Abstract
There are many transgenic GFP reporter lines that allow the visualization of specific populations of cells. Using such lines for functional studies requires a method that transforms GFP into a molecule that enables genetic manipulation. We developed a method that exploits GFP for gene manipulation, Cre recombinase dependent on GFP (CRE-DOG), a split component system that uses GFP and its derivatives to directly induce Cre/loxP recombination. Using plasmid electroporation and AAV viral vectors, we delivered CRE-DOG to multiple GFP mouse lines, which led to effective recombination selectively in GFP-labeled cells. Furthermore, CRE-DOG enabled optogenetic control of these neurons. Beyond providing a new set of tools for manipulation of gene expression selectively in GFP+ cells, we found that GFP can be used to reconstitute the activity of a protein not known to have a modular structure, suggesting that this strategy might be applicable to a wide range of proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Shimomura, O., Johnson, F.H. & Saiga, Y. Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan, Aequorea. J. Cell. Comp. Physiol. 59, 223–239 (1962).
Tsien, R.Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Chalfie, M. et al. Green fluorescent protein as a marker for gene expression. Science 263, 802–805 (1994).
Gong, S. et al. A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature 425, 917–925 (2003).
Heintz, N. Gene expression nervous system atlas (GENSAT). Nat. Neurosci. 7, 483 (2004).
Siegert, S. et al. Genetic address book for retinal cell types. Nat. Neurosci. 12, 1197–1204 (2009).
Tang, J.C. et al. A nanobody-based system using fluorescent proteins as scaffolds for cell-specific gene manipulation. Cell 154, 928–939 (2013).
Rothbauer, U. et al. Targeting and tracing antigens in live cells with fluorescent nanobodies. Nat. Methods 3, 887–889 (2006).
Kirchhofer, A. et al. Modulation of protein properties in living cells using nanobodies. Nat. Struct. Mol. Biol. 17, 133–138 (2010).
Caussinus, E., Kanca, O. & Affolter, M. Fluorescent fusion protein knockout mediated by anti-GFP nanobody. Nat. Struct. Mol. Biol. 19, 117–121 (2012).
Sadowski, I., Ma, J., Triezenberg, S. & Ptashne, M. GAL4-VP16 is an unusually potent transcriptional activator. Nature 335, 563–564 (1988).
Ho, S.N., Biggar, S.R., Spencer, D.M., Schreiber, S.L. & Crabtree, G.R. Dimeric ligands define a role for transcriptional activation domains in reinitiation. Nature 382, 822–826 (1996).
Atasoy, D., Aponte, Y., Su, H.H. & Sternson, S.M.A. FLEX switch targets Channelrhodopsin-2 to multiple cell types for imaging and long-range circuit mapping. J. Neurosci. 28, 7025–7030 (2008).
Fenno, L.E. et al. Targeting cells with single vectors using multiple-feature Boolean logic. Nat. Methods 11, 763–772 (2014).
Pivetta, C., Esposito, M.S., Sigrist, M. & Arber, S. Motor-circuit communication matrix from spinal cord to brainstem neurons revealed by developmental origin. Cell 156, 537–548 (2014).
Dymecki, S.M., Ray, R.S. & Kim, J.C. Mapping cell fate and function using recombinase-based intersectional strategies. Methods Enzymol. 477, 183–213 (2010).
Jullien, N., Sampieri, F., Enjalbert, A. & Herman, J.P. Regulation of Cre recombinase by ligand-induced complementation of inactive fragments. Nucleic Acids Res. 31, e131 (2003).
Vila-Perelló, M. & Muir, T.W. Biological applications of protein splicing. Cell 143, 191–200 (2010).
Tyszkiewicz, A.B. & Muir, T.W. Activation of protein splicing with light in yeast. Nat. Methods 5, 303–305 (2008).
Mootz, H.D. & Muir, T.W. Protein splicing triggered by a small molecule. J. Am. Chem. Soc. 124, 9044–9045 (2002).
Gill, G. & Ptashne, M. Negative effect of the transcriptional activator GAL4. Nature 334, 721–724 (1988).
Anraku, Y., Mizutani, R. & Satow, Y. Protein splicing: its discovery and structural insight into novel chemical mechanisms. IUBMB Life 57, 563–574 (2005).
Samson, M., Emerson, M.M. & Cepko, C.L. Robust marking of photoreceptor cells and pinealocytes with several reporters under control of the Crx gene. Dev. Dyn. 238, 3218–3225 (2009).
