Abstract
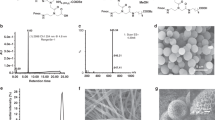
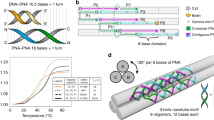
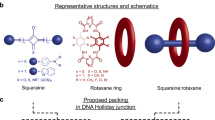
The two main branches of bionanotechnology involve the self-assembly of either peptides or DNA. Peptide scaffolds offer chemical versatility, architectural flexibility and structural complexity, but they lack the precise base pairing and molecular recognition available with nucleic acid assemblies. Here, inspired by the ability of aromatic dipeptides to form ordered nanostructures with unique physical properties, we explore the assembly of peptide nucleic acids (PNAs), which are short DNA mimics that have an amide backbone. All 16 combinations of the very short di-PNA building blocks were synthesized and assayed for their ability to self-associate. Only three guanine-containing di-PNAs—CG, GC and GG—could form ordered assemblies, as observed by electron microscopy, and these di-PNAs efficiently assembled into discrete architectures within a few minutes. The X-ray crystal structure of the GC di-PNA showed the occurrence of both stacking interactions and Watson–Crick base pairing. The assemblies were also found to exhibit optical properties including voltage-dependent electroluminescence and wide-range excitation-dependent fluorescence in the visible region.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ghadiri, M. R., Granja, J. R., Milligan, R. A., McRee, D. E. & Khazanovich, N. Self-assembling organic nanotubes based on a cyclic peptide architecture. Nature 366, 324–327 (1993).
Hartgerink, J. D., Beniash, E. & Stupp, S. I. Self-assembly and mineralization of peptide–amphiphile nanofibers. Science 294, 1684–1688 (2001).
Zhang, S. Fabrication of novel biomaterials through molecular self-assembly. Nature Biotechnol. 21, 1171–1178 (2003).
Banwell, E. F. et al. Rational design and application of responsive alpha-helical peptide hydrogels. Nature Mater. 8, 596–600 (2009).
Morris, K. L. et al. Chemically programmed self-sorting of gelator networks. Nature Commun. 4, 1480 (2013).
Hirst, A. R. et al. Biocatalytic induction of supramolecular order. Nature Chem. 2, 1089–1094 (2010).
De la Rica, R. & Matsui, H. Applications of peptide and protein-based materials in bionanotechnology. Chem. Soc. Rev. 39, 3499–3509 (2010).
Reches, M. & Gazit, E. Casting metal nanowires within discrete self-assembled peptide nanotubes. Science 300, 625–627 (2003).
Yan, X., Zhu, P. & Li, J. Self-assembly and application of diphenylalanine-based nanostructures. Chem. Soc. Rev. 39, 1877–1890 (2010).
Adler-Abramovich, L. et al. Self-assembled arrays of peptide nanotubes by vapour deposition. Nature Nanotech. 4, 849–854 (2009).
Amdursky, N. et al. Blue luminescence based on quantum confinement at peptide nanotubes. Nano Lett. 9, 3111–3115 (2009).
Adler-Abramovich, L. & Gazit, E. The physical properties of supramolecular peptide assemblies: from building block association to technological applications. Chem. Soc. Rev. 43, 6881–6893 (2014).
Seeman, N. C. Nucleic acid junctions and lattices. J. Theor. Biol. 99, 237–247 (1982).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Three-dimensional structures self-assembled from DNA bricks. Science 338, 1177–1183 (2012).
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009).
Egholm, M., Buchardt, O. & Nielsen, P. E. Peptide nucleic acids (PNA). Oligonucleotide analogues with an achiral peptide backbone. J. Am. Chem. Soc. 1, 1895–1897 (1992).
Nielsen, P. E., Egholm, M., Berg, R. H. & Buchardt, O. Sequence-selective recognition of DNA by strand displacement with a thymine-substituted polyamide. Science 254, 1497–1500 (1991).
Ura, Y., Beierle, J. M., Leman, L. J., Orgel, L. E. & Ghadiri, M. R. Self-assembling sequence-adaptive peptide nucleic acids. Science 325, 73–77 (2009).
Li, X. et al. Supramolecular nanofibers and hydrogels of nucleopeptides. Angew. Chem. Int. Ed. 50, 9365–9369 (2011).
Bonifazi, D., Carloni, L. E., Corvaglia, V. & Delforge, A. Peptide nucleic acids in materials science. Artif. DNA PNA XNA 3, 112–122 (2012).
