Abstract
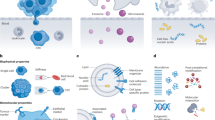
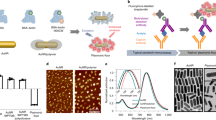
Rapid progress in identifying disease biomarkers has increased the importance of creating high-performance detection technologies. Over the last decade, the design of many detection platforms has focused on either the nano or micro length scale. Here, we review recent strategies that combine nano- and microscale materials and devices to produce large improvements in detection sensitivity, speed and accuracy, allowing previously undetectable biomarkers to be identified in clinical samples. Microsensors that incorporate nanoscale features can now rapidly detect disease-related nucleic acids expressed in patient samples. New microdevices that separate large clinical samples into nanocompartments allow precise quantitation of analytes, and microfluidic systems that utilize nanoscale binding events can detect rare cancer cells in the bloodstream more accurately than before. These advances will lead to faster and more reliable clinical diagnostic devices.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Virgin, H. W. & Todd, J. A. Metagenomics and personalized medicine. Cell 147, 44–56 (2011).
Hood, L., Heath, J. R., Phelps, M. E. & Lin, B. Systems biology and new technologies enable predictive and preventative medicine. Science 306, 640–643 (2004).
Walt, D. R. Miniature analytical methods for medical diagnostics. Science 308, 217–219 (2005).
Niemz, A., Ferguson, T. M. & Boyle, D. S. Point-of-care nucleic acid testing for infectious diseases. Trends Biotechnol. 29, 240–250 (2011).
Bradley, C. J., Given, C. W. & Roberts, C. Disparities in cancer diagnosis and survival. Cancer 91, 178–188 (2001).
Urdea, M. et al. Requirements for high impact diagnostics in the developing world. Nature 444 (suppl. 1), 73–79 (2006).
Pai, N. P. et al. Point-of-care testing for infectious diseases: diversity, complexity, and barriers in low- and middle-income countries. PLoS Med. 9, e1001306 (2012).
Rosi, N. L. & Mirkin, C. A. Nanostructures in biodiagnostics. Chem. Rev. 105, 1547–1562 (2005).
Burda, C., Chen, X., Narayanan, R. & El-Sayed, M. A. Chemistry and properties of nanocrystals of different shapes. Chem. Rev. 105, 1025–1102 (2005).
Sapsford, K. E. et al. Functionalizing nanoparticles with biological molecules: developing chemistries that facilitate nanotechnology. Chem. Rev. 113, 1904–2074 (2013).
Thorsen, T., Maerkl, S. J. & Quake, S. R. Microfluidic large-scale integration. Science 298, 580–584 (2002).
Sheehan, P. E. & Whitman, L. J. Detection limits for nanoscale biosensors. Nano Lett. 5, 803–807 (2005). This paper was one of the first to use simulations to highlight the importance of multi-length-scale engineering.
Squires, T. M., Messinger, R. J. & Manalis, S. R. Making it stick: convection, reaction and diffusion in surface-based biosensors. Nature Biotechnol. 26, 417–426 (2008).
Gosling, J. P. A decade of development in immunoassay methodology. Clin. Chem. 36, 1408–1427 (1990).
Emmadi, R. et al. Molecular methods and platforms for infectious diseases testing a review of FDA-approved and cleared assays. J. Mol. Diagn. 13, 583–604 (2011).
Nam, J. M., Thaxton, C. S. & Mirkin, C. A. Nanoparticle-based bio-bar codes for the ultrasensitive detection of proteins. Science 301, 1884–1886 (2003). Record-breaking sensitivity was reported in this paper for the detection of protein analytes.
Cutler, J. I., Auyeung, E. & Mirkin, C. A. Spherical nucleic acids. J. Am. Chem. Soc. 134, 1376–1391 (2012).
Kim, E. Y. et al. Detection of HIV-1 p24 Gag in plasma by a nanoparticle-based bio-barcode-amplification method. Nanomedicine 3, 293–303 (2008).
Thaxton, C. S. et al. Nanoparticle-based bio-barcode assay redefines “undetectable” PSA and biochemical recurrence after radical prostatectomy. Proc. Natl Acad. Sci. USA 106, 18437–18442 (2009).