Betley, J.N. & Sternson, S.M. Adeno-associated viral vectors for mapping, monitoring, and manipulating neural circuits. Hum. Gene Ther. 22, 669–677 (2011).
Rivlin-Etzion, M. et al. Transgenic mice reveal unexpected diversity of on-off direction-selective retinal ganglion cell subtypes and brain structures involved in motion processing. J. Neurosci. 31, 8760–8769 (2011).
Osterhout, J.A. et al. Cadherin-6 mediates axon-target matching in a non-image-forming visual circuit. Neuron 71, 632–639 (2011).
Osterhout, J.A., El-Danaf, R.N., Nguyen, P.L. & Huberman, A.D. Birthdate and outgrowth timing predict cellular mechanisms of axon target matching in the developing visual pathway. Cell Rep. 8, 1006–1017 (2014).
Tamamaki, N. et al. Green fluorescent protein expression and colocalization with calretinin, parvalbumin and somatostatin in the GAD67-GFP knock-in mouse. J. Comp. Neurol. 467, 60–79 (2003).
Yizhar, O., Fenno, L.E., Davidson, T.J., Mogri, M. & Deisseroth, K. Optogenetics in neural systems. Neuron 71, 9–34 (2011).
Nagai, T., Horikawa, K., Saito, K. & Matsuda, T. Genetically encoded Ca2+ indicators; expanded affinity range, color hue and compatibility with optogenetics. Front. Mol. Neurosci. 7, 90 (2014).
Sternson, S.M. & Roth, B.L. Chemogenetic tools to interrogate brain functions. Annu. Rev. Neurosci. 37, 387–407 (2014).
Wickersham, I.R. & Feinberg, E.H. New technologies for imaging synaptic partners. Curr. Opin. Neurobiol. 22, 121–127 (2012).
Luo, L., Callaway, E.M. & Svoboda, K. Genetic dissection of neural circuits. Neuron 57, 634–660 (2008).
Venken, K.J., Simpson, J.H. & Bellen, H.J. Genetic manipulation of genes and cells in the nervous system of the fruit fly. Neuron 72, 202–230 (2011).
Howard, D.B., Powers, K., Wang, Y. & Harvey, B.K. Tropism and toxicity of adeno-associated viral vector serotypes 1, 2, 5, 6, 7, 8, and 9 in rat neurons and glia in vitro. Virology 372, 24–34 (2008).
Semprini, S. et al. Cryptic loxP sites in mammalian genomes: genome-wide distribution and relevance for the efficiency of BAC/PAC recombineering techniques. Nucleic Acids Res. 35, 1402–1410 (2007).
Iwamoto, M., Bjorklund, T., Lundberg, C., Kirik, D. & Wandless, T.J. A general chemical method to regulate protein stability in the mammalian central nervous system. Chem. Biol. 17, 981–988 (2010).
Fridy, P.C. et al. A robust pipeline for rapid production of versatile nanobody repertoires. Nat. Methods 11, 1253–1260 (2014).
Matsuda, T. & Cepko, C.L. Electroporation and RNA interference in the rodent retina in vivo and in vitro. Proc. Natl. Acad. Sci. USA 101, 16–22 (2004).
Emerson, M.M. & Cepko, C.L. Identification of a retina-specific Otx2 enhancer element active in immature developing photoreceptors. Dev. Biol. 360, 241–255 (2011).
Matsuda, T. & Cepko, C.L. Controlled expression of transgenes introduced by in vivo electroporation. Proc. Natl. Acad. Sci. USA 104, 1027–1032 (2007).
Niwa, H., Yamamura, K. & Miyazaki, J. Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene 108, 193–199 (1991).
Shimshek, D.R. et al. Codon-improved Cre recombinase (iCre) expression in the mouse. Genesis 32, 19–26 (2002).
Haubensak, W. et al. Genetic dissection of an amygdala microcircuit that gates conditioned fear. Nature 468, 270–276 (2010).
Dhande, O.S. et al. Genetic dissection of retinal inputs to brainstem nuclei controlling image stabilization. J. Neurosci. 33, 17797–17813 (2013).
Huberman, A.D. et al. Architecture and activity-mediated refinement of axonal projections from a mosaic of genetically identified retinal ganglion cells. Neuron 59, 425–438 (2008).
Cruz-Martín, A. et al. A dedicated circuit links direction-selective retinal ganglion cells to the primary visual cortex. Nature 507, 358–361 (2014).
