Abstract
Susceptibility to obesity is linked to genes regulating neurotransmission, pancreatic beta-cell function and energy homeostasis. Genome-wide association studies have identified associations between body mass index and two loci near cell adhesion molecule 1 (CADM1) and cell adhesion molecule 2 (CADM2), which encode membrane proteins that mediate synaptic assembly. We found that these respective risk variants associate with increased CADM1 and CADM2 expression in the hypothalamus of human subjects. Expression of both genes was elevated in obese mice, and induction of Cadm1 in excitatory neurons facilitated weight gain while exacerbating energy expenditure. Loss of Cadm1 protected mice from obesity, and tract-tracing analysis revealed Cadm1-positive innervation of POMC neurons via afferent projections originating from beyond the arcuate nucleus. Reducing Cadm1 expression in the hypothalamus and hippocampus promoted a negative energy balance and weight loss. These data identify essential roles for Cadm1-mediated neuronal input in weight regulation and provide insight into the central pathways contributing to human obesity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Finkelstein, E.A. et al. The lifetime medical cost burden of overweight and obesity: implications for obesity prevention. Obesity (Silver Spring) 16, 1843–1848 (2008).
McCarthy, M.I. & Zeggini, E. Genome-wide association studies in type 2 diabetes. Curr. Diab. Rep. 9, 164–171 (2009).
Grant, S.F.A. et al. Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat. Genet. 38, 320–323 (2006).
Frayling, T.M. Genome-wide association studies provide new insights into type 2 diabetes aetiology. Nat. Rev. Genet. 8, 657–662 (2007).
Willer, C.J. et al. Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nat. Genet. 41, 25–34 (2009).
Speliotes, E.K. et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 42, 937–948 (2010).
Locke, A.E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Biederer, T. et al. SynCAM, a synaptic adhesion molecule that drives synapse assembly. Science 297, 1525–1531 (2002).
Fogel, A.I. et al. SynCAMs organize synapses through heterophilic adhesion. J. Neurosci. 27, 12516–12530 (2007).
Fogel, A.I. et al. N-glycosylation at the SynCAM (synaptic cell adhesion molecule) immunoglobulin interface modulates synaptic adhesion. J. Biol. Chem. 285, 34864–34874 (2010).
GTEx Consortium. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Biederer, T. Bioinformatic characterization of the SynCAM family of immunoglobulin-like domain-containing adhesion molecules. Genomics 87, 139–150 (2006).
Badman, M.K., Kennedy, A.R., Adams, A.C., Pissios, P. & Maratos-Flier, E. A very low carbohydrate ketogenic diet improves glucose tolerance in ob/ob mice independently of weight loss. Am. J. Physiol. Endocrinol. Metab. 297, E1197–E1204 (2009).
Elias, C.F. et al. Leptin differentially regulates NPY and POMC neurons projecting to the lateral hypothalamic area. Neuron 23, 775–786 (1999).
Brüning, J.C. et al. Role of brain insulin receptor in control of body weight and reproduction. Science 289, 2122–2125 (2000).
Ahima, R.S., Bjorbaek, C., Osei, S. & Flier, J.S. Regulation of neuronal and glial proteins by leptin: implications for brain development. Endocrinology 140, 2755–2762 (1999).
Tschöp, M.H. et al. A guide to analysis of mouse energy metabolism. Nat. Methods 9, 57–63 (2011).
Huo, L. et al. Leptin-dependent control of glucose balance and locomotor activity by POMC neurons. Cell Metab. 9, 537–547 (2009).
Vong, L. et al. Leptin action on GABAergic neurons prevents obesity and reduces inhibitory tone to POMC neurons. Neuron 71, 142–154 (2011).
Robbins, E.M. et al. SynCAM 1 adhesion dynamically regulates synapse number and impacts plasticity and learning. Neuron 68, 894–906 (2010).
Park, K.A. et al. Excitatory synaptic drive and feedforward inhibition in the hippocampal CA3 circuit are regulated by SynCAM 1. J. Neurosci. 36, 7464–7475 (2016).
Kerchner, G.A. & Nicoll, R.A. Silent synapses and the emergence of a postsynaptic mechanism for LTP. Nat. Rev. Neurosci. 9, 813–825 (2008).
Turrigiano, G. Too many cooks? Intrinsic and synaptic homeostatic mechanisms in cortical circuit refinement. Annu. Rev. Neurosci. 34, 89–103 (2011).
Perez de Arce, K. et al. Topographic mapping of the synaptic cleft into adhesive nanodomains. Neuron 88, 1165–1172 (2015).
van Praag, H., Kempermann, G. & Gage, F.H. Running increases cell proliferation and neurogenesis in the adult mouse dentate gyrus. Nat. Neurosci. 2, 266–270 (1999).
van Praag, H., Christie, B.R., Sejnowski, T.J. & Gage, F.H. Running enhances neurogenesis, learning, and long-term potentiation in mice. Proc. Natl. Acad. Sci. USA 96, 13427–13431 (1999).
