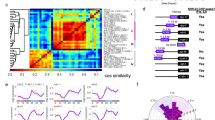
Abstract
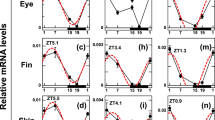
The only vertebrate clock gene identified by mutagenesis is mouse Clock , which encodes a bHLH-PAS transcription factor. We have cloned Clock in zebrafish and show that, in contrast to its mouse homologue, it is expressed with a pronounced circadian rhythm in the brain and in two defined pacemaker structures, the eye and the pineal gland. Clock oscillation was also found in other tissues, including kidney and heart. In these tissues, expression of Clock continues to oscillate in vitro . This demonstrates that self-sustaining circadian oscillators exist in several vertebrate organs, as was previously reported for invertebrates.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
King, D. P. et al. Positional cloning of the mouse circadian clock gene. Cell 89, 641–653 ( 1997).
Antoch, M. P. et al. Functional identification of the mouse circadian Clock gene by transgenic BAC rescue. Cell 89, 655– 667 (1997).
Vitaterna, M. H. et al. Mutagenesis and mapping of a mouse gene, Clock, essential for circadian behavior. Science 264, 719 –725 (1994).
Sun, Z. S. et al. RIGUI, a putative mammalian ortholog of the Drosophila period gene. Cell 90, 1003–1011 (1997).
Sassone-Corsi, P. Molecular clocks: mastering time by gene regulation. Nature 392, 871–874 (1998).
Dunlap, J. C. Genetics and molecular analysis of circadian rhythms. Annu. Rev. Genet. 30, 579–601 ( 1996).
Takahashi, J. S. Molecular neurobiology and genetics of circadian rhythms in mammals. Annu. Rev. Neurosci. 18, 531–553 (1995).
Allada, R., White, N. E., So, W. V., Hall, J. C. & Rosbash, M. A mutant Drosophila homolog of mammalian Clock disrupts circadian rhythms and transcription of period and timeless. Cell 93, 791–804 ( 1998).
Darlington, T. K. et al. Closing the circadian loop: CLOCK-induced transcription of its own inhibitors per and tim. Science 280, 1599–1603 (1998).
Bae, K., Lee, C., Sidote, D., Chuang, K.-Y. & Edery, I. Circadian regulation of a Drosophila homolog of the mammalian Clock gene: PER and TIM function as positive regulators. Mol. Cell. Biol. 18, 6142–6151 (1998).
Hardin, P. E., Hall, J. C. & Rosbash, M. Feedback of the Drosophila period gene product on circadian cycling of its messenger RNA levels. Nature 343, 536–540 (1990).
Hardin, P. E., Hall, J. C. & Rosbash, M. Circadian oscillations in period gene mRNA levels are transcriptionally regulated. Proc. Natl. Acad. Sci. USA 89, 11711–11715 (1992).
Sehgal, A. et al. Rhythmic expression of timeless: a basis for promoting circadian cycles in period gene autoregulation. Science 270, 808–810 (1995).
Zerr, D. M., Hall, J. C., Rosbash, M. & Siwicki, K. K. Circadian fluctuations of period protein immunoreactivity in the CNS and the visual system of Drosophila. J. Neurosci. 10, 2749– 2762 (1990).
Rutila, J. E. et al. CYCLE is a second bHLH-PAS clock protein essential for circadian rhythmicity and transcription of Drosophila period and timeless. Cell 93, 805–814 ( 1998).
Hogenesch, J. B., Gu, Y. Z., Jain, S. & Bradfield, C. A. The basic-helix-loop-helix-PAS orphan MOP3 forms transcriptionally active complexes with circadian and hypoxia factors. Proc. Natl. Acad. Sci. USA 95, 5474–5479 (1998).
Gekakis, N. et al. Role of the CLOCK protein in the mammalian circadian mechanism. Science 280, 1564–1569 (1998).
Plautz, J. D., Kaneko, M., Hall, J. C. & Kay, S. A. Independent photoreceptive circadian clocks throughout Drosophila. Science 278 , 1632–1635 (1997).
Moore, R. Y. & Eichler, V. B. Loss of a circadian adrenal corticosterone rhythm following suprachiasmatic lesions in the rat. Brain Res. 42, 201–206 ( 1972).
Stephan, F. K. & Zucker, I. Circadian rhythms in drinking behavior and locomotor activity of rats. Proc. Natl. Acad. Sci. USA 69, 1583–1586 (1972).
Tosini, G. & Menaker, M. Circadian rhythms in cultured mammalian retina. Science 272, 419– 421 (1996).
Tei, H. et al. Circadian oscillation of a mammalian homologue of the Drosophila period gene. Nature 389, 512– 516 (1997).
Albrecht, U., Sun, Z. S., Eichele, G. & Lee, C. C. A differential response of two putative mammalian circadian regulators, mper1 and mper2, to light. Cell 91, 1055– 1064 (1997).