Guler, M. O., Pokorski, J. K., Appella, D. H. & Stupp, S. I. Enhanced oligonucleotide binding to self-assembled nanofibers. Bioconjug. Chem. 16, 501–503 (2005).
Böhler, C., Nielsen, P. E. & Orgel, L. E. Template switching between PNA and RNA oligonucleotides. Nature 376, 578–581 (1995).
Uhlmann, E., Peyman, A., Breipohl, G. & Will, D. W. PNA: synthetic polyamide nucleic acids with unusual binding properties. Angew. Chem. Int. Ed. 37, 2796–2823 (1998).
Davis, J. T. & Spada, G. P. Supramolecular architectures generated by self-assembly of guanosine derivatives. Chem. Soc. Rev. 36, 296–313 (2007).
Menchise, V. et al. Insights into peptide nucleic acid (PNA) structural features: the crystal structure of a D-lysine-based chiral PNA–DNA duplex. Proc. Natl Acad. Sci. USA 100, 12021–12026 (2003).
Watson, J. D. & Crick, F. H. C. Molecular structure of nucleic acids: a structure for deoxyribose nucleic acid. Nature 171, 737–738 (1953).
Trushko, A., Schäffer, E. & Howard, J. The growth speed of microtubules with XMAP215-coated beads coupled to their ends is increased by tensile force. Proc. Natl Acad. Sci. USA 110, 14670–14675 (2013).
Ikezoe, Y., Washino, G., Uemura, T., Kitagawa, S. & Matsui, H. Autonomous motors of a metal–organic framework powered by reorganization of self-assembled peptides at interfaces. Nature Mater. 11, 1081–1085 (2012).
Demchenko, A. P. The red-edge effect: 30 years of exploration. Luminesence 17, 19–42 (2002).
Cushing, S. K., Li, M., Huang, F. & Wu, N. Origin of strong excitation wavelength dependent fluoresence of graphene oxide. ACS Nano 8, 1002–1013 (2014).
Muccini, M. A bright future for organic field-effect transistors. Nature Mater. 5, 605–613 (2006).
Incardona, M. F. et al. EDNA: a framework for plugin-based applications applied to X-ray experiment online data analysis. J. Synchrotron Radiat. 16, 872–879 (2009).
Spek, A. L. Structure validation in chemical crystallography. Acta Crystallogr. D 65, 148–155 (2009).
Acknowledgements
This work was supported in part by grants from the Israeli National Nanotechnology Initiative and Helmsley Charitable Trust for a focal technology area on Nanomedicine for Personalized Theranostics. The authors thank members of the Gazit Laboratory for helpful discussions, Y. Salitra for help with PNA synthesis and O. Yaniv for advice on crystallization experiments. The authors acknowledge the ESRF for synchrotron beam time and the staff scientists of the ID29 beamline for their assistance. O. Berger is supported by a fellowship from the Argentinean Friends of Tel Aviv University Association.
Author information
Authors and Affiliations
Contributions
O.B., L.A-A., L.B., Y.E. and E.G. designed the study. O.B., M.L-S., Y.L-P. and M.B. performed the experiments. O.B., A.G., T.S. and Y.E. analysed the data. F.F., L.J.W.S., E.M. and T.F. performed and analysed the X-ray diffraction experiments. F.P. performed OFET experiments. O.B., L.A-A. and E.G. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 617 kb)
Supplementary information
Supplementary Movie 1 (MP4 46352 kb)
Supplementary information
Supplementary Movie 2 (MP4 6015 kb)
Supplementary information
Supplementary information (CIF 24 kb)
Rights and permissions
About this article
Cite this article
Berger, O., Adler-Abramovich, L., Levy-Sakin, M. et al. Light-emitting self-assembled peptide nucleic acids exhibit both stacking interactions and Watson–Crick base pairing. Nature Nanotech 10, 353–360 (2015). https://doi.org/10.1038/nnano.2015.27
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2015.27
This article is cited by
-
Bio-organic adaptive photonic crystals enable supramolecular solvatochromism
Nano Research (2023)
-
Modulating vectored non-covalent interactions for layered assembly with engineerable properties
Bio-Design and Manufacturing (2022)
-
Modular self-assembly of gamma-modified peptide nucleic acids in organic solvent mixtures
Nature Communications (2020)
-
Proton-enabled activation of peptide materials for biological bimodal memory
Nature Communications (2020)
-
Hierarchically oriented organization in supramolecular peptide crystals
Nature Reviews Chemistry (2019)