Abstract
Brain function relies on fast and precisely timed synaptic vesicle (SV) release at active zones (AZs). Efficacy of SV release depends on distance from SV to Ca2+ channel, but molecular mechanisms controlling this are unknown. Here we found that distances can be defined by targeting two unc-13 (Unc13) isoforms to presynaptic AZ subdomains. Super-resolution and intravital imaging of developing Drosophila melanogaster glutamatergic synapses revealed that the Unc13B isoform was recruited to nascent AZs by the scaffolding proteins Syd-1 and Liprin-α, and Unc13A was positioned by Bruchpilot and Rim-binding protein complexes at maturing AZs. Unc13B localized 120 nm away from Ca2+ channels, whereas Unc13A localized only 70 nm away and was responsible for docking SVs at this distance. Unc13Anull mutants suffered from inefficient, delayed and EGTA-supersensitive release. Mathematical modeling suggested that synapses normally operate via two independent release pathways differentially positioned by either isoform. We identified isoform-specific Unc13-AZ scaffold interactions regulating SV-Ca2+-channel topology whose developmental tightening optimizes synaptic transmission.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Jahn, R. & Fasshauer, D. Molecular machines governing exocytosis of synaptic vesicles. Nature 490, 201–207 (2012).
Imig, C. et al. The morphological and molecular nature of synaptic vesicle priming at presynaptic active zones. Neuron 84, 416–431 (2014).
Walter, A.M., Wiederhold, K., Bruns, D., Fasshauer, D. & Sørensen, J.B. Synaptobrevin N-terminally bound to syntaxin-SNAP-25 defines the primed vesicle state in regulated exocytosis. J. Cell Biol. 188, 401–413 (2010).
Ma, C., Li, W., Xu, Y. & Rizo, J. Munc13 mediates the transition from the closed syntaxin-Munc18 complex to the SNARE complex. Nat. Struct. Mol. Biol. 18, 542–549 (2011).
Man, K.N. et al. Identification of a Munc13-sensitive step in chromaffin cell large dense-core vesicle exocytosis. eLife 4, e10635 (2015).
Varoqueaux, F. et al. Total arrest of spontaneous and evoked synaptic transmission but normal synaptogenesis in the absence of Munc13-mediated vesicle priming. Proc. Natl. Acad. Sci. USA 99, 9037–9042 (2002).
Lipstein, N. et al. Dynamic control of synaptic vesicle replenishment and short-term plasticity by Ca(2+)-calmodulin-Munc13-1 signaling. Neuron 79, 82–96 (2013).
Eggermann, E., Bucurenciu, I., Goswami, S.P. & Jonas, P. Nanodomain coupling between Ca2+ channels and sensors of exocytosis at fast mammalian synapses. Nat. Rev. Neurosci. 13, 7–21 (2011).
Wadel, K., Neher, E. & Sakaba, T. The coupling between synaptic vesicles and Ca2+ channels determines fast neurotransmitter release. Neuron 53, 563–575 (2007).
Young, S.M. Jr. & Neher, E. Synaptotagmin has an essential function in synaptic vesicle positioning for synchronous release in addition to its role as a calcium sensor. Neuron 63, 482–496 (2009).
Südhof, T.C. The presynaptic active zone. Neuron 75, 11–25 (2012).
Acuna, C., Liu, X., Gonzalez, A. & Südhof, T.C. RIM-BPs mediate tight coupling of action potentials to Ca(2+)-triggered neurotransmitter release. Neuron 87, 1234–1247 (2015).
Kaeser, P.S. et al. RIM proteins tether Ca2+ channels to presynaptic active zones via a direct PDZ-domain interaction. Cell 144, 282–295 (2011).
Kittel, R.J. et al. Bruchpilot promotes active zone assembly, Ca2+ channel clustering, and vesicle release. Science 312, 1051–1054 (2006).
Liu, K.S. et al. RIM-binding protein, a central part of the active zone, is essential for neurotransmitter release. Science 334, 1565–1569 (2011).
Matz, J., Gilyan, A., Kolar, A., McCarvill, T. & Krueger, S.R. Rapid structural alterations of the active zone lead to sustained changes in neurotransmitter release. Proc. Natl. Acad. Sci. USA 107, 8836–8841 (2010).
Han, Y., Kaeser, P.S., Südhof, T.C. & Schneggenburger, R. RIM determines Ca2+ channel density and vesicle docking at the presynaptic active zone. Neuron 69, 304–316 (2011).
Fouquet, W. et al. Maturation of active zone assembly by Drosophila Bruchpilot. J. Cell Biol. 186, 129–145 (2009).
Spangler, S.A. et al. Liprin-α2 promotes the presynaptic recruitment and turnover of RIM1/CASK to facilitate synaptic transmission. J. Cell Biol. 201, 915–928 (2013).
Zhen, M. & Jin, Y. The liprin protein SYD-2 regulates the differentiation of presynaptic termini in C. elegans. Nature 401, 371–375 (1999).
Owald, D. et al. A Syd-1 homologue regulates pre- and postsynaptic maturation in Drosophila. J. Cell Biol. 188, 565–579 (2010).
