Abstract
Models of predictive coding frame perception as a generative process in which expectations constrain sensory representations. These models account for expectations about how a stimulus will move or change from moment to moment, but do not address expectations about what other, distinct stimuli are likely to appear based on prior experience. We show that such memory-based expectations in human visual cortex are related to the hippocampal mechanism of pattern completion.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rao, R.P.N. & Ballard, D.H. Nat. Neurosci. 2, 79–87 (1999).
Friston, K. Phil. Trans. R. Soc. Lond. B 360, 815–836 (2005).
Clark, A. Behav. Brain Sci. 36, 181–204 (2013).
Smith, F.W. & Muckli, L. Proc. Natl. Acad. Sci. USA 107, 20099–20103 (2010).
Alink, A., Schwiedrzik, C.M., Kohler, A., Singer, W. & Muckli, L. J. Neurosci. 30, 2960–2966 (2010).
Schapiro, A.C., Kustner, L.V. & Turk-Browne, N.B. Curr. Biol. 22, 1622–1627 (2012).
Hawkins, J. & Blakeslee, S. On Intelligence (Times Books, 2004).
Kok, P., Jehee, J.F. & de Lange, F.P. Neuron 75, 265–270 (2012).
Marr, D. Phil. Trans. R. Soc. B. 262, 23–81 (1971).
Cohen, N.J. & Eichenbaum, H. Memory, Amnesia, and the Hippocampal System (MIT Press, 1993).
Leutgeb, S. & Leutgeb, J.K. Learn. Mem. 14, 745–757 (2007).
Ji, D. & Wilson, M.A. Nat. Neurosci. 10, 100–107 (2007).
Bosch, S.E., Jehee, J.F., Fernández, G. & Doeller, C.F. J. Neurosci. 34, 7493–7500 (2014).
Hindy, N.C. & Turk-Browne, N.B. Cereb. Cortex 10.1093/cercor/bhv030 (2015).
O'Doherty, J. et al. Science 304, 452–454 (2004).
Bar, M. et al. Proc. Natl. Acad. Sci. USA 103, 449–454 (2006).
Dale, A.M. Hum. Brain Mapp. 8, 109–114 (1999).
Yushkevich, P.A. et al. Hum. Brain Mapp. 36, 258–287 (2015).
Aly, M. & Turk-Browne, N.B. Cereb. Cortex 26, 783–796 (2016).
Duvernoy, H.M. The Human Hippocampus: Functional Anatomy, Vascularization and Serial Sections with MRI (Springer: 2005).
Carr, V.A., Rissman, J. & Wagner, A.D. Neuron 65, 298–308 (2010).
Treves, A., Tashiro, A., Witter, M.P. & Moser, E.I. Neuroscience 154, 1155–1172 (2008).
Ketz, N., Morkonda, S.G. & O'Reilly, R.C. PLoS Comput. Biol. 9, 1–9 (2013).
Nakashiba, T. et al. Cell 149, 188–201 (2012).
Duncan, K., Tompary, A. & Davachi, L. J. Neurosci. 34, 11188–11198 (2014).
Bakker, A., Kirwan, C.B., Miller, M. & Stark, C.E. Science 319, 1640–1642 (2008).
Fischl, B. et al. Cereb. Cortex 18, 1973–1980 (2008).
Hinds, O.P. et al. Neuroimage 39, 1585–1599 (2008).
Dale, A.M., Fischl, B. & Sereno, M.I. Neuroimage 9, 179–194 (1999).
Wang, L., Mruczek, R.E., Arcaro, M.J. & Kastner, S. Cereb. Cortex 25, 3911–3931 (2014).
Kriegeskorte, N., Goebel, R. & Bandettini, P. Proc. Natl. Acad. Sci. USA 103, 3863–3868 (2006).
Debas, K. et al. Neuroimage 99, 50–58 (2014).
Gabitov, E., Manor, D. & Karni, A. J. Cogn. Neurosci. 27, 736–751 (2015).
Smith, S.M. et al. Neuroimage 23 (suppl. 1), S208–S219 (2004).
Greve, D.N. & Fischl, B. Neuroimage 48, 63–72 (2009).
Jenkinson, M., Bannister, P., Brady, M. & Smith, S. Neuroimage 17, 825–841 (2002).
Andersson, J.L., Jenkinson, M. & Smith, S. FMRIB Tech. Rep. TR07JA2 (2007).
O'Reilly, R.C. & Rudy, J.W. Psychol. Rev. 108, 311–345 (2001).
Komorowski, R.W., Manns, J.R. & Eichenbaum, H. J. Neurosci. 29, 9918–9929 (2009).
Hindy, N.C., Solomon, S.H., Altmann, G.T.M. & Thompson-Schill, S.L. Cereb. Cortex 25, 884–894 (2015).
Efron, B. & Tibshirani, R. Stat. Sci. 1, 54–75 (1986).
Acknowledgements
This work was supported by NIH grants F32 EY023162, S10 OD016277, and R01 EY021755. The authors thank P. Kok and A. Schapiro for helpful comments on an earlier draft.
