Abstract
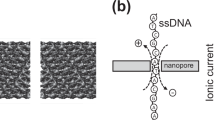
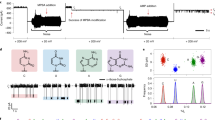
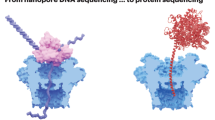
A single-molecule method for sequencing DNA that does not require fluorescent labelling could reduce costs and increase sequencing speeds. An exonuclease enzyme might be used to cleave individual nucleotide molecules from the DNA, and when coupled to an appropriate detection system, these nucleotides could be identified in the correct order. Here, we show that a protein nanopore with a covalently attached adapter molecule can continuously identify unlabelled nucleoside 5'-monophosphate molecules with accuracies averaging 99.8%. Methylated cytosine can also be distinguished from the four standard DNA bases: guanine, adenine, thymine and cytosine. The operating conditions are compatible with the exonuclease, and the kinetic data show that the nucleotides have a high probability of translocation through the nanopore and, therefore, of not being registered twice. This highly accurate tool is suitable for integration into a system for sequencing nucleic acids and for analysing epigenetic modifications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bayley, H. Sequencing single molecules of DNA. Curr. Opin. Chem. Biol. 10, 628–637 (2006).
Branton, D. et al. The potential and challenges of nanopore sequencing. Nature Biotechnol. 26, 1146–1153 (2008).
Kasianowicz, J. J., Brandin, E., Branton, D. & Deamer, D. W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl Acad. Sci. USA 93, 13770–13773 (1996).
Deamer, D. W. & Branton, D. Characterization of nucleic acids by nanopore analysis. Acc. Chem. Res. 35, 817–825 (2002).
Ashkenasy, N., Sanchez-Quesada, J., Bayley, H. & Ghadiri, M. R. Recognizing a single base in an individual DNA strand: a step toward DNA sequencing in nanopores. Angew Chem. Int. Ed. Engl. 44, 1401–1404 (2005).
Mitchell, N. & Howorka, S. Chemical tags facilitate the sensing of individual DNA strands with nanopores. Angew Chem. Int. Ed. Engl. 47, 5565–5568 (2008).
Astier, Y., Kainov, D. E., Bayley, H., Tuma, R. & Howorka, S. Stochastic detection of motor protein–RNA complexes by single-channel current recording. Chem. Phys. Chem. 8, 2189–2194 (2007).
Hornblower, B. et al. Single-molecule analysis of DNA–protein complexes using nanopores. Nature Methods 4, 315–317 (2007).
Cockroft, S. L., Chu, J., Amorin, M. & Ghadiri, M. R. A single-molecule nanopore device detects DNA polymerase activity with single-nucleotide resolution. J. Am. Chem. Soc. 130, 818–820 (2008).
Maglia, G., Restrepo, M. R., Mikhailova, E. & Bayley, H. Enhanced translocation of single DNA molecules through alpha-hemolysin nanopores by manipulation of internal charge. Proc. Natl Acad. Sci. USA 105, 19720–19725 (2008).
Jett, J. H. et al. High-speed DNA sequencing: an approach based upon fluorescence detection of single molecules. J. Biomol. Struct. Dyn. 7, 301–309 (1989).
Sauer, M. et al. Single molecule DNA sequencing in submicrometer channels: state of the art and future prospects. J. Biotechnol. 86, 181–201 (2001).
Stephan, J. et al. Towards a general procedure for sequencing single DNA molecules. J. Biotechnol. 86, 255–267 (2001).
Werner, J. H. et al. Progress towards single-molecule DNA sequencing: a one color demonstration. J. Biotechnol. 102, 1–14 (2003).
Werner, J. H., Cai, H., Keller, R. A. & Goodwin, P. M. Exonuclease I hydrolyzes DNA with a distribution of rates. Biophys. J. 88, 1403–1412 (2005).
Bayley, H. & Cremer, P. S. Stochastic sensors inspired by biology. Nature 413, 226–230 (2001).
Cheley, S., Gu, L. Q. & Bayley, H. Stochastic sensing of nanomolar inositol 1,4,5-trisphosphate with an engineered pore. Chem. Biol. 9, 829–838 (2002).
Shin, S. H., Luchian, T., Cheley, S., Braha, O. & Bayley, H. Kinetics of a reversible covalent-bond-forming reaction observed at the single-molecule level. Angew Chem. Int. Ed. Engl. 41, 3707–3709, 3523 (2002).
Gu, L. Q., Braha, O., Conlan, S., Cheley, S. & Bayley, H. Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter. Nature 398, 686–690 (1999).