Georganopoulou, D. G. et al. Nanoparticle-based detection in cerebral spinal fluid of a soluble pathogenic biomarker for Alzheimer's disease. Proc. Natl Acad. Sci. USA 102, 2273–2276 (2005).
Cattani-Scholz, A. et al. Organophosphonate-based PNA-functionalization of silicon nanowires for label-free DNA detection. ACS Nano 2, 1653–1660 (2008).
Bunimovich, Y. L. et al. Quantitative real-time measurements of DNA hybridization with alkylated nonoxidized silicon nanowires in electrolyte solution. J. Am. Chem. Soc. 128, 16323–16331 (2006).
Hahm, J. & Lieber, C. M. Direct ultrasensitive electrical detection of DNA and DNA sequence variations using nanowire nanosensors. Nano Lett. 4, 51–54 (2004).
Zhang, J. et al. Sequence-specific detection of femtomolar DNA via a chronocoulometric DNA sensor (CDS): Effects of nanoparticle-mediated amplification and nanoscale control of DNA assembly at electrodes. J. Am. Chem. Soc. 128, 8575–8580 (2006).
Wang, J., Liu, G. & Merkoçi, A. Electrochemical coding technology for simultaneous detection of multiple DNA targets. J. Am. Chem. Soc. 125, 3214–3215 (2003).
Liao, J. C. et al. Use of electrochemical DNA biosensors for rapid molecular identification of uropathogens in clinical urine specimens. J. Clin. Microbiol. 44, 561–570 (2006).
Xiang, Y. & Lu, Y. Using personal glucose meters and functional DNA sensors to quantify a variety of analytical targets. Nature Chem. 3, 697–703 (2011).
Fan, C., Plaxco, K. W. & Heeger, A. J. Electrochemical interrogation of conformational changes as a reagentless method for the sequence-specific detection of DNA. Proc. Natl Acad. Sci. USA 100, 9134–9137 (2003).
Li, D., Song, S. & Fan, C. Target-responsive structural switching for nucleic acid-based sensors. Acc. Chem. Res. 43, 631–641 (2010).
Park, S. J., Taton, T. A. & Mirkin, C. A. Array-based electrical detection of DNA with nanoparticle probes. Science 295, 1503–1506 (2002).
Drummond, T. G., Hill, M. G. & Barton, J. K. Electrochemical DNA sensors. Nature Biotechnol. 21, 1192–1199 (2003).
Lapierre, M. A., O'Keefe, M., Taft, B. J. & Kelley, S. O. Electrocatalytic detection of pathogenic DNA sequences and antibiotic resistance markers. Anal. Chem. 75, 6327–6333 (2003).
Soleymani, L. et al. Hierarchical nanotextured microelectrodes overcome the molecular transport barrier to achieve rapid, direct bacterial detection. ACS Nano 5, 3360–3366 (2011).
Soleymani, L., Fang, Z., Sargent, E. H. & Kelley, S. O. Programming the detection limits of biosensors through controlled nanostructuring. Nature Nanotech. 4, 844–848 (2009).
Bin, X., Sargent, E. H. & Kelley, S. O. Nanostructuring of sensors determines the efficiency of biomolecular capture. Anal. Chem. 82, 5928–5931 (2010).
Lam, B., Fang, Z., Sargent, E. H. & Kelley, S. O. Polymerase chain reaction-free, sample-to-answer bacterial detection in 30 minutes with integrated cell lysis. Anal. Chem. 84, 21–25 (2012).
Hill, H. D., Millstone, J. E., Banholzer, M. J. & Mirkin, C. A. The role radius of curvature plays in thiolated oligonucleotide loading on gold nanoparticles. ACS Nano 3, 418–424 (2009). This paper was the first to demonstrate that analytical sensitivity could be modulated by nanoscale surface morphology coupled with microscale sensors.
Lam, B. et al. Solution-based circuits enable rapid and multiplexed pathogen detection. Nature Commun. 4, 2001 (2013).
Creamer, E. et al. The effect of rapid screening for methicillin-resistant Staphylococcus aureus (MRSA) on the identification and earlier isolation of MRSA-positive patients. Infect. Control Hosp. Epidemiol. 31, 374–381 (2010).
