape of cells to be imaged and the virus binding events within the first millisecond of contact with the cell to be mapped at high resolution (<50 nm). We present theoretical approaches to contour the free-energy landscape of early binding events between an engineered virus and cell surface receptors. We find that the first bond formed between the viral glycoprotein and its cognate cell surface receptor has relatively low lifetime and free energy, but this increases as additional bonds form rapidly (≤1 ms). The formation of additional bonds occurs with positive allosteric modulation and the three binding sites of the viral glycoprotein are quickly occupied. Our quantitative approach can be readily applied to study the binding of other viruses to animal cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Dimitrov, D. S. Virus entry: molecular mechanisms and biomedical applications. Nat. Rev. Microbiol. 2, 109–122 (2004).
Smith, A. E. & Helenius, A. How viruses enter animal cells. Science 304, 237–242 (2004).
Brandenburg, B. & Zhuang, X. Virus trafficking—learning from single-virus tracking. Nat. Rev. Microbiol. 5, 197–208 (2007).
McMahon, H. T. & Gallop, J. L. Membrane curvature and mechanisms of dynamic cell membrane remodelling. Nature 438, 590–596 (2005).
Lingwood, D. & Simons, K. Lipid rafts as a membrane-organizing principle. Science 327, 46–50 (2010).
Chojnacki, J. et al. Maturation-dependent HIV-1 surface protein redistribution revealed by fluorescence nanoscopy. Science 338, 524–528 (2012).
Kukura, P. et al. High-speed nanoscopic tracking of the position and orientation of a single virus. Nat. Methods 6, 923–927 (2009).
Sun, E., He, J. & Zhuang, X. Live cell imaging of viral entry. Curr. Opin. Virol. 3, 34–43 (2013).
Wickersham, I. R. et al. Monosynaptic restriction of transsynaptic tracing from single, genetically targeted neurons. Neuron 53, 639–647 (2007).
Wall, N. R., Wickersham, I. R., Cetin, A., De La Parra, M. & Callaway, E. M. Monosynaptic circuit tracing in vivo through Cre-dependent targeting and complementation of modified rabies virus. Proc. Natl Acad. Sci. USA 107, 21848–21853 (2010).
Cronin, J., Zhang, X.-Y. & Reiser, J. Altering the tropism of lentiviral vectors through pseudotyping. Curr. Gene Therapy 5, 387–398 (2005).
Ginger, M., Haberl, M., Conzelmann, K. K., Schwarz, M. K. & Frick, A. Revealing the secrets of neuronal circuits with recombinant rabies virus technology. Front. Neural Circuits 7, 1–15 (2013).
Medalsy, I., Hensen, U. & Muller, D. J. Imaging and quantifying chemical and physical properties of native proteins at molecular resolution by force–volume AFM. Angew. Chem. Int. Ed. 50, 12103–12108 (2011).
Dufrêne, Y. F., Martínez-Martín, D., Medalsy, I., Alsteens, D. & Müller, D. J. Multiparametric imaging of biological systems by force–distance curve-based AFM. Nat. Methods 10, 847–854 (2013).
Alsteens, D., Trabelsi, H., Soumillion, P. & Dufrêne, Y. F. Multiparametric atomic force microscopy imaging of single bacteriophages extruding from living bacteria. Nat. Commun. 4, 2926 (2013).
Pfreundschuh, M., Martinez-Martin, D., Mulvihill, E., Wegmann, S. & Muller, D. J. Multiparametric high-resolution imaging of native proteins by force–distance curve-based AFM. Nat. Protoc. 9, 1113–1130 (2014).
Alsteens, D. et al. Imaging G protein-coupled receptors while quantifying their ligand-binding free-energy landscape. Nat. Methods 12, 845–851 (2015).
Pfreundschuh, M. et al. Identifying and quantifying two ligand-binding sites while imaging native human membrane receptors by AFM. Nat. Commun. 6, 8857 (2015).
Banerjee, I., Yamauchi, Y., Helenius, A. & Horvath, P. High-content analysis of sequential events during the early phase of influenza A virus infection. PLoS ONE 8, e68450 (2013).
Poole, K., Meder, D., Simons, K. & Müller, D. The effect of RAFT lipid depletion on microvilli formation in MDCK cells, visualized by atomic force microscopy. FEBS Lett. 565, 53–58 (2004).
Rankl, C. et al. Multiple receptors involved in human rhinovirus attachment to live cells. Proc. Natl Acad. Sci. USA 105, 17778–17783 (2008).
Sieben, C. et al. Influenza virus binds its host cell using multiple dynamic interactions. Proc. Natl Acad. Sci. USA 109, 13626–13631 (2012).
Oesterhelt, F., Rief, M. & Gaub, H. E. Single molecule force spectroscopy by AFM indicates helical structure of poly(ethylene-glycol) in water. New J. Phys. 1, 6.1–6.11 (1999).
Evans, E. & Ritchie, K. Dynamic strength of molecular adhesion bonds. Biophys. J. 72, 1541–1555 (1997).
Evans, E. A. & Calderwood, D. A. Forces and bond dynamics in cell adhesion. Science 316, 1148–1153 (2007).
