Abstract
We found that a neuron-specific isoform of LSD1, LSD1n, which results from an alternative splicing event, acquires a new substrate specificity, targeting histone H4 Lys20 methylation, both in vitro and in vivo. Selective genetic ablation of LSD1n led to deficits in spatial learning and memory, revealing the functional importance of LSD1n in neuronal activity–regulated transcription that is necessary for long-term memory formation. LSD1n occupied neuronal gene enhancers, promoters and transcribed coding regions, and was required for transcription initiation and elongation steps in response to neuronal activity, indicating the crucial role of H4K20 methylation in coordinating gene transcription with neuronal function. Our results indicate that this alternative splicing of LSD1 in neurons, which was associated with altered substrate specificity, serves as a mechanism acquired by neurons to achieve more precise control of gene expression in the complex processes underlying learning and memory.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
17 August 2015
In the version of this article initially published online, author Cagdas Tazearslan's name was misspelled Tazearsalan. The error has been corrected for the print, PDF and HTML versions of this article.
References
Sweatt, J.D. The emerging field of neuroepigenetics. Neuron 80, 624–632 (2013).
Borrelli, E., Nestler, E.J., Allis, C.D. & Sassone-Corsi, P. Decoding the epigenetic language of neuronal plasticity. Neuron 60, 961–974 (2008).
Ronan, J.L., Wu, W. & Crabtree, G.R. From neural development to cognition: unexpected roles for chromatin. Nat. Rev. Genet. 14, 347–359 (2013).
Telese, F., Gamliel, A., Skowronska-Krawczyk, D., Garcia-Bassets, I. & Rosenfeld, M.G. “Seq-ing” insights into the epigenetics of neuronal gene regulation. Neuron 77, 606–623 (2013).
Andrés, M.E. et al. CoREST, a functional corepressor required for regulation of neural-specific gene expression. Proc. Natl. Acad. Sci. USA 96, 9873–9878 (1999).
Shi, Y. et al. Coordinated histone modifications mediated by a CtBP co-repressor complex. Nature 422, 735–738 (2003).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Metzger, E. et al. LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 437, 436–439 (2005).
Garcia-Bassets, I. et al. Histone methylation–dependent mechanisms impose ligand dependency for gene activation by nuclear receptors. Cell 128, 505–518 (2007).
Wang, J. et al. Opposing LSD1 complexes function in developmental gene activation and repression programmes. Nature 446, 882–887 (2007).
Zibetti, C. et al. Alternative splicing of the histone demethylase LSD1/KDM1 contributes to the modulation of neurite morphogenesis in the mammalian nervous system. J. Neurosci. 30, 2521–2532 (2010).
Huang, J. et al. p53 is regulated by the lysine demethylase LSD1. Nature 449, 105–108 (2007).
Laurent, B. et al. A specific LSD1/KDM1A isoform regulates neuronal differentiation through H3K9 demethylation. Mol. Cell 57, 957–970 (2015).
Whyte, W.A. et al. Enhancer decommissioning by LSD1 during embryonic stem cell differentiation. Nature 482, 221–225 (2012).
Kim, T.K. et al. Widespread transcription at neuronal activity–regulated enhancers. Nature 465, 182–187 (2010).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Telese, F. et al. LRP8-Reelin-regulated neuronal (LRN) enhancer signature underlying learning and memory formation. Neuron 86, 696–710 (2015).
Core, L.J., Waterfall, J.J. & Lis, J.T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Lin, Y. et al. Activity-dependent regulation of inhibitory synapse development by Npas4. Nature 455, 1198–1204 (2008).
Schaukowitch, K. et al. Enhancer RNA facilitates NELF release from immediate early genes. Mol. Cell 56, 29–42 (2014).
Nishioka, K. et al. PR-Set7 is a nucleosome-specific methyltransferase that modifies lysine 20 of histone H4 and is associated with silent chromatin. Mol. Cell 9, 1201–1213 (2002).