Matkovic, T. et al. The Bruchpilot cytomatrix determines the size of the readily releasable pool of synaptic vesicles. J. Cell Biol. 202, 667–683 (2013).
Aravamudan, B., Fergestad, T., Davis, W.S., Rodesch, C.K. & Broadie, K. Drosophila UNC-13 is essential for synaptic transmission. Nat. Neurosci. 2, 965–971 (1999).
Basu, J., Betz, A., Brose, N. & Rosenmund, C. Munc13-1 C1 domain activation lowers the energy barrier for synaptic vesicle fusion. J. Neurosci. 27, 1200–1210 (2007).
Schotten, S. et al. Additive effects on the energy barrier for synaptic vesicle fusion cause supralinear effects on the vesicle fusion rate. eLife 4, e05531 (2015).
Shin, O.H. et al. Munc13 C2B domain is an activity-dependent Ca2+ regulator of synaptic exocytosis. Nat. Struct. Mol. Biol. 17, 280–288 (2010).
Basu, J. et al. A minimal domain responsible for Munc13 activity. Nat. Struct. Mol. Biol. 12, 1017–1018 (2005).
Stevens, D.R. et al. Identification of the minimal protein domain required for priming activity of Munc13-1. Curr. Biol. 15, 2243–2248 (2005).
Venken, K.J., He, Y., Hoskins, R.A. & Bellen, H.J. P[acman]: a BAC transgenic platform for targeted insertion of large DNA fragments in D. melanogaster. Science 314, 1747–1751 (2006).
Owald, D. et al. Cooperation of Syd-1 with Neurexin synchronizes pre- with postsynaptic assembly. Nat. Neurosci. 15, 1219–1226 (2012).
Graf, E.R., Daniels, R.W., Burgess, R.W., Schwarz, T.L. & DiAntonio, A. Rab3 dynamically controls protein composition at active zones. Neuron 64, 663–677 (2009).
Kawasaki, F., Zou, B., Xu, X. & Ordway, R.W. Active zone localization of presynaptic calcium channels encoded by the cacophony locus of Drosophila. J. Neurosci. 24, 282–285 (2004).
Weimer, R.M. et al. UNC-13 and UNC-10/rim localize synaptic vesicles to specific membrane domains. J. Neurosci. 26, 8040–8047 (2006).
Augustin, I., Rosenmund, C., Südhof, T.C. & Brose, N. Munc13-1 is essential for fusion competence of glutamatergic synaptic vesicles. Nature 400, 457–461 (1999).
Fernández-Chacón, R. et al. Synaptotagmin I functions as a calcium regulator of release probability. Nature 410, 41–49 (2001).
Meinrenken, C.J., Borst, J.G. & Sakmann, B. Calcium secretion coupling at calyx of Held governed by nonuniform channel-vesicle topography. J. Neurosci. 22, 1648–1667 (2002).
Naraghi, M. & Neher, E. Linearized buffered Ca2+ diffusion in microdomains and its implications for calculation of [Ca2+] at the mouth of a calcium channel. J. Neurosci. 17, 6961–6973 (1997).
Lou, X., Scheuss, V. & Schneggenburger, R. Allosteric modulation of the presynaptic Ca2+ sensor for vesicle fusion. Nature 435, 497–501 (2005).
Kaufmann, N., DeProto, J., Ranjan, R., Wan, H. & Van Vactor, D. Drosophila liprin-alpha and the receptor phosphatase Dlar control synapse morphogenesis. Neuron 34, 27–38 (2002).
Hallermann, S. et al. Naked dense bodies provoke depression. J. Neurosci. 30, 14340–14345 (2010).
Regehr, W.G. Short-term presynaptic plasticity. Cold Spring Harb. Perspect. Biol. 4, a005702 (2012).
Sugita, S., Shin, O.H., Han, W., Lao, Y. & Südhof, T.C. Synaptotagmins form a hierarchy of exocytotic Ca(2+) sensors with distinct Ca(2+) affinities. EMBO J. 21, 270–280 (2002).
Mohrmann, R. et al. Synaptotagmin interaction with SNAP-25 governs vesicle docking, priming, and fusion triggering. J. Neurosci. 33, 14417–14430 (2013).
Hu, Z., Tong, X.J. & Kaplan, J.M. UNC-13L, UNC-13S, and Tomosyn form a protein code for fast and slow neurotransmitter release in Caenorhabditis elegans. eLife 2, e00967 (2013).
Zhou, K., Stawicki, T.M., Goncharov, A. & Jin, Y. Position of UNC-13 in the active zone regulates synaptic vesicle release probability and release kinetics. eLife 2, e01180 (2013).
Rosenmund, C. et al. Differential control of vesicle priming and short-term plasticity by Munc13 isoforms. Neuron 33, 411–424 (2002).
Chen, Z., Das, B., Nakamura, Y., DiGregorio, D.A. & Young, S.M. Jr. Ca2+ channel to synaptic vesicle distance accounts for the readily releasable pool kinetics at a functionally mature auditory synapse. J. Neurosci. 35, 2083–2100 (2015).