Hsu, Y.-W.A. et al. Role of the dorsal medial habenula in the regulation of voluntary activity, motor function, hedonic state, and primary reinforcement. J. Neurosci. 34, 11366–11384 (2014).
Bianco, I.H. & Wilson, S.W. The habenular nuclei: a conserved asymmetric relay station in the vertebrate brain. Philos Trans R Soc Lond B Biol Sci 364, 1005–1020 (2009).
Balthasar, N. et al. Leptin receptor signaling in POMC neurons is required for normal body weight homeostasis. Neuron 42, 983–991 (2004).
Balthasar, N. et al. Divergence of melanocortin pathways in the control of food intake and energy expenditure. Cell 123, 493–505 (2005).
Hill, J.W. et al. Direct insulin and leptin action on pro-opiomelanocortin neurons is required for normal glucose homeostasis and fertility. Cell Metab. 11, 286–297 (2010).
DeFalco, J. et al. Virus-assisted mapping of neural inputs to a feeding center in the hypothalamus. Science 291, 2608–2613 (2001).
Billings, L.K. & Florez, J.C. The genetics of type 2 diabetes: what have we learned from GWAS? Ann. NY Acad. Sci. 1212, 59–77 (2010).
Loos, R.J.F. & Yeo, G.S.H. The bigger picture of FTO: the first GWAS-identified obesity gene. Nat. Rev. Endocrinol. 10, 51–61 (2014).
Moser, M.-B., Rowland, D.C. & Moser, E.I. Place cells, grid cells and memory. Cold Spring Harb. Perspect. Biol. 7, a021808 (2015).
Hartley, T., Lever, C., Burgess, N. & O'Keefe, J. Space in the brain: how the hippocampal formation supports spatial cognition. Phil. Trans. R. Soc. Lond. B 369, 20120510 (2013).
Zeltser, L.M., Seeley, R.J. & Tschöp, M.H. Synaptic plasticity in neuronal circuits regulating energy balance. Nat. Neurosci. 15, 1336–1342 (2012).
van der Weyden, L. et al. Loss of TSLC1 causes male infertility due to a defect at the spermatid stage of spermatogenesis. Mol. Cell. Biol. 26, 3595–3609 (2006).
Harno, E., Cottrell, E.C. & White, A. Metabolic pitfalls of CNS Cre-based technology. Cell Metab. 18, 21–28 (2013).
Tattikota, S.G. et al. Argonaute2 mediates compensatory expansion of the pancreatic β cell. Cell Metab. 19, 122–134 (2014).
Glynn, M.W. & McAllister, A.K. Immunocytochemistry and quantification of protein colocalization in cultured neurons. Nat. Protoc. 1, 1287–1296 (2006).
Butler, A.A. & Kozak, L.P. A recurring problem with the analysis of energy expenditure in genetic models expressing lean and obese phenotypes. Diabetes 59, 323–329 (2010).
Birkenfeld, A.L. et al. Deletion of the mammalian INDY homolog mimics aspects of dietary restriction and protects against adiposity and insulin resistance in mice. Cell Metab. 14, 184–195 (2011).
Chevalier, C. et al. Gut microbiota orchestrates energy homeostasis during cold. Cell 163, 1360–1374 (2015).
Jeffrey, M. et al. Synapse loss associated with abnormal PrP precedes neuronal degeneration in the scrapie-infected murine hippocampus. Neuropathol. Appl. Neurobiol. 26, 41–54 (2000).
Pinto, S. et al. Rapid rewiring of arcuate nucleus feeding circuits by leptin. Science 304, 110–115 (2004).
Cowley, M.A. et al. Leptin activates anorexigenic POMC neurons through a neural network in the arcuate nucleus. Nature 411, 480–484 (2001).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 57, 289–300 (1995).
