Abstract
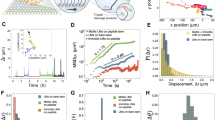
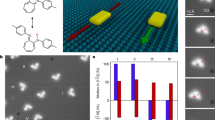
Controlled motion at the nanoscale can be achieved by using Watson–Crick base-pairing to direct the assembly and operation of a molecular transport system consisting of a track, a motor1,2,3,4,5,6,7,8,9,10,11,12 and fuel13,14,15, all made from DNA. Here, we assemble a 100-nm-long DNA track on a two-dimensional scaffold16, and show that a DNA motor loaded at one end of the track moves autonomously and at a constant average speed along the full length of the track, a journey comprising 16 consecutive steps for the motor. Real-time atomic force microscopy allows direct observation of individual steps of a single motor, revealing mechanistic details of its operation. This precisely controlled, long-range transport could lead to the development of systems that could be programmed and routed by instructions encoded in the nucleotide sequences of the track and motor. Such systems might be used to create molecular assembly lines modelled on the ribosome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Shin, J-S. & Pierce, N. A. A synthetic DNA walker for molecular transport. J. Am. Chem. Soc. 126, 10834–10835 (2004).
Sherman, W. B. & Seeman, N. C. A precisely controlled DNA biped walking device. Nano Lett. 4, 1203–1207 (2004).
Tian, Y. & Mao, C. A pair of DNA circles continuously rolls against each other. J. Am. Chem. Soc. 126, 11410–11411 (2004).
Yin, P., Yan, H., Daniell, X. G., Turberfield, A. J. & Reif, J. H. A unidirectional DNA walker that moves autonomously along a DNA track. Angew. Chem. Int. Ed. 43, 4906–4911 (2004).
Tian, Y., He, Y., Peng, Y. & Mao, C. A DNA enzyme that walks processively and autonomously along a one-dimensional track. Angew. Chem. Int. Ed. 44, 4355–4358 (2005).
Bath, J., Green, S. J. & Turberfield, A. J. A free-running DNA motor powered by a nicking enzyme. Angew. Chem. Int. Ed. 44, 4358–4361 (2005).
Pei, R. et al. Behaviour of polycatalytic assemblies in a substrate-displaying matrix. J. Am. Chem. Soc. 128, 12693–12699 (2006).
Venkataraman, S., Dirks, R. M., Rothemund, P. W. K., Winfree, E. & Pierce, N. A. An autonomous polymerization motor powered by DNA hybridization. Nature Nanotech. 2, 490–494 (2007).
Yin, P., Choi, H. M., Calvert, C. R. & Pierce, N. A. Programming biomolecular self-assembly pathways. Nature 451, 318–322 (2008).
Green, S. J., Bath, J. & Turberfield, A. J. Coordinated chemomechanical cycles: a mechanism for autonomous molecular motion. Phys. Rev. Lett. 101, 238101 (2008).
Bath, J. Green, S. J., Allen, K. E. & Turberfield, A. J. Mechanism for a directional, processive and reversible DNA motor. Small 5, 1513–1516 (2009).
Omabegho, T., Sha, R. & Seeman, N. C. A bipedal Brownian motor with coordinated legs. Science 324, 67–71 (2009).
Turberfield, A. J. et al. DNA fuel for free-running nanomachines. Phys. Rev. Lett. 90, 118102 (2003).
Bois, J. S. et al. Topological constraints in nucleic acid hybridization kinetics. Nucleic Acids Res. 33, 4090–4095 (2005).
Green, S. J., Lubrich, D. & Turberfield, A. J. DNA hairpins: fuel for autonomous DNA devices. Biophys. J. 91, 2966–2975 (2006).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Antal, T. & Krapivsky, P. L. Molecular spiders with memory. Phys. Rev. E 76, 021121 (2007).
Lund, K. et al. Molecular robots guided by prescriptive landscapes. Nature 465, 206–210 (2010).
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009).
Heiter, D., Lunnen, K. D. & Wilson, G. G. Site-specific DNA-nicking mutants of the heterodimeric restriction endonuclease R.BbvCI. J. Mol. Biol. 348, 631–640 (2005).
Yurke, B., Turberfield, A. J., Mills, A. P. Jr, Simmel, F. C. & Neumann, J. L. A DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Yurke, B. & Mills, A. P. Jr. Using DNA to power nanostructures. Genet. Program. Evol. Mach. 4, 111–122 (2003).
Ando, T. et al. High-speed atomic force microscope for studying biological macromolecules. Proc. Natl Acad. Sci. USA 98, 12468–12472 (2001).
Endo, M., Katsuda, Y., Hidaka, K. & Sugiyama, H. Regulation of DNA methylation using different tensions in the double strands constructed in a defined DNA nanostructure. J. Am. Chem. Soc. 132, 1592–1597 (2010).
Endo, M., Sugita, T., Katsuda, Y., Hidaka, K. & Sugiyama, H. Programmed-assembly system using DNA jigsaw pieces. Chem. Eur. J. 16, 5362–5368 (2010).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Gu, H., Chao, J., Xiao, S. J. & Seeman, N. C. A proximity-based programmable DNA nanoscale assembly line. Nature 465, 202–205 (2010).
Acknowledgements
This work was supported by the Engineering and Physical Sciences Research Council (EP/G037930/1), the Clarendon Fund, the Oxford–Australia Scholarship Fund, the CREST of JST and a Grant-in-Aid for Science Research from the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Author information
Authors and Affiliations
Contributions
Experiments were designed by S.W. with input from J.B. and A.J.T. Ensemble fluorescence experiments were carried out by S.W in the laboratory of A.J.T. Real-time AFM experiments were done by S.W., M.E., Y.K. and K.H in the laboratory of H.S. The manuscript was written by S.W., J.B., H.S. and A.J.T.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 4317 kb)
Supplementary information
Supplementary movie 1 (MOV 4547 kb)
Supplementary information
Supplementary movie 2 (MOV 388 kb)
Rights and permissions
About this article
Cite this article
Wickham, S., Endo, M., Katsuda, Y. et al. Direct observation of stepwise movement of a synthetic molecular transporter. Nature Nanotech 6, 166–169 (2011). https://doi.org/10.1038/nnano.2010.284
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2010.284
This article is cited by
-
Enzyme-driven Nanorobots Walking Along Predesigned Tracks on the DNA Origami for Cargo Transport and Catalysis
Chemical Research in Chinese Universities (2024)
-
Adsorbate motors for unidirectional translation and transport
Nature (2023)
-
Dissipative DNA nanotechnology
Nature Chemistry (2022)
-
Moving forward in the semantic soup of artificial molecular machine taxonomy
Nature Nanotechnology (2022)
-
Toggling Between Two Limit Cycles in a Molecular Ecosystem
New Generation Computing (2022)