Keller, D. et al. An exclusion zone for Ca2+ channels around docked vesicles explains release control by multiple channels at a CNS synapse. PLoS Comput. Biol. 11, e1004253 (2015).
Nakamura, Y. et al. Nanoscale distribution of presynaptic Ca(2+) channels and its impact on vesicular release during development. Neuron 85, 145–158 (2015).
Wang, L.Y., Neher, E. & Taschenberger, H. Synaptic vesicles in mature calyx of Held synapses sense higher nanodomain calcium concentrations during action potential-evoked glutamate release. J. Neurosci. 28, 14450–14458 (2008).
Sigrist, S.J., Reiff, D.F., Thiel, P.R., Steinert, J.R. & Schuster, C.M. Experience-dependent strengthening of Drosophila neuromuscular junctions. J. Neurosci. 23, 6546–6556 (2003).
Aberle, H. et al. wishful thinking encodes a BMP type II receptor that regulates synaptic growth in Drosophila. Neuron 33, 545–558 (2002).
Wagh, D.A. et al. Bruchpilot, a protein with homology to ELKS/CAST, is required for structural integrity and function of synaptic active zones in Drosophila. Neuron 49, 833–844 (2006).
Ullrich, A. et al. Dynamical organization of syntaxin-1a at the presynaptic active zone. PLoS Comput. Biol. 11, e1004407 (2015).
Andlauer, T.F. & Sigrist, S.J. Quantitative analysis of Drosophila larval neuromuscular junction morphology. Cold Spring Harb. Protoc. 2012, 490–493 (2012).
Göttfert, F. et al. Coaligned dual-channel STED nanoscopy and molecular diffusion analysis at 20 nm resolution. Biophys. J. 105, L01–L03 (2013).
Füger, P., Behrends, L.B., Mertel, S., Sigrist, S.J. & Rasse, T.M. Live imaging of synapse development and measuring protein dynamics using two-color fluorescence recovery after photo-bleaching at Drosophila synapses. Nat. Protoc. 2, 3285–3298 (2007).
Rasse, T.M. et al. Glutamate receptor dynamics organizing synapse formation in vivo. Nat. Neurosci. 8, 898–905 (2005).
Qin, G. et al. Four different subunits are essential for expressing the synaptic glutamate receptor at neuromuscular junctions of Drosophila. J. Neurosci. 25, 3209–3218 (2005).
Stewart, B.A., Atwood, H.L., Renger, J.J., Wang, J. & Wu, C.F. Improved stability of Drosophila larval neuromuscular preparations in haemolymph-like physiological solutions. J. Comp. Physiol. 175, 179–191 (1994).
Worseck, J.M., Grossmann, A., Weimann, M., Hegele, A. & Stelzl, U. A stringent yeast two-hybrid matrix screening approach for protein-protein interaction discovery. Methods Mol. Biol. 812, 63–87 (2012).
Hegele, A. et al. Dynamic protein-protein interaction wiring of the human spliceosome. Mol. Cell 45, 567–580 (2012).
Matveev, V., Sherman, A. & Zucker, R.S. New and corrected simulations of synaptic facilitation. Biophys. J. 83, 1368–1373 (2002).
Allbritton, N.L., Meyer, T. & Stryer, L. Range of messenger action of calcium ion and inositol 1,4,5-trisphosphate. Science 258, 1812–1815 (1992).
Helmchen, F., Borst, J.G. & Sakmann, B. Calcium dynamics associated with a single action potential in a CNS presynaptic terminal. Biophys. J. 72, 1458–1471 (1997).
Xu, T., Naraghi, M., Kang, H. & Neher, E. Kinetic studies of Ca2+ binding and Ca2+ clearance in the cytosol of adrenal chromaffin cells. Biophys. J. 73, 532–545 (1997).
Nägerl, U.V., Novo, D., Mody, I. & Vergara, J.L. Binding kinetics of calbindin-D(28k) determined by flash photolysis of caged Ca(2+). Biophys. J. 79, 3009–3018 (2000).
Klingauf, J. & Neher, E. Modeling buffered Ca2+ diffusion near the membrane: implications for secretion in neuroendocrine cells. Biophys. J. 72, 674–690 (1997).
Neher, E. & Sakaba, T. Combining deconvolution and noise analysis for the estimation of transmitter release rates at the calyx of held. J. Neurosci. 21, 444–461 (2001).
Acknowledgements
This work was supported by grants from the Deutsche Forschungsgemeinschaft (DFG) to S.J.S. (Exc 257, TP A3 and A6 SFB 958, TP B9/SFB665; TP09/SFB740), and J.H.D., M.C.W. and B.L. (TP A6/SFB958). A.M.W. was supported by a DFG Emmy Noether grant. M.A.B. was supported by a PhD fellowship from the graduate school GRK 1123 funded by the DFG and by the SFB 740. A.T.G. was supported by a NeuroCure PhD fellowship within the International Graduate Program Medical Neurosciences funded by the DFG (Exc 257). We thank M. Brünner and A. Stawrakakis for excellent technical assistance, S. Sweeney for help with the EMS screen and T. Matkovic for help with cloning.