Hughes, D.I. et al. Morphological, neurochemical and electrophysiological features of parvalbumin-expressing cells: a likely source of axo-axonic inputs in the mouse spinal dorsal horn. J. Physiol. (Lond.) 590, 3927–3951 (2012).
Drobizhev, M., Makarov, N.S., Tillo, S.E., Hughes, T.E. & Rebane, A. Two-photon absorption properties of fluorescent proteins. Nat. Methods 8, 393–399 (2011).
Acknowledgements
We thank C. Wang of the Z. He laboratory (Boston Children's Hospital) for rAAV production (core service supported by grant NEI 5P30EY012196-17), D. Goz and S. Zhao for technical assistance, B. Huang, D. Meijer, and the Cepko, Tabin and Dymecki laboratory members for input on the manuscript, and the Neurobiology Imaging Facility (supported by NINDS P30 Core Center grant NS072030) for consultation and instrument availability. We are grateful to U. Rothbauer and H. Leonhardt (Ludwig Maximilian University of Munich) for providing GBPs. This work was funded by the Howard Hughes Medical Institute (C.L.C.), the Nancy Lurie Marks Foundation (W.G.R.), the Lefler Foundation (W.G.R.), the Knights Templar Eye Foundation (O.S.D.), the McKnight Foundation (A.D.H.), The Pew Charitable Trusts (A.D.H.), the Glaucoma Research Foundation (A.D.H.), and grants from the US National Institutes of Health (R01 EY022157-01 to A.D.H. and R01 NS32405 to W.G.R.). S.R. is funded by an Alice and Joseph Brooks fellowship and a F32 NS087708 training grant and E.D. is funded by a F31 AG041582 (NIA) training grant.
Author information
Authors and Affiliations
Contributions
J.C.Y.T. and C.L.C. initiated and coordinated the entire project. J.C.Y.T. originated the idea and carried out all of the in vitro molecular biology, electroporation and proof-of-principle rAAV experiments to validate the concept. S.R. coordinated brain injections, and performed electrophysiological recordings of cerebellar slices, immunohistochemistry, imaging and analysis of cerebellar and cortical data. O.S.D. and A.D.H. performed viral injections into retinas of GFP lines and subsequent tissue processing, imaging and analysis. V.E.A. and S.C. performed viral injections into spinal cords and subsequent tissue processing, imaging and analysis. E.D. and S.W.L. constructed the ChAG-loxP-TagBFP-loxP-mCherry construct. S.W.L. performed electroporation into the Tg(PROX1-GFP) line and subsequent tissue processing. I.R.D. performed viral injections into the cortex and cerebellum and provided feedback on experiments. A.D.H. and W.G.R. supervised aspects of the project involving rAAV injection into transgenic GFP retinas and brains, respectively. C.L.C. supervised the entire project. J.C.Y.T. and C.L.C. wrote the paper, with contributions from all of the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Comparison of GFP-dependent transcription and Cre recombination systems.
(a) Transcription Devices Dependent on GFP (T-DDOG) can be used to turn on Cre expression, but 3 components have to be delivered to cells. (b) In comparison, Cre Recombinase Dependent On GFP (CRE-DOG) functions via 2 components that directly interact with GFP to initiate Cre recombination. DBD, DNA Binding Domain. AD, Activation Domain. UAS, Upstream Activating Sequence. Pacman shapes – GFP binding proteins.
Supplementary Figure 2 The original GFP-dependent Cre recombinase (CRE-DOGOG).
