Abstract
Neuroscientists increasingly analyze the joint activity of multineuron recordings to identify population-level structures believed to be significant and scientifically novel. Claims of significant population structure support hypotheses in many brain areas. However, these claims require first investigating the possibility that the population structure in question is an expected byproduct of simpler features known to exist in data. Classically, this critical examination can be either intuited or addressed with conventional controls. However, these approaches fail when considering population data, raising concerns about the scientific merit of population-level studies. Here we develop a framework to test the novelty of population-level findings against simpler features such as correlations across times, neurons and conditions. We apply this framework to test two recent population findings in prefrontal and motor cortices, providing essential context to those studies. More broadly, the methodologies we introduce provide a general neural population control for many population-level hypotheses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Cunningham, J.P. & Yu, B.M. Dimensionality reduction for large-scale neural recordings. Nat. Neurosci. 17, 1500–1509 (2014).
Gao, P. & Ganguli, S. On simplicity and complexity in the brave new world of large-scale neuroscience. Curr. Opin. Neurobiol. 32, 148–155 (2015).
Stevenson, I.H. & Kording, K.P. How advances in neural recording affect data analysis. Nat. Neurosci. 14, 139–142 (2011).
Pillow, J.W. et al. Spatio-temporal correlations and visual signalling in a complete neuronal population. Nature 454, 995–999 (2008).
Stopfer, M., Jayaraman, V. & Laurent, G. Intensity versus identity coding in an olfactory system. Neuron 39, 991–1004 (2003).
Saha, D. et al. A spatiotemporal coding mechanism for background-invariant odor recognition. Nat. Neurosci. 16, 1830–1839 (2013).
Machens, C.K., Romo, R. & Brody, C.D. Functional, but not anatomical, separation of “what” and “when” in prefrontal cortex. J. Neurosci. 30, 350–360 (2010).
Mante, V., Sussillo, D., Shenoy, K.V. & Newsome, W.T. Context-dependent computation by recurrent dynamics in prefrontal cortex. Nature 503, 78–84 (2013).
Churchland, M.M. et al. Neural population dynamics during reaching. Nature 487, 51–56 (2012).
Sadtler, P.T. et al. Neural constraints on learning. Nature 512, 423–426 (2014).
Raposo, D., Kaufman, M.T. & Churchland, A.K. A category-free neural population supports evolving demands during decision-making. Nat. Neurosci. 17, 1784–1792 (2014).
Morcos, A.S. & Harvey, C.D. History-dependent variability in population dynamics during evidence accumulation in cortex. Nat. Neurosci. 19, 1672–1681 (2016).
Elsayed, G.F., Lara, A.H., Kaufman, M.T., Churchland, M.M. & Cunningham, J.P. Reorganization between preparatory and movement population responses in motor cortex. Nat. Commun. 7, 13239 (2016).
Hubel, D.H. & Wiesel, T.N. Receptive fields and functional architecture in two nonstriate visual areas (18 and 19) of the cat. J. Neurophysiol. 28, 229–289 (1965).
Georgopoulos, A.P., Schwartz, A.B. & Kettner, R.E. Neuronal population coding of movement direction. Science 233, 1416–1419 (1986).
Rigotti, M. et al. The importance of mixed selectivity in complex cognitive tasks. Nature 497, 585–590 (2013).
Murray, J.D. et al. Stable population coding for working memory coexists with heterogeneous neural dynamics in prefrontal cortex. Proc. Natl. Acad. Sci. USA 114, 394–399 (2017).
Broome, B.M., Jayaraman, V. & Laurent, G. Encoding and decoding of overlapping odor sequences. Neuron 51, 467–482 (2006).
Sussillo, D., Churchland, M.M., Kaufman, M.T. & Shenoy, K.V. A neural network that finds a naturalistic solution for the production of muscle activity. Nat. Neurosci. 18, 1025–1033 (2015).
Maass, W., Natschläger, T. & Markram, H. Real-time computing without stable states: a new framework for neural computation based on perturbations. Neural Comput. 14, 2531–2560 (2002).
Cunningham, J.P., Gilja, V., Ryu, S.I. & Shenoy, K.V. Methods for estimating neural firing rates, and their application to brain-machine interfaces. Neural Netw. 22, 1235–1246 (2009).