Acknowledgements
The authors would like to thank M. Gruhn and the Biozentrum Imaging Facility, University of Cologne, for access to Amira Software, and N. Zampieri, A. Plested, T. Breiderhoff, D. Matthäus, I. Park, C. Teng, T. Klüssendorf, H. Wessels, T. Willnow and M. Gotthardt for helpful discussions and assistance in the conduct of this work. This work was funded by the Helmholtz Gemeinschaft, the Helmholtz Metabolic Dysfunction Consortium, the Helmholtz Alliance ICEMED (Project 1210251 to T.N.), the European Research Council (ERC-2010-StG-260744 to M.N.P., ERC-2015-CoG-682422 to J.F.A.P., ERC-2011-StG-280565 to J.S., and ERC-2013-StG-336607 to M.T.), the US National Institutes of Health (R01-DK-111178, 1P01-AG-051459 and 1R56-AG-052986 to T.L.H.) the Swiss National Science Foundation Professorship (PP00P3_144886 to M.T.), the Deutsche Forschungsgemeinschaft (FOR-2143-Interneuron to J.F.A.P., Exc-257-NeuroCure to J.F.A.P. and V.H., and SFB958/A01 to V.H., and DFG BI1292/4-2 and DFG IRTG2251 to A.L.B.), the Federal Ministry of Education and Research (BMBF, Germany) (Project eMed:symAtrial (01ZX1408D to M.H.), the European Foundation for the Study of Diabetes (EFSD, Germany), the Thyssen Foundation, and the Kay Kendall Leukemia Foundation (KKLF Fellowship to L.v.d.W.). AAV reagents were provided by the UNC Vector Core facility and used with permission by K. Deisseroth (Stanford University).
Author information
Authors and Affiliations
Contributions
T.R. and M.N.P. conceived the study. T.R., X.Y., N.L.K., M.-C.K., K.S.-B., L.F., V.T., D.P., G.K., S.B., L.V., K.S., C.-X.Y., S.C.S., S.G.T., A.S.C., M.M., J.S., A.H., L.v.d.W., A.L.B., T.N., J.F.A.P., T.L.H., M.H.T., M.H., M.T., V.H. and M.N.P. designed and performed the experiments with help from all of the authors. V.H. and M.N.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 BMI risk SNPs associate with increased CADM1 and CADM2 expression in the cerebellum of human subjects.
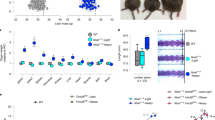
Boxplots show the 25% and 75% quantiles of normalized mRNA expression levels (y-axis), solid horizontal lines indicate the median, and whiskers indicate the 10% and 90% quantiles. (a) Elevated expression of CADM1 associates with risk allele (G) of rs12286929 in cerebellum. (b) Genotype dependent expression levels of CADM2 for the SNP rs13078807. The risk allele (G) is associated with higher expression levels in human cerebellum. Statistical analyses are described in the Methods and Supplementary Table 3.
Supplementary Figure 2 Increased Cadm1 expression in multiple brain regions of Lepob/ob mice is reversed upon ketogenic diet feeding.
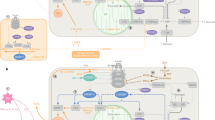
(a) Schematic representation of Cadm1 and Cadm2 engaged in homophilic (homo) and heterophilic (het) interactions. (b) (c) Western blot analysis of Cadm1 and Cadm2 in total and synaptosome-enriched lysates from cerebellum and hippocampus of 4-week-old Lepob/ob mice and littermate controls. (d) Quantification of western blot analysis of Cadm1 in multiple brain regions of 16-week-old WT and Lepob/ob mice on chow or ketogenic diet for 45 days. Cortex (ctx), striatum (str), hippocampus (hpc), hypothalamus (hyp), midbrain (mid), hindbrain (hind), and cerebellum (cer) are shown. (e, f) Western blot analysis of Cadm1 in cerebellum and hippocampus of 16-week-old wild-type on normal chow (WT/chow) and Lepob/ob mice on normal chow (Lepob/ob/chow) or ketogenic diet (Lepob/ob/keto). (g) Western blot analysis of Cadm1 from isolated hippocampus of 12-week old C57Bl/6 mice after administration of ketogenic diet for 4 or 8 days. Results are presented as mean ± s.e.m. *P<0.05 and **P<0.01. Boxplots show median, lower and upper quartiles (box), and minimum and maximum (whiskers). Statistical analyses are described in the Methods and Supplementary Table 3.
Supplementary Figure 3 Cadm1 and Cadm2 total knockout mice exhibit decreased body weight, and increased insulin sensitivity and energy expenditure.