Schotta, G. et al. A chromatin-wide transition to H4K20 monomethylation impairs genome integrity and programmed DNA rearrangements in the mouse. Genes Dev. 22, 2048–2061 (2008).
Qi, H.H. et al. Histone H4K20/H3K9 demethylase PHF8 regulates zebrafish brain and craniofacial development. Nature 466, 503–507 (2010).
Liu, W. et al. PHF8 mediates histone H4 lysine 20 demethylation events involved in cell cycle progression. Nature 466, 508–512 (2010).
Stender, J.D. et al. Control of proinflammatory gene programs by regulated trimethylation and demethylation of histone H4K20. Mol. Cell 48, 28–38 (2012).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Saha, R.N. et al. Rapid activity-induced transcription of Arc and other IEGs relies on poised RNA polymerase II. Nat. Neurosci. 14, 848–856 (2011).
Hargreaves, D.C., Horng, T. & Medzhitov, R. Control of inducible gene expression by signal-dependent transcriptional elongation. Cell 138, 129–145 (2009).
Kwak, H. & Lis, J.T. Control of transcriptional elongation. Annu. Rev. Genet. 47, 483–508 (2013).
Trojer, P. et al. L3MBTL1, a histone-methylation–dependent chromatin lock. Cell 129, 915–928 (2007).
Yang, Z. et al. Recruitment of P-TEFb for stimulation of transcriptional elongation by the bromodomain protein Brd4. Mol. Cell 19, 535–545 (2005).
West, A.E. & Greenberg, M.E. Neuronal activity–regulated gene transcription in synapse development and cognitive function. Cold Spring Harb. Perspect. Biol. 3, a005744 (2011).
Nam, H.J. et al. Phosphorylation of LSD1 by PKCα is crucial for circadian rhythmicity and phase resetting. Mol. Cell 53, 791–805 (2014).
Lee, C.T. & Duerre, J.A. Changes in histone methylase activity of rat brain and liver with aging. Nature 251, 240–242 (1974).
Evertts, A.G. et al. H4K20 methylation regulates quiescence and chromatin compaction. Mol. Biol. Cell 24, 3025–3037 (2013).
Schaefer, A. et al. Control of cognition and adaptive behavior by the GLP/G9a epigenetic suppressor complex. Neuron 64, 678–691 (2009).
Gupta-Agarwal, S. et al. G9a/GLP histone lysine dimethyltransferase complex activity in the hippocampus and the entorhinal cortex is required for gene activation and silencing during memory consolidation. J. Neurosci. 32, 5440–5453 (2012).
Li, W. et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 498, 516–520 (2013).
Beck, D.B., Oda, H., Shen, S.S. & Reinberg, D. PR-Set7 and H4K20me1: at the crossroads of genome integrity, cell cycle, chromosome condensation, and transcription. Genes Dev. 26, 325–337 (2012).
Laumonnier, F. et al. Mutations in PHF8 are associated with X linked mental retardation and cleft lip/cleft palate. J. Med. Genet. 42, 780–786 (2005).
Wilhelm, B.T. et al. Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 453, 1239–1243 (2008).
Sando, R. et al. HDAC4 governs a transcriptional program essential for synaptic plasticity and memory. Cell 151, 821–834 (2012).
Li, X. et al. Comparison of the effects of the GABAB receptor positive modulator BHF177 and the GABAB receptor agonist baclofen on anxiety-like behavior, learning, and memory in mice. Neuropharmacology 70, 156–167 (2013).
Lee, H.S. et al. Astrocytes contribute to gamma oscillations and recognition memory. Proc. Natl. Acad. Sci. USA 111, E3343–E3352 (2014).
Semenova, S., Contet, C., Roberts, A.J. & Markou, A. Mice lacking the b4 subunit of the nicotinic acetylcholine receptor show memory deficits, altered anxiety- and depression-like behavior, and diminished nicotine-induced analgesia. Nicotine Tob. Res. 14, 1346–1355 (2012).