London, M., Roth, A., Beeren, L., Häusser, M. & Latham, P.E. Sensitivity to perturbations in vivo implies high noise and suggests rate coding in cortex. Nature 466, 123–127 (2010).
Gerstein, G.L. & Perkel, D.H. Simultaneously recorded trains of action potentials: analysis and functional interpretation. Science 164, 828–830 (1969).
Cohen, M.R. & Kohn, A. Measuring and interpreting neuronal correlations. Nat. Neurosci. 14, 811–819 (2011).
Gawne, T.J. & Richmond, B.J. How independent are the messages carried by adjacent inferior temporal cortical neurons? J. Neurosci. 13, 2758–2771 (1993).
Yu, B.M. et al. Gaussian-process factor analysis for low-dimensional single-trial analysis of neural population activity. J. Neurophysiol. 102, 614–635 (2009).
Schneidman, E., Berry, M.J. II, Segev, R. & Bialek, W. Weak pairwise correlations imply strongly correlated network states in a neural population. Nature 440, 1007–1012 (2006).
Romo, R., Brody, C.D., Hernández, A. & Lemus, L. Neuronal correlates of parametric working memory in the prefrontal cortex. Nature 399, 470–473 (1999).
Brody, C.D., Hernández, A., Zainos, A. & Romo, R. Timing and neural encoding of somatosensory parametric working memory in macaque prefrontal cortex. Cereb. Cortex 13, 1196–1207 (2003).
Shenoy, K.V., Sahani, M. & Churchland, M.M. Cortical control of arm movements: a dynamical systems perspective. Annu. Rev. Neurosci. 36, 337–359 (2013).
Tang, A. et al. A maximum entropy model applied to spatial and temporal correlations from cortical networks in vitro. J. Neurosci. 28, 505–518 (2008).
Shlens, J. et al. The structure of multi-neuron firing patterns in primate retina. J. Neurosci. 26, 8254–8266 (2006).
Shlens, J. et al. The structure of large-scale synchronized firing in primate retina. J. Neurosci. 29, 5022–5031 (2009).
Kimmel, D., Elsayed, G.F., Cunningham, J.P., Rangel, A. & Newsome, W.T. Encoding of value and choice as separable, dynamic neural dimensions in orbitofrontal cortex. Cosyne 2016.
Churchland, M.M. & Shenoy, K.V. Temporal complexity and heterogeneity of single-neuron activity in premotor and motor cortex. J. Neurophysiol. 97, 4235–4257 (2007).
Hernández, A. et al. Decoding a perceptual decision process across cortex. Neuron 66, 300–314 (2010).
Kobak, D. et al. Demixed principal component analysis of neural population data. eLife 5, e10989 (2016).
Romo, R. & Salinas, E. Flutter discrimination: neural codes, perception, memory and decision making. Nat. Rev. Neurosci. 4, 203–218 (2003).
Churchland, M.M., Cunningham, J.P., Kaufman, M.T., Ryu, S.I. & Shenoy, K.V. Cortical preparatory activity: representation of movement or first cog in a dynamical machine? Neuron 68, 387–400 (2010).
Fetz, E.E. Are movement parameters recognizably coded in the activity of single neurons. Behav. Brain Sci. 15, 679–690 (1992).
Scott, S.H. Population vectors and motor cortex: neural coding or epiphenomenona? Nat. Neurosci. 3, 307–308 (2000).
Churchland, M.M., Afshar, A. & Shenoy, K.V. A central source of movement variability. Neuron 52, 1085–1096 (2006).
Kaufman, M.T. et al. Roles of monkey premotor neuron classes in movement preparation and execution. J. Neurophysiol. 104, 799–810 (2010).
Kaufman, M.T., Churchland, M.M., Ryu, S.I. & Shenoy, K.V. Cortical activity in the null space: permitting preparation without movement. Nat. Neurosci. 17, 440–448 (2014).
Michaels, J.A., Dann, B. & Scherberger, H. Neural population dynamics during reaching are better explained by a dynamical system than representational tuning. PLOS Comput. Biol. 12, e1005175 (2016).