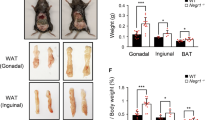
(a) Body weight curves of Cadm1KO mice (n=10) and littermate controls (n=12). (b) Body weight-to-length ratio in Cadm1KO mice (n=10) and control littermates at age 12-weeks (n=13). (c) Glucose measurements during an insulin tolerance test (ITT) on 12-week old Cadm1KO mice (n=5) and control littermates (n=5). (d) Glucose measurements during an intraperitoneal glucose tolerance test (GTT) on 12-week old Cadm1KO mice (n=5) and control littermates (n=6). (e) Glucose measurements during a pyruvate tolerance test (PTT) on 12-week old Cadm1KO mice (n=5) and control littermates (n=6). (f) Quantification of food intake in 12-week old Cadm1KO (n=5) and littermate controls (n=6) during leptin challenge. Daily food intake and body weight was measured for 5 days prior to leptin administration for base line. Differences in daily food intake were normalized to daily body weight. (g) Quantification of body weight change in 12-week old Cadm1KO (n=5) and littermate controls (n=6) during leptin challenge. Differences in daily body weight were normalized to daily body weight. (h) Quantification of daily food intake in 12-week old Cadm1KO (n=10) and littermate controls (n=10). (i) Quantification of O2 consumption, CO2 production, and RER, respectively in 12-week old male Cadm1KO mice (KO) (n=9) and control littermates (WT) (n=11). (j) Quantification of core body temperature (CBT) in 12-week old male Cadm1KO mice (KO) (n=5) and control littermates (WT) (n=5). (k) Quantification of energy expenditure during day and night phases in 12-week old male Cadm1KO mice (n=9) and control littermates (n=11). (l) Locomotor activity measured during day and night phases in 12-week old male Cadm1KO mice (n=8) and control littermates (n=8). (m) Energy expenditure of individual animals plotted against locomotor activity in 12-week old Cadm1KO (n=7) and littermate controls (n=11). (n) Western blot analysis of Cadm2 and Cadm1 in cortex, cerebellum (cereb), hippocampus (hpc), hypothalamus (hyp), liver and lung from Cadm2KO (KO) mice compared to littermate controls (WT). (o) Representative confocal images of primary hippocampal neurons isolated from Cadm2KO and littermate control mice, stained for Cadm2 (green) and MAP2 (red). (p) Body weight curves of Cadm2KO mice (n=8) and littermate controls (n=8) from age post-natal day 3-21. (q) Glucose measurements during an ITT on 9-week old Cadm2KO mice (n=6) and control littermates (n=6). (r) Glucose measurements during a GTT on Cadm2KO mice (n=3) and littermates (n=8) and quantification of area under the curve (AUC). (s) Quantification of food intake in 10-week old Cadm2KO mice (n=7) and littermate controls (n=9). (t, u) Quantification of O2 consumption, CO2 production, and RER, respectively in 10-week old Cadm2KO mice (n=6) and littermate controls (n=8). (v) Quantification of energy expenditure in 10-week old Cadm2KO mice (n=6) and littermate controls (n=8). (w) Quantification of locomotor activity in 10-week old Cadm2KO mice (n=6) and littermate controls (n=8). (x) Energy expenditure of individual animals plotted against lean body mass (LBM) in 10-week old Cadm2KO mice (n=7) and littermate controls (n=8). (y) Energy expenditure of individual animals plotted against locomotor activity in 10-week old Cadm2KO mice (n=7) and littermate controls (n=8). Results in panels a, c, d, e, f, g, p, g, and r are presented as mean ± s.e.m. *P<0.05, and **P<0.01. Boxplots show median, lower and upper quartiles (box), and minimum and maximum (whiskers). Statistical analyses are described in the Methods and Supplementary Table 3.
Supplementary Figure 4 Loss of Cadm1 reduces inhibitory post-synaptic currents in POMC neurons.
(a) Immunostaining of Cadm1 (red), POMC-eGFP (green) and VGLUT2 (magenta) identifies limited Cadm1 and VGLUT2 expression in POMC neurons in the arcuate nucleus region (ARC) of the hypothalamus. High magnification images are outlined by white boxes. (b) Immunostaining quantification of POMC-GFP and Cadm1-positive cells in the ARC (GFP-positive cells: 188±28; GFP- and CADM1-positive cells: 23±5). (c) qRT-PCR analysis of Pomc, Npy and Agrp expression in the ARC region of 12-week old Cadm1KO mice (n=3), Lepob/ob mice (n=3) and control littermates (WT) (n=4). (d) Summary of miniature IPSC frequency in POMC neurons from Cadm1KO and control littermates at the age of 3 weeks (top left). Example traces of IPSCs recorded from POMC neurons of Cadm1KO or control littermates (top right). Mean amplitude and cumulative probability distribution of IPSC frequencies from POMC neurons showing a significant shift in Cadm1KO mice (n=15) compared to littermate control mice (n=12) (lower left and right). (e) Summary of miniature EPSC frequency in POMC neurons from Cadm1KO and control littermates (top left). Example traces of EPSCs recorded from POMC neurons of Cadm1KO or control littermates (top right). Mean amplitude and cumulative probability distribution of EPSC frequencies from POMC neurons showing no change in Cadm1KO mice (n=7) compared to littermate control mice (n=11) (lower left and right). (f, g) Representative electron micrographs showing perikaryal membranes of eGFP-expressing POMC neurons from Cadm1KO (KO) and wild-type littermates (WT) mice identify marked differences in the asymmetrical (excitatory) synaptic inputs contacting POMC neurons (Scale bar, 1 μm). (h) Quantification of asymmetrical, symmetrical and total synapse numbers in in Cadm1KO mice (n=6) and littermate controls (n=8). Results are presented as mean ± s.e.m. *P<0.05, **P<0.01 and ***P<0.001. Boxplots show median, lower and upper quartiles (box), and minimum and maximum (whiskers). Statistical analyses are described in the Methods and Supplementary Table 3.