Author information
Authors and Affiliations
Contributions
N.C.H., F.Y.N., and N.B.T.-B. designed the experiment, reviewed the analyses, and discussed the results. N.C.H. and F.Y.N. collected the data. N.C.H. performed the analyses. N.C.H. and N.B.T.-B. wrote and revised the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Task procedure
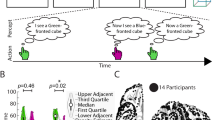
(a) The stimuli consisted of 12 fractal-like images. (b) During initial training and scanning, full-sequence trials began with a cue at fixation. A double-headed arrow prompted participants to press a button with their choice of left or right hand, at which point an outcome replaced the cue. Cue+action trials contained the same structure, but a blank screen appeared for 1000 ms instead of the outcome. Outcome-only trials contained just an outcome for 1000 ms without a preceding cue or action. (c) Behavioral tests were conducted to assess learning at different stages. Each test trial involved a verbal “top” or “bottom” response to select which of the two outcomes associated with the cue and action seemed most probable.
Supplementary Figure 2 Resampled decoding performance
Subject-level bootstrap resampling41 was used to confirm the random-effects significance of classification for each ROI, without the assumptions of parametric tests. (a) Sequence decoding was reliable in CA2–CA3–DG (P = 0.008) and CA1 (P = 0.004), but not in subiculum, V1, or V2 (P > 0.77). (b) Outcome decoding was reliable in V1 (P = 0.0005) and V2 (P = 0.01), but not in CA2–CA3–DG, CA1, or subiculum (P > 0.55). Error bars indicate 95% confidence intervals. *P < 0.05, **P < 0.01, ***P < 0.001
Supplementary Figure 3 Unpredictable trials
No reliable effects were obtained in a control condition where actions did not predict outcomes, providing evidence that predictive actions were required to observe sequence decoding and outcome decoding. (a) Sequence decoding for unpredictable trials was not reliable in either CA–DG (t23 = 0.85, P = 0.40) or V1–V2 (t23 = 0.13, P = 0.90). (b) Outcome decoding for unpredictable trials was also not reliable in either CA–DG (t23 = 0.61, P = 0.55) or V1–V2 (t23 = 0.60, P = 0.56). Error bars depict ±1 s.e.m.
Supplementary Figure 4 Action decoding
Predictable and unpredictable trials were used to examine action information in the hippocampus and the right putamen. (a) Classifiers trained to distinguish predictable full-sequence trials with left vs. right actions for one cue (subscript 1) could not reliably decode predictable full-sequence trials with corresponding left vs. right actions for the other cue (subscript 2) in either CA–DG (t23 = -0.39, P = 0.70) or the right putamen (t23 = 1.50, P = 0.15). (b) These classifiers also could not reliably decode predictable cue+action trials with left vs. right actions for another cue in CA–DG (t23 = 1.66, P = 0.11), but could marginally decode them in the right putamen (t23 =2.05, P = 0.05). (c) Classifiers trained to distinguish unpredictable full-sequence trials with left vs. right actions (orthogonal to outcomes) could not reliably decode unpredictable cue+action trials with corresponding left vs. right actions in CA–DG (t23 = 0.11, P = 0.92), but could decode them in the right putamen (t23 = 2.85, P = 0.009). Error bars depict ±1 s.e.m. ~P < 0.10, **P < 0.01
Supplementary Figure 5 Within-action classification
Classifiers trained to distinguish predictable full-sequence trials with different cues and outcomes but identical left or right actions reliably decoded predictable cue+action trials with different cues but identical actions in CA–DG (t23 = 2.20, P = 0.04), but not in the right putamen (t23 = -0.65, P = 0.52). Error bars depict ±1 s.e.m. *P < 0.05
Supplementary Figure 6 Brain–behavior correlations
For the predictable condition, voice RT in the behavioral tests (a) had a marginally negative correlation across participants with sequence decoding in CA–DG (r22 = -0.36, P = 0.09) and (b) had a reliably negative correlation across participants with outcome decoding in V1–V2 (r22 = -0.54, P = 0.007). That is, better outcome decoding and to some extent better sequence decoding were associated with faster outcome identification at test. For the unpredictable condition, test RT was not correlated with either (c) sequence decoding in CA–DG (r22 = -0.10, P = 0.63) or (d) outcome decoding in V1–V2 (r22 = -0.02, P = 0.92). ~ P < 0.10, ** P < 0.01
Supplementary Figure 7 Cross-classification