Das, J. et al. An ultrasensitive universal detector based on neutralizer displacement. Nature Chem. 4, 642–648 (2012).
Fang, Z. et al. Direct profiling of cancer biomarkers in tumor tissue using a multiplexed nanostructured microelectrode integrated circuit. ACS Nano 3, 3207–3213 (2009).
Vasilyeva, E. et al. Direct genetic analysis of ten cancer cells: tuning sensor structure and molecular probe design for efficient mRNA capture. Angew. Chem. Int. Ed. 50, 4137–4141 (2011).
Yang, H. et al. Direct, electronic microRNA detection for the rapid determination of differential expression profiles. Angew. Chem. Int. Ed. 48, 8461–8464 (2009).
Ivanov, I. et al. Chip-based nanostructured sensors enable accurate identification and classification of circulating tumor cells in prostate cancer patient blood samples. Anal. Chem. 85, 398–403 (2013).
Pei, H. et al. A DNA nanostructure-based biomolecular probe carrier platform for electrochemical biosensing. Adv. Mater. 22, 4754–4758 (2010).
Lu, N. et al. Charge transport within a three-dimensional DNA nanostructure framework. J. Am. Chem. Soc. 134, 13148–13151 (2012).
Ge, Z. et al. Hybridization chain reaction amplification of microRNA detection with a tetrahedral DNA nanostructure-based electrochemical biosensor. Anal. Chem. 86, 2124–2130 (2014).
Wen, Y. et al. DNA nanostructure-based interfacial engineering for PCR-free ultrasensitive electrochemical analysis of microRNA. Sci. Rep. 2, 867 (2012).
Pei, H. et al. Regenerable electrochemical immunological sensing at DNA nanostructure-decorated gold surfaces. Chem. Commun. 47, 6254–6256 (2011).
Wen, Y. et al. DNA nanostructure-decorated surfaces for enhanced aptamer-target binding and electrochemical cocaine sensors. Anal. Chem. 83, 7418–7423 (2011).
Sykes, P. J. et al. Quantitation of targets for PCR by use of limiting dilution. Biotechniques 13, 444–449 (1992).
Vogelstein, B. & Kinzler, K. W. Digital PCR. Proc. Natl Acad. Sci. USA 96, 9236–9241 (1999).
Baker, M. Digital PCR hits its stride. Nature Methods 9, 541–544 (2012).
Heyries, K. A. et al. Megapixel digital PCR. Nature Methods 8, 649–651 (2011).
Qin, J., Jones, R. C. & Ramakrishnan, R. Studying copy number variations using a nanofluidic platform. Nucl. Acids Res. 36, e116 (2008).
White, R. A., Quake, S. R. & Curr, K. Digital PCR provides absolute quantitation of viral load for an occult RNA virus. J. Virol. Methods 179, 45–50 (2012).
Strain, M. C. et al. Highly precise measurement of HIV DNA by droplet digital PCR. PLoS ONE 8, e55943 (2013).
Warren, L., Bryder, D., Weissman, I. L. & Quake, S. R. Transcription factor profiling in individual hematopoietic progenitors by digital RT-PCR. Proc. Natl Acad. Sci. USA 103, 17807–17812 (2006).
Goh, H. G. et al. Sensitive quantitation of minimal residual disease in chronic myeloid leukemia using nanofluidic digital polymerase chain reaction assay. Leuk. Lymphoma 52, 896–904 (2011).
Fan, H. C. et al. Microfluidic digital PCR enables rapid prenatal diagnosis of fetal aneuploidy. Am. J. Obs. Gynecol. 200, 543.e541–543.e547 (2009).
Lo, Y. M. D. et al. Digital PCR for the molecular detection of fetal chromosomal aneuploidy. Proc. Natl Acad. Sci. USA 104, 13116–13121 (2007).
Fan, H. C. et al. Non-invasive prenatal measurement of the fetal genome. Nature 487, 320–324 (2012).
Gansen, A. et al. Digital LAMP in a sample self-digitization (SD) chip. Lab Chip 12, 2247–2254 (2012).
Shen, F. et al. Multiplexed quantification of nucleic acids with large dynamic range using multivolume digital RT-PCR on a rotational SlipChip tested with HIV and hepatitis C viral load. J. Am. Chem. Soc. 133, 17705–17712 (2011). This paper highlights the large dynamic ranges that can be achieved with quantitative digital methods.