Supplementary Figure 5 Induction of Cadm1 in excitatory neurons increases co-localization with Cadm2.
(a) Confocal images revealing partial co-localization of Cadm1 and VGLUT2 in the different mouse brain regions. Cadm1 (green) and VGLUT2 (red) co-localize in hippocampus (Hpc), medial habenula (MHb), and paraventricular hypothalamus (PVH) regions. High magnification images are identified by white boxes. Fluorescent intensity profiles of Cadm1 and VGLUT2 (measured along white lines) quantify the co-localization of Cadm1 and VGLUT2. (b, c) Representative confocal images of Cadm1 and Cadm2 expression in Slc17a6-Cre and (d, e) Slc32a1-Cre-positive primary hippocampal neurons. Immunostaining for Cadm1 or Cadm2 (red), ZsGreen (green), and Map2 (cyan). (f) Representative confocal images of Cadm1 and Cadm2 expression in primary hippocampal neurons of Cadm1KO, Tg-Cadm1, and WT mice. (g) Quantification of O2 consumption, CO2 production, RER and energy expenditure (EE), in 14-week old Tg-Cadm1 mice (n=6) and littermate controls (n=8). (h) Representative confocal images of Cadm1, Cadm2, and VGLUT2 expression in primary hippocampal neurons of Cadm1KO, Tg-Cadm1, and WT mice. Immunofluorescence detection of Cadm1 (red), Cadm2 (green), and VGLUT2 (cyan) in dendritic branches identifies areas of co-localization. (i) Fluorescence intensity profiles of Cadm1 and Cadm2 (measured along white dashed lines in (h) quantify areas of co-localization. (j) Pearson correlation indicates increased association between Cadm1 and Cadm2 expression in Tg-Cadm1 (n=9) mice compared to wild-type control littermates (n=10). Results are presented as mean ± s.e.m. *P<0.05. Boxplots show median, lower and upper quartiles (box), and minimum and maximum (whiskers). Statistical analyses are described in the Methods and Supplementary Table 3.
Supplementary Figure 6 Loss of Cadm1 in excitatory neurons improves glucose homeostasis and insulin sensitivity.
(a) Confocal imaging of coronal brain slice identifying ZsGreen expression in Slc17a6-ires-Cre, lox-ZsGreen mice at age 12 weeks. (b) Western blot analysis of Slc17a6-Cre-mediated reduction of Cadm1 expression in hippocampus (hpc), cortex, and olfactory bulb (Olf bulb). (c) Confocal imaging of coronal brain slice identifying eGFP expression in Slc32a1-ires-Cre, lox-ZsGreen mice at age 12 weeks. (d) Western blot analysis of Slc32a1-Cre-mediated reduction of Cadm1 expression in hypothalamus (Hypoth), striatum (Striat), and olfactory bulb (Olf bulb). (e) Body weight curves in Slc17a6-Cre, Cadm1flox/flox mice (n=6) and littermate controls (n=6) from age post-natal day 0-21. (f) Body weight-to-length ratio in Slc17a6-Cre, Cadm1flox/flox mice (n=4) and control littermates (n=8) at age 3-weeks. (g) Body weight curves in Slc32a1-Cre, Cadm1flox/flox mice (n=8) and littermate controls from 4-12 weeks of age (n=8). (h) Glucose measurements during an ITT on 12-week old Slc32a1-Cre, Cadm1flox/flox mice (n=4) and control littermates (n=6) and quantification of area under the curve (AUC). (i) Glucose measurements during a GTT on Slc32a1-Cre, Cadm1flox/flox mice (n=5) and control littermates (n=8). (j) Energy expenditure of individual animals plotted against lean body mass in 12-week old Slc32a1-Cre, Cadm1flox/flox mice (n=7) and littermate controls (n=8). (k) Energy expenditure of individual animals plotted against locomotor activity in 12-week old Slc32a1-Cre, Cadm1flox/flox mice (n=7) and littermate controls (n=8). (l) Quantification of locomotor activity and food intake measured in 12-week old Slc32a1-Cre, Cadm1flox/flox mice (n=8) and littermate controls (n=8). (m) Body weight, blood glucose, and plasma insulin values before initiating hyperinsulinemic-euglycemic clamp studies on 12-week old Slc17a6-Cre, Cadm1flox/flox mice (n=6) and control littermates (n=4). (n) Plasma glucose concentrations and glucose infusion rate during hyperinsulinemic-euglycemic clamp studies on 12-week old Slc17a6-Cre, Cadm1flox/flox mice (n=6) and control littermates (n=4). (o) Endogenous glucose production in the basal and the clamped state and suppression of hepatic glucose production in Slc17a6-Cre, Cadm1flox/flox mice (n=6) and control littermates (n=4). (p) Peripheral glucose uptake, glycolysis rate, glycogen synthesis, plasma insulin levels during the hyperinsulinemic-euglycemic clamp studies in 12-week old Slc17a6-Cre, Cadm1flox/flox mice (n=6) and control littermates (n=4). Results in panels e, g, h, I, and n are presented as mean ± s.e.m. *P<0.05, **P<0.01 and ***P<0.001. Boxplots show median, lower and upper quartiles (box), and minimum and maximum (whiskers). Statistical analyses are described in the Methods and Supplementary Table 3.