Astier, Y., Braha, O. & Bayley, H. Toward single molecule DNA sequencing: direct identification of ribonucleoside and deoxyribonucleoside 5′-monophosphates by using an engineered protein nanopore equipped with a molecular adapter. J. Am. Chem. Soc. 128, 1705–1710 (2006).
Brody, R. S., Doherty, K. G. & Zimmerman, P. D. Processivity and kinetics of the reaction of exonuclease I from Escherichia coli with polydeoxyribonucleotides. J. Biol. Chem. 261, 7136–7143 (1986).
Enderlein, J. Nucleotide specificity versus complex heterogeneity in exonuclease activity measurements. Biophys. J. 92, 1556–1558 (2007).
Cheley, S., Braha, O., Lu, X., Conlan, S. & Bayley, H. A functional protein pore with a ‘retro’ transmembrane domain. Protein Sci. 8, 1257–1267 (1999).
Song, L. et al. Structure of staphylococcal alpha-hemolysin, a heptameric transmembrane pore. Science 274, 1859–1866 (1996).
Gu, L. Q., Cheley, S. & Bayley, H. Capture of a single molecule in a nanocavity. Science 291, 636–640 (2001).
Wu, H. C., Astier, Y., Maglia, G., Mikhailova, E. & Bayley, H. Protein nanopores with covalently attached molecular adapters. J. Am. Chem. Soc. 129, 16142–16148 (2007).
Sanchez-Quesada, J., Ghadiri, M. R., Bayley, H. & Braha, O. Cyclic peptides as molecular adapters for a pore-forming protein. J. Am. Chem. Soc. 122, 11757–11766 (2000).
Brena, R. M., Huang, T. H. & Plass, C. Toward a human epigenome. Nature Genet. 38, 1359–1360 (2006).
Schaefer, C. B., Ooi, S. K., Bestor, T. H. & Bourc'his, D. Epigenetic decisions in mammalian germ cells. Science 316, 398–399 (2007).
Ooi, S. K. & Bestor, T. H. The colorful history of active DNA demethylation. Cell 133, 1145–1148 (2008).
Hayatsu, H. The bisulfite genomic sequencing used in the analysis of epigenetic states, a technique in the emerging environmental genotoxicology research. Mutat. Res. 659, 77–82 (2008).
Howorka, S., Cheley, S. & Bayley, H. Sequence-specific detection of individual DNA strands using engineered nanopores. Nature Biotechnol. 19, 636–639 (2001).
Acknowledgements
The authors would like to thank O. Braha, S. Cheley, T. Reid and G. Sanghera for valuable discussion, S. Lewis for cyclodextrin preparation and analysis, M. Crawford, G. Hall and L. Woodward for nanopore testing, J. Kilgour and J. White for protein production, Z. McDougall for assistance with the manuscript, and all the staff at ONT. This work was supported by grants from the NIH, the European Commission's seventh Framework Programme (FP7) READNA Consortium and Oxford Nanopore Technologies. H.B. is the holder of a Royal Society Wolfson Research Merit Award.
Author information
Authors and Affiliations
Contributions
J.C. and H.B. conceived the experiments and wrote the manuscript. J.C. designed the mutant constructs and analysed the data. H.W. designed and synthesized the cyclodextrin. L.J. engineered the proteins. A.P. performed the single-channel recordings. S.R. wrote data analysis algorithms and software.
Corresponding author
Ethics declarations
Competing interests
H.B. is the founder, a director and a shareholder of Oxford Nanopore Technologies Ltd, a company engaged in the development of nanopore sequencing technology. J.C., L.J., A.P. and S.R. are employees of Oxford Nanopore Technologies Ltd.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1892 kb)
Rights and permissions
About this article
Cite this article
Clarke, J., Wu, HC., Jayasinghe, L. et al. Continuous base identification for single-molecule nanopore DNA sequencing. Nature Nanotech 4, 265–270 (2009). https://doi.org/10.1038/nnano.2009.12
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2009.12
This article is cited by
-
Peptide sequencing based on host–guest interaction-assisted nanopore sensing
Nature Methods (2024)
-
Biological big-data sources, problems of storage, computational issues, and applications: a comprehensive review
Knowledge and Information Systems (2024)
-
Comparison of methylation estimates obtained via MinION nanopore sequencing and sanger bisulfite sequencing in the TRPA1 promoter region
BMC Medical Genomics (2023)
-
High-resolution Nanopore methylome-maps reveal random hyper-methylation at CpG-poor regions as driver of chemoresistance in leukemias
Communications Biology (2023)
-
Gene sequencing and result analysis of balanced translocation carriers by third-generation gene sequencing technology
Scientific Reports (2023)