Hennequin, G., Vogels, T.P. & Gerstner, W. Optimal control of transient dynamics in balanced networks supports generation of complex movements. Neuron 82, 1394–1406 (2014).
Ecker, A.S. et al. State dependence of noise correlations in macaque primary visual cortex. Neuron 82, 235–248 (2014).
Park, I.M., Meister, M.L., Huk, A.C. & Pillow, J.W. Encoding and decoding in parietal cortex during sensorimotor decision-making. Nat. Neurosci. 17, 1395–1403 (2014).
Kaufman, M.T., Churchland, M.M., Ryu, S.I. & Shenoy, K.V. Vacillation, indecision and hesitation in moment-by-moment decoding of monkey motor cortex. Elife 4, e04677 (2015).
Victor, J.D. Spike train metrics. Curr. Opin. Neurobiol. 15, 585–592 (2005).
Boumal, N., Mishra, B., Absil, P.A. & Sepulchre, R. Manopt, a Matlab toolbox for optimization on manifolds. J. Mach. Learn. Res. 15, 1455–1459 (2014).
Cunningham, J.P. & Ghahramani, Z. Linear dimensionality reduction: survey, insights, and generalizations. J. Mach. Learn. Res. 16, 2859–2900 (2015).
Gilboa, E., Saatçi, Y. & Cunningham, J.P. Scaling multidimensional inference for structured Gaussian processes. IEEE Trans. Pattern Anal. Mach. Intell. 37, 424–436 (2015).
Acknowledgements
We thank the laboratories of L. Paninski, M. Churchland and K. Shenoy for discussions. We thank M. Churchland, L. Abbott and K. Miller for comments on the manuscript. We thank D. Kobak and C. Machens for discussions about and assistance with the dPCA algorithm. We thank M. Kaufman for help with Figure 1a. We thank M. Churchland, M. Kaufman, S. Ryu and K. Shenoy for the motor cortex data. We thank R. Romo and C. Brody for the prefrontal cortex data, downloaded from the CRCNS (available at the time of publication at https://crcns.org/data-sets/pfc/pfc-4). We thank T. Requarth for comments on the manuscript. We thank the 2016 Modeling Neural Activity conference for discussions and for a travel grant to GFE (MH 064537, NSF-DMS 1612914 and the Burroughs-Wellcome Fund). This work was funded by NIH CRCNS R01 NS100066-01, the Sloan Research Fellowship, the McKnight Fellowship, the Simons Collaboration on the Global Brain SCGB325233, the Grossman Center for the Statistics of Mind, the Center for Theoretical Neuroscience, the Gatsby Charitable Trust and the Zuckerman Mind Brain Behavior Institute.
Author information
Authors and Affiliations
Contributions
G.F.E. and J.P.C. contributed to all aspects of this study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Corrected Fisher randomization (CFR) and tensor maximum entropy (TME) methods.
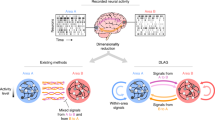
(a) Creating S surrogate datasets using CFR (top) and TME methods (bottom). Sample CFR: original neural responses are shuffled across conditions, yielding a randomized dataset with distorted primary features. Optimize CFR: find the best neural readout that operates on the shuffled data to retain the primary features of the original dataset. Optimize TME: find the maximum entropy distribution of tensor-valued datasets that has the same primary features (first and second marginal moments) as the original dataset. Sample TME: use efficient Kronecker methods to sample from the maximum entropy distribution. (b) Using surrogate datasets from CFR or TME to test the null hypothesis that population structure is an expected byproduct of given primary features. Quantify structure: evaluate a summary statistic such as R2 or variance explained that quantifies the degree that a hypothesized population structure exists in population data. Evaluate the summary statistic from the original neural data and from S surrogate datasets. Use the S values of the summary statistics from the surrogate datasets as the null distribution of population structure arising from the primary features alone. The summary statistic from the original neural data is then compared to that null distribution to obtain a p-value for the null hypothesis.
Supplementary Figure 2 Quantification of primary features in the surrogate datasets based on motor cortex data.