Dobrowsky, T. M., Zhou, Y., Sun, S. X., Siliciano, R. F. & Wirtz, D. Monitoring early fusion dynamics of human immunodeficiency virus type 1 at single-molecule resolution. J. Virol. 82, 7022–7033 (2008).
Friddle, R. W., Noy, A. & De Yoreo, J. J. Interpreting the widespread nonlinear force spectra of intermolecular bonds. Proc. Natl Acad. Sci. USA 109, 13573–13578 (2012).
Damico, R. L., Crane, J. & Bates, P. Receptor-triggered membrane association of a model retroviral glycoprotein. Proc. Natl Acad. Sci. USA 95, 2580–2585 (1998).
Mammen, M., Choi, S. K. & Whitesides, G. M. Polyvalent interactions in biological systems: implications for design and use of multivalent ligands and inhibitors. Angew. Chem. Int. Ed. 37, 2754–2794 (1998).
Stegmann, T., White, J. M. & Helenius, A. Intermediates in influenza induced membrane fusion. EMBO J. 9, 4231–4241 (1990).
Sattentau, Q. J. & Moore, J. P. Conformational changes induced in the human immunodeficiency virus envelope glycoprotein by soluble CD4 binding. J. Exp. Med. 174, 407–415 (1991).
Thomas, D. et al. Mass and molecular composition of vesicular stomatitis virus: a scanning transmission electron microscopy analysis. J. Virol. 54, 598–607 (1985).
Osakada, F. & Callaway, E. M. Design and generation of recombinant rabies virus vectors. Nat. Protoc. 8, 1583–1601 (2013).
Boulant, S., Stanifer, M. & Lozach, P. Y. Dynamics of virus–receptor interactions in virus binding, signaling, and endocytosis. Viruses 7, 2794–2815 (2015).
Stencel-Baerenwald, J. E., Reiss, K., Reiter, D. M., Stehle, T. & Dermody, T. S. The sweet spot: defining virus–sialic acid interactions. Nat. Rev. Microbiol. 12, 739–749 (2014).
Evans, E. & Williams, P. in Physics of Bio-Molecules and Cells (eds Flyvbjerg, H., Jülicher, F., Orms, P. & David, F.) 145–204 (Springer, 2002).
Barde, I., Salmon, P. & Trono, D. Production and titration of lentiviral vectors. Curr. Protoc. Neurosci. 4, 12.10 (2010).
Freshney, R. I. Animal Cell Culture: A Practical Approach (Oxford Univ. Press, 1992).
Wildling, L. et al. Linking of sensor molecules with amino groups to amino-functionalized AFM tips. Bioconjug. Chem. 22, 1239–1248 (2011).
Hutter, J. L. & Bechhoefer, J. Calibration of atomic-force microscope tips. Rev. Sci. Instrum. 64, 1868–1873 (1993).
Acknowledgements
The authors thank T. Lopez and V. Jäggin for assistance with fluorescence-activated cell sorting operation and analysis, M. Mohr for producing eGFP-encoding lentiviruses and E. Bieler, D. Mathys and S. Erpel for assistance with scanning electron microscopy. The plasmid pAAV-EF1a-FLEX-TVA-mCherry was a gift from N. Uchida. The EnvA expressing BHK cell line was a gift from E. Callaway. The Swiss National Science Foundation (SNF; grant no. 310030B_160225 to D.J.M.), NCCR Molecular Systems Engineering and the European Molecular Biology Organization (EMBO; ALTF 265-2013 to D.A. and ALTF 506-2012 to D.M.M.) supported this work. D.A. is Research Associate of FRS-FNRS.
Author information
Authors and Affiliations
Contributions
D.A., D.J.M., B.R. and R.N. designed the experiments. R.N. produced viruses, modified cell lines and validated virus-binding and transduction. D.A. and R.S. performed confocal microscopy. D.A. and D.M.-M. set up the AFM chamber. D.A. set up and performed AFM experiments and developed strategies to chemically functionalize the AFM tip. M.D., R.N. and R.S. performed scanning electron microscopy. D.A., D.J.M. and R.N. co-analysed the experimental and calculated data. All authors wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
D.A., D.M.M. and D.J.M. have applied for a patent for the chamber enabling AFM and optical microscopy under cell culture conditions (EP15002176.4). The other authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 1716 kb)
Supplementary information
Supplementary Movie (MOV 2871 kb)
Rights and permissions
About this article
Cite this article
Alsteens, D., Newton, R., Schubert, R. et al. Nanomechanical mapping of first binding steps of a virus to animal cells. Nature Nanotech 12, 177–183 (2017). https://doi.org/10.1038/nnano.2016.228
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2016.228
This article is cited by
-
Monitoring the mass, eigenfrequency, and quality factor of mammalian cells
Nature Communications (2024)
-
Single-molecule force stability of the SARS-CoV-2–ACE2 interface in variants-of-concern
Nature Nanotechnology (2024)
-
Paired immunoglobulin-like receptor B is an entry receptor for mammalian orthoreovirus
Nature Communications (2023)
-
Mathematical model for force and energy of virion-cell interactions during full engulfment in HIV: Impact of virion maturation and host cell morphology
Biomechanics and Modeling in Mechanobiology (2023)
-
Multivalent 9-O-Acetylated-sialic acid glycoclusters as potent inhibitors for SARS-CoV-2 infection
Nature Communications (2022)