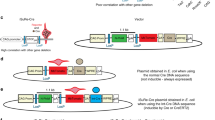
(a) CRE-DOGOG is made of N-CreintG and C-CreintG. CAG-loxP-Neo-loxP-luc2 (CALNL-luc2) is a Cre-dependent luciferase reporter. N-VMA and C-VMA are N- and C-terminal split intein portions from yeast VMA1 gene, respectively. GBP, GFP-binding protein. NLS, nuclear localization signal. (b-c) Luciferase assays showing specificity of CRE-DOGOG activity for each component of the system (b) and for different fluorescent proteins (c). n=9 replicate samples each condition (see Online Methods for n definition). In box plots, boxes show 25th to 75th percentile, line in box shows median, whiskers show minimum to maximum range. Scatter plots show median, interquartile range. (d-j) GFP directly controls CRE-DOGOG activity in a cell-specific manner in vivo. (d) Schematic of experiment. CAG-LNL-DsRed is a Cre-dependent reporter. CAG-nlacZ, which expresses n-βgal, is a substitution plasmid for GFP. (e-g) CRE-DOGOG induces expression of CALNL-DsRed reporter in a GFP-dependent manner in electroporated retinas. DsRed expression pattern depends on whether the broadly active CAG (e) or rod-specific Rho (f) promoters drives GFP expression. Scale bar, 20 μm. (h) Efficiency of CRE-DOGOG recombination in electroporated retinas. An electroporated cell is defined as either GFP+ or n-βgal+. Rod photoreceptors in the outer nuclear layer (ONL) were quantified. Sample size: GFP+ condition - At least 50 GFP+ cells/retina (n=3 retinas). Rho-GFP condition - At least 50 GFP+ cells/retina (n=4 retinas). No GFP condition - At least 100 n-βgal+ cells/retina (n=3 retinas). (i) GFP specificity of floxed DsRed+ ONL cells. Sample size: At least 30 to 50 cells/retina (CAG-GFP, n=3 retinas. Rho-GFP, n=4 retinas) (j) Expression pattern of floxed DsRed+ cells compared to GFP+ cells in ONL versus inner nuclear layer (INL). Sample size: CAG-GFP, at least 550 GFP+ cells/retina, at least 180 DsRed+ cells/retina, n=3 retinas; Rho-GFP, at least 200 GFP+ cells/retina, at least 100 DsRed+ cells/retina, n=4 retinas. For (e-j), consistent results were obtained in two independent experiments.
Supplementary Figure 3 CRE-DOG appears to act independently of protein splicing.
(a-b) Luciferase assays testing various CRE-DOGOG variants for GFP-dependent activation of a CALNL-luc2 reporter. (a) Mutations known to disrupt the protein splicing ability of VMA intein did not drastically alter CRE-DOGOG activity. N-splicing site refer to a cysteine residue in the N-intein portions of N-CreintG of CRE-DOGOG. C-splicing site refer to a cysteine residue in the C-intein portions of C-CreintG of CRE-DOGOG. n=6 or 9 replicate samples each condition. (b) Both intein portions were required for the tight GFP-dependent recombination activity of CRE-DOGOG. “Int” indicates presence of intein portion. Removal of either intein portion adversely affected CRE-DOGOG specificity for GFP. Each portion independently contributed to the GFP-specificity of CRE-DOGOG. n=9 each condition. In box plots, boxes show 25th to 75th percentile, line in box shows median, whiskers show minimum to maximum range. (c) Testing for protein splicing mechanism for CRE-DOGOG function. FLAG tagged C-terminal component of CRE-DOGOG is ~52 kDa. If protein splicing occurs, then the ~37 kDa full-length Cre product should be detected by anti-FLAG. (d) Western blot testing the protein splicing model in (c). 293T cells transfected with indicated components (CRE-DOGOG-FLAG and GFP) were lysed 1 day post-transfection. Lysate supernatant (nuclear and cytoplasmic content) were probed for protein products labeled by FLAG tag. The results failed to support a protein splicing mechanism, due to absence of ~37 kDa band. Consistent results were obtained in an independent experiment with whole cell lysate loading.
Supplementary Figure 4 Truncation of N-VMA intein fragment promotes enhanced GFP-specificity and activity of CRE-DOG.
(a-b) Luciferase assays performed with transfected 293T cells, 24 hours post-transfection. Split Cre constructs were adjusted to be equimolar in amount of transfected plasmids. (a) Broad truncation scans along N-VMA intein fragment revealed a 43 amino acid fragment that promotes increased GFP-dependent recombination and specificity in the context of the split-Cre/GBP7 fusion construct. This truncated construct is code named “121trc”. n=14, 15 or 18 replicate samples per condition, pooled from 2 independent experiments. Each condition value is obtained from 5 or 6 independent transfections assayed in 2 or 3 technical replicates. Two-tailed Student’s t test assuming unequal variance. *, p<0.001 (0.0002). **, p<0.0001 (1,2x10-13) (b) Finer resolution truncation scan along the 43 amino acid N-VMA fragment. An additional truncation of 3 amino acid increased GFP-dependent recombination and specificity in the context of the 121trc construct. This construct is named N-CretrcintG. n=19 or 21 per condition, pooled from 3 independent experiments. Each condition value from 6 or 7 independent transfections assayed in 2 or 3 technical replicates. Two-tailed Student’s t test assuming unequal variance. *, p<0.05 (0.019). **, p<0.0001 (5.3x10-8). Green and grey dotted lines denote activity levels of control CRE-DOG for comparison, with and without GFP, respectively. In box plots, boxes show 25th to 75th percentile, line in box shows median, whiskers show minimum to maximum range.