Author information
Authors and Affiliations
Contributions
M.A.B., S.R.-A., A.M.W. and S.J.S. conceived the project. M.A.B. and M.M.M. performed all confocal and in vivo imaging experiments and analyzed the data. M.A.B. performed STED experiments and analyzed the data. C.B., S.R.-A. and E.R. performed all electrophysiological experiments, and C.B. and E.R. analyzed the data. J.L., U.S. and J.H.D. performed and analyzed Y2H experiments. M.C.W. and B.L. contributed to the Y2H experiments and analysis. C.B. and D.D.B. performed HPF experiments and C.B. analyzed the data. A.T.G. performed modeling and simulation. H.B. and M.M.M. created antibodies and isoform-specific mutants. I.M.R. and C.J.O. performed the EMS screen. F.G. and S.W.H. developed and built the STED microscope. M.A.B., A.M.W. and S.J.S. wrote the paper with input from all coauthors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Liprin-ɑ/Syd-1 scaffold complexes organize the AZ localization of Unc13B
(a,b) Muscle 4 NMJs of segments A2-A4 from 3rd instar larvae of the displayed genotypes labelled with the antibodies (ABs) indicated. (c) Mean BRP intensity measured over the whole NMJ was unchanged in Syd-1null and Liprin-αnull in comparison to the Wild type (Wild type (n=13 NMJs from 4 larvae) vs. Syd-1null (n=11 NMJs from 4 larvae) vs. Liprin-αnull (n=12 NMJs from 4 larvae): p >0.05 for Wild type vs Syd-1null; p >0.05 for Wild type vs. Liprin-αnull; p >0.05 for Syd-1null vs. Liprin-αnull; p=0.2883 (F(2,33)=1.29)). (d) BRP spots per µm² NMJ were slightly reduced in Syd-1null and significantly reduced in Liprin-αnull in comparison to the Wild type (Wild type (n=13 NMJs from 4 larvae) vs. Syd-1null (n=11 NMJs from 4 larvae) vs. Liprin-αnull (n=12 NMJs from 4 larvae): p >0.05 for Wild type vs Syd-1null; p ≤0.01 for Wild type vs. Liprin-αnull; p >0.05 for Syd-1null vs. Liprin-αnull; p=0.0108 (F(2,33)=5.207)). (e) Mean Unc13A intensity measured over the whole NMJ was unchanged in Syd-1null and Liprin-αnull in comparison to the Wild type (Wild type (n=13 NMJs from 4 larvae) vs. Syd-1null (n=11 NMJs from 4 larvae) vs. Liprin-αnull (n=12 NMJs from 4 larvae): p >0.05 for Wild type vs Syd-1null; p >0.05 for Wild type vs. Liprin-α null; p >0.05 for Syd-1null vs. Liprin-αnull; p=0.2105 (F(2,33)=1.63)). (f) Unc13A spots per µm² NMJ were slightly reduced in Syd-1null and Liprin-αnull in comparison to the Wild type (Wild type (n=13 NMJs from 4 larvae) vs. Syd-1null (n=11 NMJs from 4 larvae) vs. Liprin-αnull (n=12 NMJs from 4 larvae): p ≤0.05 for Wild type vs Syd-1null; p >0.05 for Wild type vs. Liprin-αnull; p >0.05 for Syd-1null vs. Liprin-αnull; p=0.0278 (F(2,33)=4.00)). (g,h) Muscle 4 NMJs of segments A2-A4 from 3rd instar larvae of the displayed genotypes labelled with the ABs indicated. (i) Mean Unc13B intensity measured over the whole NMJ was slightly reduced in Syd-1null but severely reduced in Liprin-αnull in comparison to the Wild type (Wild type (n=13 NMJs from 5 larvae) vs. Syd-1null (n=11 NMJs from 5 larvae) vs. Liprin-αnull (n=15 NMJs from 5 larvae): p >0.05 for Wild type vs Syd-1null; p ≤0.001 for Wild type vs. Liprin-αnull; p ≤0.01 for Syd-1null vs. Liprin-αnull; p <0.0001 (F(2,36)=15.13)). (j) Unc13B spots per µm² NMJ were significantly reduced in Syd-1null but severely reduced in Liprin-αnull in comparison to the Wild type (Wild type (n=13 NMJs from 5 larvae) vs. Syd-1null (n=11 NMJs from 5 larvae) vs. Liprin-αnull (n=15 NMJs from 5 larvae): p ≤0.01 for Wild type vs Syd-1null; p ≤0.001 for Wild type vs. Liprin-αnull; p ≤0.001 for Syd-1null vs. Liprin-αnull; p <0.0001 (F(2,36)=42.93)). Number and p values are listed in Supplementary Table 1. Statistics for c-f, i, j: nonparametric one-way analysis of variance (ANOVA) test, followed by a Turkey’s multiple comparison test. All panels show mean ± SEM; *, p ≤0.05; **, p ≤0.01; ***, p ≤0.001; ns, not significant, p >0.05. Scale bars: 5 μm.