Bach, M.E., Hawkins, R.D., Osman, M., Kandel, E.R. & Mayford, M. Impairment of spatial but not contextual memory in CaMKII mutant mice with a selective loss of hippocampal LTP in the range of the theta frequency. Cell 81, 905–915 (1995).
Barnes, C.A. Memory deficits associated with senescence: a neurophysiological and behavioral study in the rat. J. Comp. Physiol. Psychol. 93, 74–104 (1979).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Basnet, H. et al. Tyrosine phosphorylation of histone H2A by CK2 regulates transcriptional elongation. Nature 516, 267–271 (2014).
Acknowledgements
We thank J. Chen (University of California San Diego, UCSD) for pLNL vector for LSD1n gene targeting, and S. Wu and M. Capecchi (University of Utah) for pCAG-LSL vector for generation of Lsd1 transgenic mice. We thank J. Zhao and E. Kothari (UCSD transgenic core) for generating knockout and transgenic mice, S. Roberts (The Scripps Research Institute Mouse Behavioral Core) for behavioral assessment, H. Karten (UCSD) for brain anatomy analysis, M. Ghassemian (UCSD) for MALDI-TOF mass spectrometry analysis, A. Gamliel, R. McEvilly, I. Garcia-Bassets, B. Bloodgood and C.K. Glass (UCSD) for discussion, comments, suggestions and critical reading of the manuscript, R. Pardee for proofreading of the manuscript, and J. Hightower for help with figure preparation. J.W. receives funding from a US National Institutes of Health T32 Postdoctoral Fellowship. F.T. was supported by grants from the Roche Extending Innovation Network Program. W.L. was supported by a Department of Defense postdoctoral fellowship. S.L.P. (Benjamin H. Lewis Chair in Neuroscience) and M.G.R. receive funding from the Howard Hughes Medical Institute. This research was supported by grants from the National Institute of Neurological Disorders and Stroke (R37NS5037116) to S.L.P. and by grants from the US National Institutes of Health and the National Cancer Institute (DK018477, NS034934, DK039949, HL065445 and CA173903) to M.G.R. Y.S. received funding from the US National Institutes of Health and the National Institute of General Medical Sciences (R01GM104459-01).
Author information
Authors and Affiliations
Contributions
J.W., F.T., Y.T. and M.G.R. conceived the project. J.W. performed the biochemical characterization of LSD1n, with assistance from C.J., X.H., H.B., Z.L. and X.Z. J.W. generated the murine genetic models, with assistance from H.T. F.T. performed all of the analyses using primary cortical neuronal cultures and helped to coordinate all of the behavioral studies. Y.T. performed GRO-seq experiments and bioinformatics analyses, with assistance from D.M. and Q.M. W.L. performed GRO-seq experiments, and K.O. and J.Z. performed deep-sequencing experiments. R.R.W., C.T. and Y.S. performed gene expression analysis in young and old mice. T.S.M. and S.L.P. performed gene expression analysis in embryonic stem cells.J.W., F.T., Y.T. and M.G.R. wrote the manuscript. All of the authors reviewed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 LSD1n removes H4K20 methylation in vitro
a) Diagram of LSD1 locus, revealing two alternatively-spliced exons: E2a and E8a;
b) RT-PCR detecting alternatively spliced isoforms of LSD1 in mouse embryonic stem cells and Neuro2A cell lines. LSD1 E8a is induced upon RA-induced differentiation of mouse embryonic stem cells;
c) Amino acid sequence showing the alternative splicing events of LSD1 identified in mammals, turtles and fishes;
d) Coomassie blue staining of different isoforms of LSD1 recombinant proteins obtained by two-step affinity purification;
e) In vitro histone demethylase assay on core histones substrates using control (without enzyme), LSD1 WT, E8A, E2A or E2A&8A recombinant proteins in the presence of recombinant CoREST protein. Western blot analysis of histone H3 and H4 methylations revealing the removal of H4K20 methylation specifically by E8A or E2&8A. The amount of specific modification is quantified using ImageJ software; the 2x control lane has an arbitrarily assigned value of 100;
f) In vitro histone demethylase assay on core histones substrates using LSD1n or LSD1m (K685A) mutant recombinant proteins. Western blot analysis of histone H3 and H4 methylations revealing the removal of H4K20 methylation specifically by LSD1n but not by LSD1m mutant. The amount of specific modification is quantified using ImageJ software; the first control lane has an arbitrarily assigned value of 100;
g) In vitro histone demethylase assay on core histones substrates using LSD1n or LSD1m (K685A) mutant recombinant proteins. Western blot analysis of histone H3 and H4 methylations revealing the removal of H4K20 methylation specifically by LSD1n but not by LSD1m mutant. The amount of specific modification is quantified using ImageJ software; the first control lane has an arbitrarily assigned value of 100.