Supplementary Figure 5 Comparison of original CRE-DOG and optimized CRE-DOG.
Lucifearse assay comparing CRE-DOGOG and CRE-DOGOPT activity in transfected 293T cells. The CALNL-luc2 reporter was used. CRE-DOGOPT had greater activity and lower background than CRE-DOGOG. < * indicates CRE-DOGOPT has significantly greater activity than CRE-DOGOG. n=12 per condition. Samples from 4 independent transfection assayed in triplicates. * indicates p<0.05 (0.04). ** indicates p<0.0001 (4.6x10-8). Two-tailed Student’s t test assuming unequal variance. Cells were harvested 1 day post-transfection. In box plots, boxes show 25th to 75th percentile, line in box shows median, whiskers show minimum to maximum range.
Supplementary Figure 6 Quantification of electroporation experiment in Figure 2 a-c.
(a-b) Efficiency and GFP-dependency of CRE-DOGOPT activity in electroporated Tg(CRX-GFP)+ or – retinas. Electroporated ONL (a) or INL (b) cells were defined by expression of n-βgal+ cells (from co-electroporated CAG-nlacZ plasmids). Sample size: for (a), both GFP+ and GFP- conditions - at least 300 n-βgal+ ONL cells/retina, at least 15 n-βgal+ INL cells/retina; n=3 or 4 retinas per condition; for (b), at least 100 TagBFP+ cells/retina in all conditions; n=3 retinas per condition. (c) Quantification of floxed DsRed expression intensity in Tg(CRX-GFP)+, DsRed+ ONL and INL cells. n=28 ONL cells and 15 INL cells. (d-e) Efficiency and GFP-dependency of CRE-DOGOPT activity in electroporated Tg(PROX1-GFP)+ or – retinas. Electroporated ONL (d) or INL (e) cells are defined by staining for Anti-TagBFP (TagBFP is from ChAG-loxP-TagBFP-loxP-mCherry cassette. Antibody cross-reacts with mCherry too). (f) Quantification of floxed mCherry expression intensity in Tg(PROX1-GFP)+, mCherry+ ONL and INL cells. n=65 ONL cells and 67 INL cells. (g) GFP specificity of ONL DsRed (Tg(CRX-GFP) set) or INL mCherry (Tg(PROX1-GFP) set) cells. n=4 retinas per condition, at least 50 DsRed+ or mCherry+ cells counted per retina. All scatter plots show median and interquartile range. In box plots, boxes show 25th to 75th percentile, line in box shows median, whiskers show minimum to maximum range.
Supplementary Figure 7 Delivery of CRE-DOGOPT to the mouse nervous system by rAAV.
(a) Co-injection of rAAV (serotype 2/8) encoding EF1α-driven N-CretrcintG and C-CreintG and rAAV-FLEX-tdT into the P0 murine retina along with either rAAV encoding EF1α-driven GFP (top row) or ZsGreen (bottom row). Retinas were harvested between P21–30. Top row: GFP-dependent activation of tdT in the horizontal cell layer of the retina. Bottom row: little background CRE-DOGOPT activity was observed. Note: ZsGreen was found to aggregate heavily in the retina. Images representative of at least 3 retinas per condition. Scale bar, 30 μm. (b) Little to no background FLEX-tdT activity in rAAV infected cells of the retinal GCL. Co-injection of rAAV (serotype 2/8) encoding EF1α-driven N-CretrcintG and C-CreintG and rAAV-FLEX-tdT into the murine retina. Mice were of age P77 at time of injection. Retinas were harvested 3 weeks post-infection and immunostained for polyclonal Cre antibody (left panel). No tdT expression was detected in Cre-immunopositive cells of the ganglion cell layer (GCL) (center panel). Merge of all channels with DAPI (right panel) suggests that faint red dots were background signals or debris and were not cells. Images are representative of 3 retinas. Scale bar, 20 μm.
Supplementary Figure 8 Cre immunostaining establish GFP specificity of CRE-DOGOPT in rAAV-infected cerebella (Related to Figure 6).