Consistent with our hypotheses that full-sequence trials would most effectively target conjunctive representations in the hippocampus and that outcome-only trials would most effectively target outcome representations in early visual cortex, cross-classification in CA–DG was reliable only when full-sequence trials were part of either the training or testing set and cross-classification in V1–V2 was reliable only when outcome-only trials were part of either the training or testing set. (a) Cross-classification from full-sequence to cue+action trials (sequence decoding elsewhere) was reliable in CA–DG (t23 = 2.64, P = 0.01), but not in V1–V2 (t23 = -0.97, P = 0.34). In contrast, cross-classification from full-sequence to outcome-only trials was reliable in both CA–DG (t23 = 2.29, P = 0.03) and V1–V2 (t23 = 3.00, P = 0.006). (b) Cross-classification from outcome-only to cue+action trials (outcome decoding elsewhere) was reliable in V1–V2 (t23 = 2.99, P = 0.007), but not in CA–DG (t23 = 0.09, P = 0.93), and cross-classification from outcome-only to full-sequence trials was likewise reliable in V1–V2 (t23 = 2.35, P = 0.03), but not in CA–DG (t23 = 1.60, P = 0.12). (c) Similar to sequence decoding, cross-classification from cue+action to full-sequence trials was reliable in CA–DG (t23 = 2.69, P = 0.01), but not in V1–V2 (t23 = -0.03, P = 0.97). Unlike outcome decoding, cross-classification from cue+action to outcome-only trials was not reliable in V1–V2 (t23 = 1.24, P = 0.23), and still not in CA–DG (t23 = 1.14, P = 0.27). The difference in V1–V2 cross-classification for [outcome-only → cue+action] vs. [cue+action → outcome-only] may be due to worse classifier training with cue+action trials: there were more outcome-only than cue+action training examples in the design, cue+action trials contained an additional uninformative stimulus (the cue), and the outcome representation on cue+action trials may have been weaker because it reflected an internal expectation rather than an external stimulus. Error bars depict ±1 s.e.m. *P < 0.05, **P < 0.01
Supplementary Figure 8 Resampled hippocampal–visual relationships
Subject-level bootstrap resampling41 was used to confirm the random-effects significance of within- and across-participant relationships between sequence decoding in the hippocampus and outcome decoding in early visual cortex, without the assumptions of parametric tests. (a) Outcome decoding in V1–V2 was more reliable (P = 0.02) on trials where sequence decoding in CA–DG was correct (vs. 50% chance: P = 0.0006) vs. incorrect (P = 0.28). (b) Individual differences in V1–V2 outcome decoding could be predicted from CA–DG sequence decoding (P = 0.004). Error bars and bands indicate 95% confidence intervals. *P < 0.05, **P < 0.01, ***P < 0.001
Supplementary Figure 9 Timecourse analysis
The temporal precedence of CA–DG sequence decoding vs. V1–V2 outcome decoding might provide tentative evidence about the directionality of the relationship between these regional processes. (a) If sequence information in the hippocampus precedes outcome information in visual cortex, then CA–DG sequence decoding at TR 3 should be predictive of V1–V2 outcome decoding on TR 4 in a multinomial regression (purple). (b) If outcome information in visual cortex precedes sequence information in the hippocampus, then V1–V2 outcome decoding at TR 3 should be predictive of CA–DG sequence decoding at TR 4 (green). (c) Classifiers for CA–DG were trained on patterns of GLM beta parameters from the full-sequence trials, whereas classifiers for V1–V2 were trained on patterns of GLM beta parameters from the outcome-only trials. All classifiers were tested on raw activity patterns from cue+action trials that were z-scored and extracted from timepoints around the peak response (TRs 3 and 4 after trial onset). CA–DG sequence decoding at TR 3 predicted V1–V2 outcome decoding at TR 4 (purple; t23 = 2.30, P = 0.03), whereas V1–V2 outcome decoding at TR 3 did not reliably predict CA–DG sequence decoding at TR 4 (green; t23 = 0.84, P = 0.41). In contrast, CA–DG sequence decoding did not reliably predict V1–V2 outcome decoding (gray) within either TR 3 (t23 = 1.50, P = 0.15) or TR 4 (t23 = -0.31, P = 0.76). CA–DG sequence decoding at TR 3 was more predictive of V1–V2 outcome decoding at TR 4 than was CA–DG sequence decoding at TR 4 (t23 = 2.21, P = 0.04). CA–DG sequence decoding at TR 3 did not predict V1–V2 outcome decoding at TR 4 more reliably than V1–V2 outcome decoding at TR 3 (t23 = 1.25, P = 0.23), although the difference was in the same direction. Mean parameter estimates are shown in bold font, with s.e.m. in parentheses. *P < 0.05
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 (PDF 2902 kb)
Rights and permissions
About this article
Cite this article
Hindy, N., Ng, F. & Turk-Browne, N. Linking pattern completion in the hippocampus to predictive coding in visual cortex. Nat Neurosci 19, 665–667 (2016). https://doi.org/10.1038/nn.4284
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4284
This article is cited by
-
From cognitive maps to spatial schemas
Nature Reviews Neuroscience (2023)
-
Hippocampal representations switch from errors to predictions during acquisition of predictive associations
Nature Communications (2022)
-
Moment-by-moment tracking of naturalistic learning and its underlying hippocampo-cortical interactions
Nature Communications (2021)
-
Object representations in the human brain reflect the co-occurrence statistics of vision and language
Nature Communications (2021)
-
Cross-modal auditory priors drive the perception of bistable visual stimuli with reliable differences between individuals
Scientific Reports (2021)