Each surrogate dataset Xsurr(i) (for i ∈ {1, …,S}, S=100) has marginal covariance ΣT (i), ΣN (i), and ΣC (i), which, in the surrogate-TNC control (right column of each panel), should match the specified primary features ΣT, ΣN, and ΣC of the original neural data. The right column demonstrates that fit: the top two rows (panel a) of the right column shows that CFR and TME match the true covariances in expectation (see equation on vertical axis); the bottom two rows (panel b) show that CFR has very minor variance around that mean, whereas TME has meaningful variance (see equation on vertical axis), as expected. Each dot corresponds to one of the nine separate datasets collected in the motor cortex (A, B, J1, J2, J3, J4, J-Array, N, and N-Array; see Methods for data details). The left and middle columns of each panel of the figure correspond to the surrogate-T (left) and surrogate-TN (middle) controls. Here we see as expected that all covariances that are specified in the control ΣT in the left column; both ΣT and ΣN in the middle column) are very well matched. Correspondingly, we also see that covariances that are not specified in the control do not match the moments of the data. Taken together, these data demonstrate quantitatively that the CFR and TME methods behave as desired, both in terms of preserving the specified structure, and in terms of destroying structure not specified.
Supplementary Figure 3 Quantification of primary features in the surrogate datasets based on PFC data.
Each surrogate dataset Xsurr(i) (for i {1, …,S}, S=100) has marginal covariances ΣT (i), ΣN (i), and ΣC (i), which, in the surrogate-TNC control (right column of each panel), should match the specified primary features ΣT, ΣN, and ΣC of the original neural data. The right column demonstrates that fit: the top two rows (panel a) of the right column shows that CFR and TME match the true covariances in expectation (see equation on vertical axis); the bottom two rows (panel b) show that CFR has very minor variance around that mean, whereas TME has meaningful variance (again see equation on vertical axis), as expected. Each dot corresponds to one of the two datasets collected in the prefrontal cortex (RR15 and RR14; see Methods for data details). The left and middle columns of each panel of the figure correspond to the surrogate-T (left) and surrogate-TN (middle) controls. Here we see as expected that all covariances that are specified in the control (ΣT in the left column; both ΣT and ΣN in the middle column) are very well matched. Correspondingly, we also see that covariances that are not specified in the control do not match the moments of the data. Taken together, these data demonstrate quantitatively that the CFR and TME methods behave as desired, both in terms of preserving the specified structure, and in terms of destroying structure not specified.
Supplementary Figure 4 Neural responses in prefrontal cortex vs. surrogates.
This figure is similar to Fig. 3 from the main text but showing preprocessed data with soft-normalization and mean-condition subtraction. (a) Example neuron (neuron number 15 of 571 total) from prefrontal cortex. Each trace is the trial-averaged firing of the twelve task conditions (six stimuli and two decisions). The traces color and style reflect the twelve possible experimental conditions. Horizontal bars denote the times of first (F1) and second (F2) vibrotactile stimuli. (b-d) Example neurons from one surrogate-T, surrogate-TN, and surrogate-TNC dataset, respectively. b-d panels follow the same convention as panel a. Top panels in b-d are surrogate datasets generated using Corrected Fisher Randomization (CFR) and bottom panels in b-d are surrogate datasets generated using Tensor Maximum Entropy (TME).
Supplementary Figure 5 Neural responses in motor cortex vs. surrogates.
This figure is showing preprocessed data with soft-normalization and mean-condition subtraction. (a) Example neuron (neuron number 175 of 218 total) recorded from the motor cortex of one monkey during the delayed-reach task. Each trace is the trial-averaged normalized rate during one of 108 reaching conditions (neuron 175 from monkey N-Array; see Methods). The trace color indicates the reach condition from Fig. 5a in the main text. (b-d) Example neurons from one surrogate-T, surrogate-TN, and surrogate-TNC dataset, respectively. b-d panels follow the same convention as panel a. Top panels in b-d are surrogate datasets generated using Corrected Fisher Randomization (CFR) and bottom panels in b-d are surrogate datasets generated using Tensor Maximum Entropy (TME). Scale bars and time markers in b-d match the scale bar and time markers in a.
Supplementary Figure 6 Decision (or stimulus) readouts in PFC are not (or are) an expected byproduct.