Supplementary Figure 2 Unc13B interacts with Syd-1/Liprin-α; Unc13A interacts with BRP/RBP
(a) Schematic representation of Unc13B N-terminus including three fragments (1-3) that were used in the Y2H screen; Liprin-α domain structure containing three C-terminal SAM domains (I-III from the N terminus); Syd-1 domain structure containing an N-terminal PDZ domain, a C2 and a Rho-GAP domain. The corresponding fragments of each protein used in the Y2H screen are indicated. A central N-terminal fragment of Unc13B interacted with an N-terminal part of Liprin-α. Both very N-terminal fragments of Unc13B interacted with a central stretch of Syd-1 located in-between PDZ- and C2-domain. (b) Schematic representation of Unc13A N-terminus including three fragments (1-3) that were used in the Y2H screen. The RBP-binding PxxP motif is indicated; RBP domain structure containing three SH3 domains (I-III from the N terminus) and three Fibronectin 3 (FN3) domains; BRP domain structure containing several coiled-coil (CC) domains. The corresponding fragments of each protein used in the Y2H screen are indicated. The most N-terminal fragment of Unc13A (including the RBP binding PxxP motif) interacted with both C-terminal fragments of RBP including the SH3-domains II and III, and with an N-terminal part of BRP.
Supplementary Figure 3 BRP/RBP scaffold complexes organize the AZ-localization of Unc13A
(a-c) Muscle 4 NMJs of segments A2-A4 from 3rd instar larvae of the displayed genotypes labelled with the ABs indicated. BRP as well as Unc13A were severely decreased upon motoneuronal downregulation of BRP or in Rbpnull mutants. (d,e) BRP as well as Unc13A intensity were severely decreased upon motoneuronal downregulation of BRP or in Rbpnull mutants with the strongest downregulation upon Brp knockdown in Rbpnull (BRP intensity: Ctrl (n=14 NMJs from 5 larvae) vs. BrpRNAi (n=15 NMJs from 5 larvae) vs. Rbpnull (n=15 NMJs from 5 larvae) vs. BrpRNAi;RBPnull (n=13 NMJs from 5 larvae): p ≤0.001 for Ctrl vs BrpRNAi; p ≤0.001 for Ctrl vs. Rbpnull; p ≤0.001 for Ctrl vs. BrpRNAi;RBPnull; p ≤0.05 for BrpRNAi vs. Rbpnull; p >0.05 for BrpRNAi vs. BrpRNAi;RBPnull; p ≤0.05 for Rbpnull vs. BrpRNAi;RBPnull; p <0.0001 (F(3,53)=31.96)); Unc13A intensity: Ctrl (n=14 NMJs from 5 larvae) vs. BrpRNAi (n=15 NMJs from 5 larvae) vs. Rbpnull (n=15 NMJs from 5 larvae) vs. BrpRNAi;RBPnull (n=13 NMJs from 5 larvae): p ≤0.001 for Ctrl vs BrpRNAi; p ≤0.001 for Ctrl vs. Rbpnull; p ≤0.001 for Ctrl vs. BrpRNAi;RBPnull; p >0.05 for BrpRNAi vs. Rbpnull; p >0.05 for BrpRNAi vs. BrpRNAi;RBPnull; p >0.05 for Rbpnull vs. BrpRNAi;RBPnull; p <0.0001 (F(3,53)=30.07)). (f) Unc13A spots per µm² NMJ were decreased upon motoneuronal downregulation of BRP or in Rbpnull mutants with the strongest downregulation upon Brp knockdown in Rbpnull (Ctrl (n=14 NMJs from 5 larvae) vs. BrpRNAi (n=15 NMJs from 5 larvae) vs. Rbpnull (n=15 NMJs from 5 larvae) vs. BrpRNAi;RBPnull (n=13 NMJs from 5 larvae): p ≤0.001 for Ctrl vs BrpRNAi; p ≤0.001 for Ctrl vs. Rbpnull; p ≤0.001 for Ctrl vs. BrpRNAi;RBPnull; p ≤0.01 for BrpRNAi vs. Rbpnull; p >0.05 for BrpRNAi vs. BrpRNAi;RBPnull; p ≤0.001 for Rbpnull vs. BrpRNAi;RBPnull; p <0.0001 (F(3,53)=63.28)). (g-i) Muscle 4 NMJs of segments A2-A4 from 3rd instar larvae of the genotypes indicated labelled with the ABs indicated. BRP but not Unc13B were severely decreased upon motoneuronal downregulation of BRP as well as in Rbpnull mutants. (j) Mean Unc13B intensity measured over the whole NMJ was not affected upon downregulation of BRP or in Rbpnull mutants or even upon Brp knockdown in Rbpnull (Ctrl (n=12 NMJs from 5 larvae) vs. BrpRNAi (n=11 NMJs from 5 larvae) vs. Rbpnull (n=13 NMJs from 5 larvae) vs. BrpRNAi;RBPnull (n=10 NMJs from 5 larvae): p >0.05 for Ctrl vs BrpRNAi; p >0.05 for Ctrl vs. Rbpnull; p >0.05 for Ctrl vs. BrpRNAi;RBPnull; p >0.05 for BrpRNAi vs. Rbpnull; p >0.05 for BrpRNAi vs. BrpRNAi;RBPnull; p >0.05 for Rbpnull vs. BrpRNAi;RBPnull; p=0.3491 (F(3,42)=1.127)). (k) Unc13B spots per µm² NMJ were unchanged upon downregulation of BRP or in Rbpnull mutants or even upon Brp knockdown in Rbpnull (Ctrl (n=12 NMJs from 5 larvae) vs. BrpRNAi (n=11 NMJs from 5 larvae) vs. Rbpnull (n=13 NMJs from 5 larvae) vs. BrpRNAi;RBPnull (n=10 NMJs from 5 larvae): p >0.05 for Ctrl vs BrpRNAi; p >0.05 for Ctrl vs. Rbpnull; p >0.05 for Ctrl vs. BrpRNAi;RBPnull; p >0.05 for BrpRNAi vs. Rbpnull; p >0.05 for BrpRNAi vs. BrpRNAi;RBPnull; p >0.05 for Rbpnull vs. BrpRNAi;RBPnull; p=0.6570 (F(3,42)=0.5408)). Number and p values are listed in Supplementary Table 1. Statistics for d-f, j,k: nonparametric one-way analysis of variance (ANOVA) test, followed by a Turkey’s multiple comparison test. All panels show mean ± SEM; *, p ≤0.05; **, p ≤0.01; ***, p ≤0.001; ns, not significant, p >0.05. Scale bars: 5 μm.