Sun, B. et al. Mechanistic evaluation of the pros and cons of digital RT-LAMP for HIV-1 viral load quantification on a microfluidic device and improved efficiency via a two-step digital protocol. Anal. Chem. 85, 1540–1546 (2013).
Byrnes, S. et al. A portable, pressure driven, room temperature nucleic acid extraction and storage system for point of care molecular diagnostics. Anal. Methods 5, 3177–3184 (2013).
Selck, D. A., Karymov, M. A., Sun, B. & Ismagilov, R. F. Increased robustness of single-molecule counting with microfluidics, digital isothermal amplification, and a mobile phone versus real-time kinetic measurements. Anal. Chem. 85, 11129–11136 (2013).
Rissin, D. M. et al. Single-molecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations. Nature Biotechnol. 28, 595–599 (2010).
Rissin, D. M. & Walt, D. R. Digital concentration readout of single enzyme molecules using femtoliter arrays and Poisson statistics. Nano Lett. 6, 520–523 (2006).
Rissin, D. M. & Walt, D. R. Digital readout of target binding with attomole detection limits via enzyme amplification in femtoliter arrays. J. Am. Chem. Soc. 128, 6286–6287 (2006). The first example of a digital quantitation approach applied to protein analytes.
Li, Z., Hayman, R. B. & Walt, D. R. Detection of single-molecule DNA hybridization using enzymatic amplification in an array of femtoliter-sized reaction vessels. J. Am. Chem. Soc. 130, 12622–12623 (2008).
Song, L. et al. Direct detection of bacterial genomic DNA at sub-femtomolar concentrations using single molecule arrays. Anal. Chem. 85, 1932–1939 (2013).
Zetterberg, H. et al. Plasma tau levels in Alzheimer's disease. Alzheimers Res. Ther. 5, 9 (2013).
Neselius, S. et al. Olympic boxing is associated with elevated levels of the neuronal protein tau in plasma. Brain Inj. 27, 425–433 (2013).
Randall, J. et al. Tau proteins in serum predict neurological outcome after hypoxic brain injury from cardiac arrest: results of a pilot study. Resuscitation 84, 351–356 (2013).
Zetterberg, H. et al. Hypoxia due to cardiac arrest induces a time-dependent increase in serum amyloid beta levels in humans. PLoS ONE 6, e28263 (2011).
Duffy, D. C. Ultra-sensitive protein detection using single molecule arrays (Simoa): the potential for detecting single molecules of botulinum toxin. The Botulinum 2, 164–167 (2012).
Chang, L. et al. Simple diffusion-constrained immunoassay for p24 protein with the sensitivity of nucleic acid amplification for detecting acute HIV infection. J. Virol. Methods 188, 153–160 (2013).
Lepor, H. et al. Clinical evaluation of a novel method for the measurement of prostate-specific antigen, AccuPSA(TM), as a predictor of 5-year biochemical recurrence-free survival after radical prostatectomy: results of a pilot study. BJU Int. 109, 1770–1775 (2012).
Song, L. et al. Single molecule measurements of tumor necrosis factor α and interleukin-6 in the plasma of patients with Crohn's disease. J. Immunol. Methods 372, 177–186 (2011).
Miltenyi, S., Muller, W., Weichel, W. & Radbruch, A. High gradient magnetic cell separation with MACS. Cytometry 11, 231–238 (1990).
Allard, W. J. et al. Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases. Clin. Cancer Res. 10, 6897–6904 (2004).
Nagrath, S. et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450, 1235–1239 (2007).
Stott, S. L. et al. Isolation and characterization of circulating tumor cells from patients with localized and metastatic prostate cancer. Sci. Transl. Med. 2, 25ra23 (2010). This study reported significant progress in high-sensitivity CTC enumeration.
Miyamoto, D. T. et al. Androgen receptor signaling in circulating tumor cells as a marker of hormonally responsive prostate cancer. Cancer Disc. 2, 995–1003 (2012).
Yu, M. et al. RNA sequencing of pancreatic circulating tumour cells implicates WNT signalling in metastasis. Nature 487, 510–513 (2012).