Shearman, L. P., Zylka, M. J., Weaver, D. R., Kolakowski, L. F. Jr & Reppert, S. M. Two period homologs: circadian expression and photic regulation in the suprachiasmatic nuclei. Neuron 19, 1261– 1269 (1997).
Zylka, M. J., Shearman, L. P., Weaver, D. R. & Reppert, S. M. Three period homologs in mammals: differential light responses in the suprachiasmatic circadian clock and oscillating transcripts outside of brain. Neuron 20, 1103–1110 ( 1998).
Balsalobre, A., Damiola, F. & Schibler, U. A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell 93, 929– 937 (1998).
Cahill, G. M. Circadian regulation of melatonin production in cultured zebrafish pineal and retina. Brain Res. 708, 177– 181 (1996).
Rajendran, R. R. et al. Zebrafish interphotoreceptor retinoid-binding protein: differential circadian expression among cone subtypes. J. Exp. Biol. 199, 2775–2787 (1996).
Gonzalez, G. A. & Montminy, M. R. Cyclic AMP stimulates somatostatin gene transcription by phosphorylation of CREB at serine 133. Cell 59, 675–680 (1989).
Foulkes, N. S. & Sassone-Corsi, P. Transcription factors coupled to the cAMP-signalling pathway. Biochim. Biophys. Acta 1288, F101–121 ( 1996).
Shigeyoshi, Y. et al. Light-induced resetting of a mammalian circadian clock is associated with rapid induction of the mPer1 transcript. Cell 91, 1043–1053 (1997).
Reppert, S. M. & Weaver, D. R. Forward genetic approach strikes gold: cloning of a mammalian clock gene. Cell 89, 487–490 ( 1997).
Reppert, S. M. & Sauman, I. Period and timeless tango: a dance of two clock genes. Neuron 15, 983–986 (1995).
Fonjallaz, P., Ossipow, V., Wanner, G. & Schibler, U. The two PAR leucine zipper proteins, TEF and DBP, display similar circadian and tissue-specific expression, but have different target promoter preferences. Embo J. 15, 351–362 ( 1996).
Servillo, G., Della Fazia, M. A. & Viola-Magni, M. Variation of tyrosine aminotransferase expression during the day in rats of different ages. Biochem. Biophys. Res. Comm. 175, 104–109 (1991).
Reppert, S. M. & Weaver, D. R. Melatonin madness. Cell 83, 1059–1062 (1995).
Miyamoto, Y. & Sancar, A. Vitamin B2-based blue-light photoreceptors in the retinohypothalamic tract as the photoactive pigments for setting the circadian clock in mammals. Proc. Natl. Acad. Sci. USA 95, 6097–6102 (1998).
Klein, D. C., Moore, R. Y. & Reppert, S. M. (eds) Suprachiasmatic Nucleus—The Mind's Clock (Oxford Univ. Press, Oxford, 1991).
Chomczynski, P. & Sacchi, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162, 156– 159 (1987).
Foulkes, N. S., Borrelli, E. & Sassone-Corsi, P. CREM gene: use of alternative DNA-binding domains generates multiple antagonists of cAMP-induced transcription. Cell 64, 739–749 (1991).
Mellström, B., Naranjo, J. R., Foulkes, N. S., Lafarga, M. & Sassone-Corsi, P. Transcriptional response to cAMP in brain: specific distribution and induction of CREM antagonists. Neuron 10, 655–665 ( 1993).
Acknowledgements
We thank Joseph S. Takahashi, Nicolas Cermakian, Dario De Cesare, Lucia Monaco, Jean-Marie Garnier, Pilar Garcia-Villalba and Patrick Blader for discussions, advice and gifts of materials. We also acknowledge the technical assistance of Estelle Heitz, as well as Dominique Biellman, Odile Nkundwa, Nadine Fisher, Serge Vicaire and Frank Ruffenach. D.W. was supported by an EEC TMR fellowship. Our studies are funded by grants from CNRS, INSERM, CHUR, Rhône-Poulenc Rorer (Bioavenir), Fondation pour la Recherche Médicale and Association pour la Recherche sur le Cancer (P. S.-C.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Whitmore, D., Foulkes, N., Strähle, U. et al. Zebrafish Clock rhythmic expression reveals independent peripheral circadian oscillators. Nat Neurosci 1, 701–707 (1998). https://doi.org/10.1038/3703
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/3703
This article is cited by
-
Circadian regulation of developmental synaptogenesis via the hypocretinergic system
Nature Communications (2023)
-
Cloning, tissue distribution, mRNA expression and functional analysis of circadian clock gene per2 from the high-latitude Amur minnow (Phoxinus lagowskii)
Aquaculture International (2023)
-
Daily rhythms in gene expression of the human parasite Schistosoma mansoni
BMC Biology (2021)
-
Adaptation and convergence in circadian‐related genes in Iberian freshwater fish
BMC Ecology and Evolution (2021)
-
Finding Nemo’s clock reveals switch from nocturnal to diurnal activity
Scientific Reports (2021)