Supplementary Figure 4 TEVC analysis of Unc13A and Unc13B mutant terminals
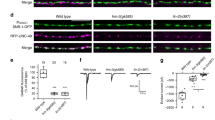
(a,b) The time-to-peak (time difference between stimulation pulse to the afferent nerve and the eEJC minimum) and eEJC decay, which is the time constant τ resulting from a single exponential fit in the range from 60% of the eEJC amplitude back to baseline, both are similar in Ctrl (black) and Unc13Bnull (blue) (time to peak: Ctrl (n=12 NMJs from 12 larvae) vs Unc13Bnull (n=12 NMJs from 12 larvae), p=0.1333 (t(22)=1.559); eEJC decay: Ctrl (n=12 NMJs from 12 larvae) vs Unc13Bnull (n=11 NMJs from 11 larvae), p=0.2413 (t(21)=1.206)). (c) The time to peak is significantly prolonged in Unc13Anull mutant synapses (Wild type (n=12 NMJs from 12 larvae) vs Unc13Anull (n=12 NMJs from 12 larvae), p=0.0162 (t(22)=2.605)). (d) The eEJC decay is similar in Wild type and Unc13Anull (Wild type (n=12 NMJs from 12 larvae) vs Unc13Anull (n=9 NMJs from 9 larvae), p=0.2136 (U=36)). (e) The mEJC rise time is unaltered in Unc13Bnull mutant synapses compared to Ctrl (Ctrl (n=14 NMJs from 10 larvae) vs Unc13Bnull (n=7 NMJs from 5 larvae), p=0.7652 (U=44.5)). (f) In Unc13Bnull the mEJC decay is significantly increased compared to Ctrl (Ctrl (n=14 NMJs from 10 larvae) vs Unc13Bnull (n=7 NMJs from 5 larvae), p=0.0480 (U=22.00)). (g,h) mEJC kinetics do not differ between Wild type and Unc13Anull (mEJC rise time: Wild type (n=12 NMJs from 6 larvae) vs Unc13Anull (n=11 NMJs from 6 larvae), p=0.1914 (t(21)=1.350); mEJC decay: Wild type (n=12 NMJs from 6 larvae) vs Unc13Anull (n=11 NMJs from 6 larvae), p=0.2546 (t(21)=1.171)). All recordings were performed in the presence of 1.5 mM extracellular Ca2+. Number and p values are listed in Supplementary Table 1. Statistics: Student's t-test besides panels d,e,f where a Mann Whitney U-test was performed. All panels show mean ± SEM; *, p ≤0.05; **, p ≤0.01; ***, p ≤0.001; ns, not significant, p >0.05.