Kuo, J. S. et al. Deformability considerations in filtration of biological cells. Lab Chip 10, 837–842 (2010).
Hosokawa, M. et al. Size-selective microcavity array for rapid and efficient detection of circulating tumor cells. Anal. Chem. 82, 6629–6635 (2010).
Lee, H. J. et al. Efficient isolation and accurate in situ analysis of circulating tumor cells using detachable beads and a high-pore-density filter. Angew. Chem. Int. Ed. 52, 8337–8340 (2013).
Baccelli, I. et al. Identification of a population of blood circulating tumor cells from breast cancer patients that initiates metastasis in a xenograft assay. Nature Biotechnol. 31, 539–544 (2013).
Zhang, L. et al. The identification and characterization of breast cancer CTCs competent for brain metastasis. Sci. Transl. Med. 5, 180ra148 (2013).
Yu, M. et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339, 580–584 (2013). This study highlights the importance of capturing CTCs with diverse phenotypes.
Kirby, B. J. et al. Functional characterization of circulating tumor cells with a prostate-cancer-specific microfluidic device. PLoS ONE 7, e35976 (2012).
Casavant, B. P. et al. A negative selection methodology using a microfluidic platform for the isolation and enumeration of circulating tumor cells. Methods 64, 137–143 (2013).
Ozkumur, E. et al. Inertial focusing for tumor antigen-dependent and -independent sorting of rare circulating tumor cells. Sci Transl. Med. 5, 179ra147 (2013).
Issadore, D. et al. Ultrasensitive clinical enumeration of rare cells ex vivo using a micro-hall detector. Sci. Transl. Med. 4, 141ra92 (2012).
Zhao, M. et al. An automated high-throughput counting method for screening circulating tumor cells in peripheral blood. Anal. Chem. 85, 2465–2471 (2013).
Prigodich, A. E. et al. Multiplexed nanoflares: mRNA detection in live cells. Anal. Chem. 84, 2062–2066 (2012).
Acknowledgements
S.O.K. and E.H.S. acknowledge Genome Canada, the Canadian Institute of Health Research, the Natural Sciences and Engineering Research Council, and the Ontario Research Fund for support of their work. M.T. acknowledges the National Institute of Health (NIH) P41 Resource Center, NIH National Institute of Biomedical Imaging and Bioengineering Quantum Grant. R.F.I acknowledges NIH grant R01EB012946 and the Defense Advanced Research Projects Agency (DARPA) Cooperative Agreement HR0011-11-2-0006 for support. D.R.W. acknowledges generous support from DARPA (HR0011-12-2,0001: SUB #5-55065) and a Department of Defense Innovator Award BC100510(W81XWH-11-1-0814). C.A.M. acknowledges support from the Center for Cancer Nanotechnology Excellence (CCNE) initiative of the National Institutes of Health (NIH), the Nanoscale Science and Engineering Centers (NSEC) initiative of the National Science Foundation, the Prostate Cancer Foundation, National Institute of Arthritis and Musculosketal and Skin Diseases/NIH, and DARPA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
R.F.I. is a scientific founder, a Director, and has equity in SlipChip. D.R.W. is a scientific founder, a Director, and has equity in Quanterix. S.O.K. is a founder, a Director, and has equity in Xagenic. E.H.S. holds equity in Xagenic.
Rights and permissions
About this article
Cite this article
Kelley, S., Mirkin, C., Walt, D. et al. Advancing the speed, sensitivity and accuracy of biomolecular detection using multi-length-scale engineering. Nature Nanotech 9, 969–980 (2014). https://doi.org/10.1038/nnano.2014.261
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2014.261
This article is cited by
-
Biomolecular sensors for advanced physiological monitoring
Nature Reviews Bioengineering (2023)
-
Direct digital sensing of protein biomarkers in solution
Nature Communications (2023)
-
Multifunctional Perovskite Photodetectors: From Molecular-Scale Crystal Structure Design to Micro/Nano-scale Morphology Manipulation
Nano-Micro Letters (2023)
-
Lossless enrichment of trace analytes in levitating droplets for multiphase and multiplex detection
Nature Communications (2022)
-
Nano-omics: nanotechnology-based multidimensional harvesting of the blood-circulating cancerome
Nature Reviews Clinical Oncology (2022)