2/1 serotype rAAV-CRE-DOGOPT and rAAV-FLEX-tdT were injected into the cerebellum of Tg(GAD67-GFP)+ or GFP negative animals and harvested 3 weeks later. Same experiment as in Fig. 6. (a) Top panel: Images of rAAV-injected lobule V in Tg(GAD67-GFP)+ animal. Anti-Cre immunostain and GFP-dependent expression of tdT indicate successful virus injection. Bottom Panel: Images of uninfected lobule in the same animal show no expression of Cre and tdT. Scale bar, 50 μm (b) Top Panel: Images of rAAV injected lobule V in Tg(GAD67-GFP) negative animal. Anti-Cre immunostain indicates successful infection, but tdT is not expressed in the absence of GFP. Bottom Panel: Images of uninfected lobule in the same animal shows no expression of Cre and tdT. In all Merged panels, green is GFP, blue is Anti-Cre staining, red is tdT. Note the infected lobule panels in (b) were also shown in main text Fig. 6. All images are representative of 4 Tg(GAD-GFP)+ animals and 3 GFP-negative animals.
Supplementary Figure 9 Characterization and quantification of rAAV-infected cerebellum of Tg(GAD67-GFP) and GFP negative animals (Related to Figure 6).
a) Images of rAAV-CRE-DOGOPT and rAAV-FLEX-tdT-infected lobule V, immunostained for GFP (blue). (b) Quantification of anti-GFP immunostained sections. (c) Immunostaining of rAAV-infected lobule V for the molecular layer interneuron marker Parvalbumin (PV) (blue, top), or the Purkinje cell marker Calbindin (Cb) (blue, bottom). Scale bar for (a) and (c), 50 μm. (d) Quantification of tdT expression in different cell types. In densely infected areas, tdT+ was expressed in (mean +/- s.d.): 65 +/- 16 % of Purkinje Cells (108 Cb+ cells), 44 +/- 22 % molecular layer interneurons (394 PV+ cells) and 53 +/- 21 % of granular layer interneurons (99 GLIs, identified based on location and anatomy, not shown). n=4 animals. (e) Sagittal cerebellar section showing low leakage of FLEX-tdT in GFP negative brains. Arrowhead indicates a single tdT+ Purkinje cell in infected lobule V. Scale bar, 1 mm. (f) Example of tdT+ cells in GFP negative tissue. Left panel: Bergmann glia cell (BG). Scale bar, 50 μm. Top right: molecular layer interneuron (MLI) Scale bar in BG panel also applies to MLI panel, 50 μm. Bottom right: granule cell (GC). Scale bar, 30 μm. (g) Summary quantification of cell types (n=84 cells, 22 sections, 3 animals) labeled by tdT in GFP negative cerebellum infected with rAAV-CRE-DOGOPT and FLEX-tdT mix. MLI, molecular layer interneuron. PC, Purkinje cell. BG, Bergmann glia. Astro, astrocyte. GC, granule cell. Other, identities include oligodendrocyte, unipolar brush cell, NG2 cell. (h) CRE-DOGOPT system enables visualization of fine neuronal processes in the brain. Left panel: tdT expression in GFP+ Purkinje cells allows tracing of axons (arrowheads). Middle panel: higher magnification of Purkinje cells shows tdT labels fine dendritic processes, dendritic spines and synaptic boutons on cell bodies. Right panel: tdT-expression in GLI cell bodies (asterisks) and their axonal projections throughout the molecular layer (arrowheads). Scale bar, 30 μm. All images reported are gathered from or representative of 4 Tg(GAD-GFP)+ animals and 3 GFP negative animals. Scatter plots indicate median and interquartile range.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Tables 1 and 2 (PDF 1589 kb)
Source data
Rights and permissions
About this article
Cite this article
Tang, J., Rudolph, S., Dhande, O. et al. Cell type–specific manipulation with GFP-dependent Cre recombinase. Nat Neurosci 18, 1334–1341 (2015). https://doi.org/10.1038/nn.4081
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4081
This article is cited by
-
Adolescent stress impairs postpartum social behavior via anterior insula-prelimbic pathway in mice
Nature Communications (2023)
-
A CRISPR toolbox for generating intersectional genetic mouse models for functional, molecular, and anatomical circuit mapping
BMC Biology (2022)
-
Nanobody-based RFP-dependent Cre recombinase for selective anterograde tracing in RFP-expressing transgenic animals
Communications Biology (2022)
-
Noninvasive optical activation of Flp recombinase for genetic manipulation in deep mouse brain regions
Nature Communications (2019)
-
Large scale matching of function to the genetic identity of retinal ganglion cells
Scientific Reports (2017)