Supplementary Figure 5 Ca2+ sensitivity and release probability is altered upon loss of Unc13A but not -B
(a-f) Plot of eEJC amplitude as a function of extracellular Ca2+ concentrations [Ca2+]ext fitted with Hill equations to determine the values for slope and KD. A clear shift can be observed in (a) Unc13Anull mutant synapses (red) compared to Wild type (black), whereas in (d) there is no change upon loss of Unc13Bnull (blue) compared to Ctrl (black) (a: Wild type (n=12 NMJs from 12 larvae per Ca2+ concentration) vs Unc13Anull (n=10 NMJs from 10 larvae per Ca2+ concentration): 0.75 mM [Ca2+]ext: p=0.0092 (U=20); 1.5 mM [Ca2+]ext: p <0.0001 (U=0); 3 mM [Ca2+]ext: p=0.0005 (U=7); 6 mM [Ca2+]ext: p=0.0272 (U=26); 10 mM [Ca2+]ext: p=0.0062 (U=18)); d: Ctrl (n=12 NMJs from 12 larvae per Ca2+ concentration) vs Unc13Bnull (n=12 NMJs from 12 larvae per Ca2+ concentration): 0.75 mM [Ca2+]ext: p=0.1971 (t(22)=1.330); 1.5 mM [Ca2+]ext: p=0.2652 (t(22)=1.143); 3 mM [Ca2+]ext: p=0.9269 (t(22)=0.09278); 6 mM [Ca2+]ext: p=0.5181 (t(22)=0.6569); 10 mM [Ca2+]ext: p=0.6284 (t(22)=0.4908)). The values for I/Imax can be found in Supplementary Table 1. (b) Ca2+-dependence of release analysis revealed an increased Ca2+ requirement (KD, obtained from fitting with the Hill function) in Unc13Anull mutant synapses (Wild type (n=12 NMJs from 12 larvae) vs Unc13Anull (n=10 NMJs from 10 larvae), p=0.0004 (U=6)). (c) The apparent Ca2+ cooperativity of release (slope, obtained from fitting with the Hill function) is not different in Unc13Anull relative to Wild type (Wild type (n=12 NMJs from 12 larvae) vs Unc13Anull (n=10 NMJs from 10 larvae), p=0.6682 (U=53)). (e,f) The Ca2+-dependence and Ca2+-cooperativity of release are both unaltered upon loss of Unc13B (KD: Ctrl (n=12 NMJs from 12 larvae) vs Unc13Bnull (n=12 NMJs from 12 larvae), p=0.9566 (t(22)=0.05502); slope: Ctrl (n=12 NMJs from 12 larvae) vs Unc13Bnull (n=12 NMJs from 12 larvae), p=0.1574 (t(22)=1.464)). (g) Sample traces of paired pulse stimulation for Ctrl (black) and Unc13Bnull (blue) at 10 ms ISI show no differences between genotypes. (h) The paired pulse ratios were not significantly changed in Unc13Bnull at 10 ms ISI, in all Ca2+ concentrations (Ctrl (n=12 NMJs from 12 larvae per Ca2+ concentration) vs Unc13Bnull (n=12 NMJs from 12 larvae per Ca2+ concentration): 0.75 mM [Ca2+]ext: p=0.1971 (t(22)=1.33); 1.5 mM [Ca2+]ext: p=0.1678 (t(22)=1.426); 3 mM [Ca2+]ext: p=0.474 (t(22)=0.7284); 6 mM [Ca2+]ext: p=0.3726 (t(22)=0.9102); 10 mM [Ca2+]ext: p=0.2602 (t(22)=1.156)). Values can be found in Supplementary Table 1. (i) Unc13Anull (gold: control with DMSO, red: with EGTA-AM/DMSO) shows faster and stronger inhibition of eEJC amplitudes after addition of 200 µM EGTA-AM to the extra-cellular solution compared to Wild type (black: control with DMSO, blue: with EGTA-AM/DMSO). Amplitudes are normalized to average eEJC amplitudes obtained during 1 min of baseline recording prior to the addition of EGTA-AM/DMSO or DMSO, each with Pluronic F-127. Synaptic transmission was stimulated by single action potentials every 10 s. Experiments were performed in the presence of 2.5 mM extracellular Ca2+. Values can be found in Supplementary Table 1. (j) The time constant of the inhibition caused by EGTA-AM application was determined by fitting a single exponential decay function to 100 peak amplitude values after addition of EGTA-AM in individual cells. This revealed a significantly faster inhibition in Unc13Anull compared to Wild type animals (Wild type + EGTA (n=10 NMJs from 10 larvae) vs Unc13Anull + EGTA (n=10 NMJs from 10 larvae), p=0.0012 (t(18)=3.835)). (k) The asymptotic inhibition is captured in the exponential fit as the plateau value which was significantly decreased in Unc13Anull in comparison to Wild type (Wild type + EGTA (n=10 NMJs from 10 larvae) vs Unc13Anull + EGTA (n=10 NMJs from 10 larvae), p=0.016 (t(18)=2.6508)). (l) 30 min incubation with the fast Ca2+-buffer Bapta-AM reduced eEJC amplitudes in both genotypes to a similar extent. Sample traces for Wild type (black with DMSO, blue with Bapta-AM/DMSO) and Unc13Anull (gold with DMSO, red with Bapta-AM/DMSO) exhibit similar Bapta-sensitivity for both genotypes. For clarity, the stimulation artefact was removed and replaced by a straight line. (m,n) The significant reduction of the eEJC amplitude after 30 min Bapta-AM incubation is similar in Wild type (m) and Unc13Anull (n) compared to DMSO incubated cells (m: Wild type + DMSO (n=15 NMJs from 9 larvae) vs Wild type + Bapta (n=14 NMJs from 9 larvae), p <0.0001 (t(27)=12.59); n: Unc13Anull + DMSO (n=14 NMJs from 10 larvae) vs Unc13Anull + Bapta (n=14 NMJs from 8 larvae), p=0.0004 (t(26)=4.095)). Values can be found in Supplementary Table 1. (o) The Bapta sensitivity is calculated as the ratio of eEJC amplitude size in the presence of Bapta-AM/DMSO to the eEJC amplitude size in the presence of DMSO. The Bapta-sensitivity does not differ between Wild type and Unc13Anull (Bapta sensitivity: Wild type (n=14 NMJs from 9 larvae) vs Unc13Anull (n=14 NMJs from 8 larvae), p=0.304 (t(26)=1.049)). Values can be found in Supplementary Table 1. Statistics: Student's t-test except for panels (a-c) where a Mann-Whitney U-test was performed. All panels show mean ± SEM; *, p ≤0.05; **, p ≤0.01; ***, p ≤0.001; ns, not significant, p >0.05.