Supplementary Figure 7 Loss of Cadm1 in excitatory neurons increases long-term potentiation and long-term depression in hippocampal neurons.
(a) Representative electron micrograph images from hippocampus of Slc17a6-Cre, Cadm1flox/flox mice and littermate controls. (b) Quantification of excitatory synapse number per volume fraction or per area and (c) Relative post-synaptic densities (PSD) length in hippocampus in Slc17a6-Cre, Cadm1flox/flox mice (n=3) and littermate controls (n=3). (d) Western blotting analysis of Cadm1 and PSD95 in total and synaptosome-enriched lysates from hippocampus of 12-week old Slc17a6-Cre, Cadm1flox/flox mice (n=2) and control littermates (n=2). (e) Cadm1 glutamatergic deficiency results in increased basal synaptic transmission. Representative fEPSPs responses at increasing stimulation intensities (10-100 μA, color coded) show higher amplitudes for Slc17a6-Cre, Cadm1flox/flox as compared to WT controls. Synaptic transmission was measured as the relationship of fEPSP slope vs fiber volley (FV) amplitudes. Two-way RM ANOVA revealed no significant changes for FV amplitudes (P=0.739). Slope of fEPSP was significantly facilitated (P=0.029) in Slc17a6-Cre, Cadm1flox/flox mice (WT n=18, N=9; Slc17a6-Cre, Cadm1flox/flox n=20, N=9). (f) LTP induction in MPP-DG synapses is facilitated in Slc17a6-Cre, Cadm1flox/flox mice. Representative traces are the average of 20 fEPSPs recorded 10 min before (black) and 50–60 min after LTP induction (grey). Higher LTP values measured 50–60 min after induction were detected in Slc17a6-Cre, Cadm1flox/flox as compared to WT control (P=0.021, WT n=8, N=8; Slc17a6-Cre, Cadm1flox/flox n=9, N=8). (g) LTD induction in MPP-DG synapses is facilitated in Slc17a6-Cre, Cadm1flox/flox mice. Representative traces are the average of 20 fEPSPs recorded 10 min before (black) and 50–60 min after LTD induction (grey). Facilitated LTD values measured 50–60 min after induction were detected in Slc17a6-Cre, Cadm1flox/floxas compared to WT control (P=0.032, WT n=9, N=8; Slc17a6-Cre, Cadm1flox/flox n=10, N=9). Results are presented as mean ± s.e.m. *P<0.05. Boxplots show median, lower and upper quartiles (box), and minimum and maximum (whiskers). Statistical analyses are described in the Methods and Supplementary Table 3.
Supplementary Figure 8 Tracing analysis identifies VGLUT2-positive afferent inputs to the ARC originating from the PVH, MHb, and Hpc regions.