This figure is similar to Fig. 4e,f but with reconstruction variance (termed explained variance in Kobak et al.37) instead of the conventional percentage variance explained. (a) Percent reconstruction variance of the population projection onto the top decision readout. Black lines show the percent variance explained from the original neural data, colored box-whisker plots show the variance explained distribution from 200 surrogate samples (same convention as Fig. 4 e,f; stars denote significantly high variance; upper-tail test). The variance of the decision projection is calculated during the decision epoch (100 ms after the second stimulus onset, until the second stimulus offset). (b) Same as a but for reconstruction variance of the population projection onto the top stimulus readout. The variance of the stimulus projection is calculated during the stimulus epoch (100 ms after first stimulus onset, until the second stimulus onset).
Supplementary Figure 7 Single-neuron tuning to stimulus is prominent in PFC.
Left column: the lower triangle is the unique values of the condition covariance matrix (lower triangle of ΣC) of the original neural data (normalized to have unit norm); the upper triangle is the unique values of the outer product of the stimulus vector with itself (again normalized to have unit norm, shown above the diagonal line). Right column: lower triangle is the same as the left panel but the upper triangle is now the unique values of the outer product of the decision vector (right). Top row is for monkey RR15 and bottom row is for monkey RR14 (see Methods for data details). The left column has a closer qualitative match between upper and lower triangles, indicating stimulus tuning is similar to the condition covariance matrix. Compare to the right column, which looks highly non-symmetric, meaning that ΣC does not capture decision tuning well.
Supplementary Figure 8 Dimensionality of the linear dynamical system structure in motor cortex.
We measured the fit quality (R2) of the motor cortex data to a linear dynamical system with different dimensionalities using leave-one-condition-out cross-validation. Model dimensionalities in the horizontal axis reflect the number of principal components retained in the population response before fitting the dynamical system. Colored traces show the R2 from different datasets from the motor cortex (nine datasets). Black dot is the highest cross-validated R2 value, which is used in Fig. 6a and Supplementary Fig. 9. Note that all test results are robust to this choice of dimensionality over a wide range (all but the smallest dimensionalities; see Fig. 6b).
Supplementary Figure 9 Population dynamics in motor cortex from many datasets are consistently significant.
Neural responses in the motor cortex from the 400 ms duration reflecting movement response were projected onto the top PCs (see Supplementary Fig. 8), and then fit to a linear dynamical system. (a) Quality of fit (R2) of the original neural responses to the dynamical system model. Black bar denotes the R2 from the original neural data. Colored bars denote the median R2 from 100 different surrogates from each control type (error bars denote the 95th percentile of the distribution). Stars denote significantly higher R2 value in the original data than in the surrogates (P<0.001; upper-tail test). (b) Same as a, but using leave-one-condition-out cross-validated R2.
Supplementary Figure 10 Quasirhythmic population response in motor cortex is not an expected byproduct.
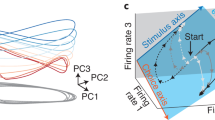
Churchland et al.9 identified a quasi-rhythmic population response during reaching from motor cortex data. They designed a dimensionality reduction method (jPCA; Churchland et al.9) to identify a population projection, which revealed the presence or absence of rotational dynamics in the neural trajectories. Some have expressed concern that the observed population oscillations may be an expected byproduct of the primary features of neural data. (a) Projections of population responses onto the top jPCA plane. Each trace shows neural trajectories reflecting the first 200 ms of movement-related activity for one of 108 reaching conditions. The colors of the traces are based on the variance of the preparatory state along jPC1. (b) jPCA projections from one sample of each surrogate type. (c) Quality of fit (R2) of the jPCA oscillatory dynamical system model of the original neural data and the surrogate datasets (same convention as Fig. 6a). This figure validates the claim that quasi-rhythmic dynamics are not an expected byproduct.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Notes 1–4. (PDF 1904 kb)
Rights and permissions
About this article
Cite this article
Elsayed, G., Cunningham, J. Structure in neural population recordings: an expected byproduct of simpler phenomena?. Nat Neurosci 20, 1310–1318 (2017). https://doi.org/10.1038/nn.4617
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4617