Supplementary Figure 6 Increased Ca2+ channel abundance at Unc13Anull mutant AZs
(a) Two-color STED images of multiple AZs from 3rd instar larvae of the displayed genotypes labelled with the indicated ABs. BRP rings were larger in Unc13Anull. (b) BRP ring diameters were increased in Unc13Anull in comparison to the Wild type (Wild type (n=9 NMJs from 3 larvae) vs Unc13Anull (n=12 NMJs from 3 larvae), p=0.0001 (U=0)). (c) Two-color STED images of multiple AZs from 3rd instar larvae of the displayed genotypes labelled with the indicated ABs. BRP ring structure appeared normal in Unc13Bnull (d) BRP ring diameters were unchanged in Unc13Bnull in comparison to Ctrl (Ctrl (n=8 NMJs from 3 larvae) vs Unc13Bnull (n=8 NMJs from 3 larvae), p=0.9591 (U=31)). (e) Muscle 4 NMJs of segments A2-A4 from 3rd instar larvae of the displayed genotypes labelled with the ABs indicated. (f) Cac-GFP spot sizes were increased in Unc13Anull but not Unc13Bnull in comparison to Ctrl (Ctrl (n=19 NMJs from 5 larvae) vs. Unc13Anull (n=15 NMJs from 5 larvae) vs. Unc13Bnull (n=21 NMJs from 5 larvae): p >0.05 for Ctrl vs Unc13Bnull; p ≤0.001 for Ctrl vs. Unc13Anull; p ≤0.001 for Unc13Bnull vs. Unc13Anull; p <0.0001 (F(2,52)=54.12)). (g) Mean Cac-GFP intensity measured over the whole NMJ was similar in Unc13Anull, Unc13Bnull and Ctrl (Ctrl (n=19 NMJs from 5 larvae) vs. Unc13Anull (n=15 NMJs from 5 larvae) vs. Unc13Bnull (n=21 NMJs from 5 larvae): p >0.05 for Ctrl vs Unc13Bnull; p >0.05 for Ctrl vs. Unc13Anull; p >0.05 for Unc13Bnull vs. Unc13Anull; p=0.166 (F(2,52)=1.855)). Number and p values are listed in Supplementary Table 1. Statistics: Mann-Whitney U-test, except for f and g where a nonparametric one-way analysis of variance (ANOVA) test, followed by a Turkey’s multiple comparison test was performed. All panels show mean ± SEM; *, p ≤0.05; **, p ≤0.01; ***, p ≤0.001; ns, not significant, p >0.05. Scale bar: (a,c) 500 nm; (e) 5 μm.
Supplementary Figure 7 Sketch of de novo synapse formation
During the process of AZ assembly, clusters of Syd-1 and Liprin-α undergo rounds of assembly and disassembly at the presynaptic membrane. Unc13B localizes to sites of de novo synapse formation via the Syd-1/Liprin-α scaffold. At nascent synapses, this induces a loose SV-Ca2+ channel coupling. Later during the AZ maturation process, Unc13A localizes to more mature synapses via a second, central RBP/BRP scaffold that concentrates Unc13A at the center of the AZ. Unc13A facilitates a close localization of SVs to the presynaptic Ca2+ channels and therefore maintains a tight stimulus/secretion coupling.
Supplementary Figure 8 Allosteric five-site binding model of Ca2+-driven exocytosis
Reaction scheme (derived from the ‘allosteric model’; Lou et al., 2005. Nature. 435:497-501) depicts the sequential binding of up to five Ca2+ ions to a single vesicle (RCa0-5).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1 and 2 (PDF 2087 kb)
Rights and permissions
About this article
Cite this article
Böhme, M., Beis, C., Reddy-Alla, S. et al. Active zone scaffolds differentially accumulate Unc13 isoforms to tune Ca2+ channel–vesicle coupling. Nat Neurosci 19, 1311–1320 (2016). https://doi.org/10.1038/nn.4364
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4364
This article is cited by
-
UNC13A Gene Brings New Hope for ALS Disease-Modifying Drugs
Neuroscience Bulletin (2022)
-
TDP-43 represses cryptic exon inclusion in the FTD–ALS gene UNC13A
Nature (2022)
-
Unc13A and Unc13B contribute to the decoding of distinct sensory information in Drosophila
Nature Communications (2021)
-
Advanced imaging and labelling methods to decipher brain cell organization and function
Nature Reviews Neuroscience (2021)
-
PKC-phosphorylation of Liprin-α3 triggers phase separation and controls presynaptic active zone structure
Nature Communications (2021)