(a) Western blot analysis of Cadm1 expression in total lysates from hippocampus and cortex after delivery of AAV-Cre to floxed Cadm1 mice (flox/flox) and control littermates (+/?). Total hippocampal lysates from Cadm1KO (KO) and Slc17a6-Cre, Cadm1flox/flox (Vglut2) serve as positive controls for loss of Cadm1 expression. (b) Quantification of O2 consumption, CO2 production, energy expenditure, locomotor activity and RER in 9-week old floxed Cadm1 mice (flox/flox) (n=7) and control littermates (+/?) (n=6). (c) Representative confocal images of coronal sections through the brain of Slc17a6-Cre/POMC-eGFP transgenic mice stereotaxically injected with AAV2/EF1a-DIO-hChR2(H134R)-mCherry into the paraventricular hypothalamus (PVH), habenular nuclei (Hb), and hippocampus (Hpc). Nuclei have been visualized with DAPI staining. High magnification images are identified by white boxes. (d) Representative coronal brain section of Cadm1 floxed mice after stereotaxic rAAV8-CaMKIIa-mCherry-Cre injection to hypothalamic neurons in the paraventricaular hypothalamus (PVH), and ventromedial hypothalamus (VMH) regions to target Cre expression to excitatory neurons in these areas; confocal imaging of mCherry and the neuronal marker NeuN. (e) Representative western blot analysis showing AAV-Cre-mediated deletion of Cadm1 after stereotaxic injection of rAAV8-CaMKIIa-mCherry-Cre into the PVH region of the hypothalamus. (f) Quantification of locomotor activity, O2 consumption, CO2 production, RER, and energy expenditure from 9-week old floxed Cadm1 mice (n=9) and control littermates (n=8) after stereotaxic injection of rAAV8/CamKII-mCherry-Cre injection into the hypothalamus. (g) Glucose measurements during an ITT on 9-week floxed Cadm1 mice (n=6) and control littermates (n=6) after stereotaxic injection of rAAV8/CamKII-mCherry-Cre injection into the hypothalamus. (h) Representative confocal images of coronal sections through the brain of Slc17a6-Cre/POMC-eGFP transgenic mice stereotaxically injected with AAV2/EF1a-DIO-hChR2(H134R)-mCherry into the hippocampus (Hpc). High magnification image of the PVH is identified by dashed box. (i) Representative confocal images of coronal sections through the brain of Slc17a6-Cre/POMC-eGFP transgenic mice stereotaxically injected with AAV2/EF1a-DIO-hChR2(H134R)-mCherry into the paraventricular hypothalamus (PVH). High magnification image of the hippocampus is identified by dashed box. (j) Confocal image analysis after stereotaxic injection of AAV2/EF1a-DIO-hChR2(H134R)-mCherry to the habenular nuclei (Hb) of Slc17a6-Cre/POMC-eGFP transgenic mice. Axonal varicosities (red) and POMC-positive neurons (green) are visualized within the ARC region of the hypothalamus. Dotted white circles outline eGFP-POMC-positive cell body, used for 3D reconstruction analysis. (k) Surface rendering of Amira 3D reconstruction of eGFP-POMC-positive cell body receiving afferent inputs from the MHb. Cell is represented in red, while the synaptic input is color-coded, with the cold to warm colors spreading from 0 to 250nm distance between axonal varicosities and the soma (see color-coded horizontal bar for the distance definition). (l) Histogram shows the number of anterograde AAV-mCherry-labelled MHb axonal varicosities found within 250nm distance from the POMC cell body. (m) Double immunostaining of Cadm1 and mCherry in MHb region identify Cadm1-positive anterograde projections to the ARC region of hypothalamus. Results in panel g are presented as mean ± s.e.m. *P<0.05, and **P<0.01. Boxplots show median, lower and upper quartiles (box), and minimum and maximum (whiskers). Statistical analyses are described in the Methods and Supplementary Table 3.
Supplementary Figure 9 Lepr-Cre-mediated deletion of Cadm1 impacts insulin sensitivity.
(a) Glucose measurements during an ITT on 12-week old Agrp-Cre, Cadm1flox/flox mice (n=3) and control littermates (n=5). (b) Glucose measurements during an ITT on 12-week old Pomc-Cre, Cadm1flox/flox mice (n=6) and control littermates (n=6). (c) Glucose measurements during an ITT on 12-week old Sim1-Cre, Cadm1flox/flox mice (n=6) and control littermates (n=10). (d) Body weight in Lepr-Cre, Cadm1flox/flox mice (n=4) and littermate controls (n=7). (e) Glucose measurements during an ITT on 12-week old Lepr-Cre, Cadm1flox/flox mice (n=3) and control littermates (n=5). (f) Correlation of energy expenditure of individual animals plotted against lean body mass in in 12-week old male Lepr-Cre, Cadm1flox/flox mice (n=7) and littermate controls (n=8). (g) Quantification of locomotor activity and daily food intake in 12-week old Lepr-Cre, Cadm1flox/flox mice (n=12) and littermate controls (n=12). (h) Body weight in ob/Lepr-Cre, Cadm1flox/flox mice (n=3) and Lepob/ob littermates (n=3). (i) Random-fed and fasted blood glucose and random-fed plasma insulin in 12-week old ob/Lepr-Cre, Cadm1flox/flox mice (n=5) and Lepob/ob littermates (n=7). (j) Glucose measurements during an insulin tolerance test on 12-week old ob/Lepr-Cre, Cadm1flox/flox mice (n=5) and Lepob/ob littermates (n=7). (k) Glucose measurements during a glucose tolerance test on 12-week old ob/Lepr-Cre, Cadm1flox/flox mice (n=5) and Lepob/ob littermates (n=6). (l) Glucose measurements during a pyruvate tolerance test on 12-week old ob/Lepr-Cre, Cadm1flox/flox mice (n=3) and Lepob/ob littermates (n=4). (m) Pancreatic β-cell mass and insulin content measurements in Lepr-Cre, Cadm1flox/flox (n=5), ob/Lepr-Cre, Cadm1flox/flox (n=7), Lepob/ob (n=5), and wild-type (WT) littermate controls (n=6) from 12 weeks of age. (n) Energy expenditure per individual animals plotted against lean body mass in 12-week old ob/Lepr-Cre, Cadm1flox/flox mice (n=6) and Lepob/ob littermates (n=7). (o) Quantification of locomotor activity and daily food intake in 12-week old ob/Lepr-Cre, Cadm1flox/flox mice (n=6) and Lepob/ob littermates. (n=8). Results in panels a-e, h, and j-l are presented as mean ± s.e.m. *P<0.05, and **P<0.01. Boxplots show median, lower and upper quartiles (box), and minimum and maximum (whiskers). Statistical analyses are described in the Methods and Supplementary Table 3.
Supplementary Figure 10 Immunostaining of Cadm1 with Vglut2-positive afferent projections in contact with POMC neurons.
(a) Quadruple immunostaining within the ARC region showing Cadm1 and Vglut2-positive afferent projections in contact with POMC neuron. Arrows indicate colocalization of mCherry punctas with Cadm1 and Vglut2. Slc17a6-Cre-positive mice received stereotaxic injection of AAV2-mCherry to the PVH, and were subsequently immunostained for POMC (green), mCherry (red), Cadm1 (magenta) and Vglut2 (cyan). (b) Visualization of Cadm1, mCherry, and Vglut2. Arrows indicate colocalization of mCherry punctas with Cadm1 and Vglut2.
Supplementary Figure 11 Original western blot panels with gel markers for Figures 2a, 2b, and 2f.
Complete scanned gels for western blots shown in Figures 2a, 2b, and 2f. Western images are overlaid onto image containing MW markers. White dashed line identifies cropped region shown in respective figure. Replicate experiments are shown at the right and unlabeled lanes contain samples not included in this study. All results are presented as mean ± s.e.m.
Supplementary Figure 12 Original western blot panels with gel markers for Figures 2f and 4e.
Complete scanned gels for western blots shown in Figures 2f, and 4e. Western images are overlaid onto image containing MW markers. White dashed line identifies cropped region shown in respective figure. All results are presented as mean ± s.e.m.
Supplementary Figure 13 Original western blot panels with gel markers for Supplementary Figures 2b, 2c, and 2e.
Complete scanned gels for western blots shown in Supplementary Figures 2b, 2c, and 2e. Western images are overlaid onto image containing MW markers. White dashed line identifies cropped region shown in respective figure. All results are presented as mean ± s.e.m.
Supplementary Figure 14 Original western blot panels with gel markers for Supplementary Figures 2g, 3n, 6b, and 6d.
Complete scanned gels for western blots shown in Supplementary Figures 2g, 3n, 6b, and 6d. Western images are overlaid onto image containing MW markers. White dashed line identifies cropped region shown in respective figure. All results are presented as mean ± s.e.m.
Supplementary Figure 15 Original western blot panels with gel markers for Supplementary Figures 7d, 8a, and 8e.
Complete scanned gels for western blots shown in Supplementary Figures 7d, 8a, and 8e. Western images are overlaid onto image containing MW markers. White dashed line identifies cropped region shown in respective figure. All results are presented as mean ± s.e.m.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15 and Supplementary Tables 1–3 (PDF 5060 kb)
Rights and permissions
About this article
Cite this article
Rathjen, T., Yan, X., Kononenko, N. et al. Regulation of body weight and energy homeostasis by neuronal cell adhesion molecule 1. Nat Neurosci 20, 1096–1103 (2017). https://doi.org/10.1038/nn.4590
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4590
This article is cited by
-
Brain-based gene expression of putative risk genes for anorexia nervosa
Molecular Psychiatry (2023)
-
Genetics and epigenetics in the obesity phenotyping scenario
Reviews in Endocrine and Metabolic Disorders (2023)
-
The mediating effect of DNA methylation in the association between maternal sleep during pregnancy and offspring adiposity status: a prospective cohort study
Clinical Epigenetics (2022)
-
Genetics and neurobiology of eating disorders
Nature Neuroscience (2022)
-
The genetics of obesity: from discovery to biology
Nature Reviews Genetics (2022)