Supplementary Figure 2 LSD1n removes H4K20 methylation in vitro
a) MALDI-TOF results of in vitro histone demethylase assay on H3K4me1 and H3K4me2 peptides substrates using control (without enzyme), LSD1c, LSD1n or LSD1m (K685A mutant) recombinant proteins in the presence of CoREST recombinant protein. Spectra analysis reveals generation of H3K4me1 and/or H3K4me0 from H3K4me1 or H3K4me2 peptides by LSD1c and LSD1n but not by LSD1m;
b) MALDI-TOF results of in vitro histone demethylase assay on H3K9me1 and H3K9me2 peptides substrates. Spectra analysis reveals no enzymatic activity of LSD1c or LSD1n toward H3K9 methylation;
c) MALDI-TOF results of in vitro histone demethylase assay on H4K20me1 and H4K20me2 peptides as substrates. Spectra analysis reveals generation of H4K20me1 and/or H4K20me0 from H4K20me1 or H4K20me2 peptides by LSD1n, but not by LSD1c or LSD1m.
Supplementary Figure 3 Both LSD1n and LSD1c interact with histone H3 and H4 tails
a) Histone peptide pull-down assay using histone H3 and H4 peptides and recombinant LSD1c protein revealing binding between LSD1c and histone H3 or H4 by Western blot using anti-His probe. * Truncated form of His-LSD1c;
b) Histone peptide pull-down assay using histone H3 and H4 peptides and recombinant LSD1n protein, revealing binding between LSD1n and histone H3 or H4 by Western blot using anti-His probe. * Truncated form of His-LSD1n;
c) Histone peptide pull-down assay using histone H3 and H4 peptides and recombinant CoREST protein, revealing specific binding between CoREST and H4 (1-23) by Western blot using anti-His probe;
d) Coomassie staining and Western blot using anti-GST or anti-His probe antibodies, showing expression of the recombinant GST-CoREST-ELM2-His protein;
e) MODified histone peptide array using GST-CoREST-ELM2-His recombinant protein revealing specific interaction between ELM2 domain of CoREST and histone H4 tails by Western blot using anti-GST antibody.
Supplementary Figure 4 Both LSD1n and LSD1c occupy gene promoters and enhancers
a) Schematic diagram of the generation of FLAG-LSD1n and FLAG-LSD1c transgenic mice;
b) Western blot analysis of the expression of FLAG-LSD1 upon Cre-mediated recombination, assessed by Western blot using anti-LSD1 or anti-FLAG antibody;
c) LSD1 genome occupancy in cortical neurons is changed after KCl treatment. Heatmaps display of LSD1 (total), FLAG-LSD1c and FLAG-LSD1n genome occupancies before and after KCl-mediated depolarization, centered on LSD1-peaks in resting cortical neurons. Additional 3kb from the center of the peaks is shown;
d) Box-and-whisker plots of LSD1n/LSD1c tag-density ratio of LSD1 binding on promoters and enhancers revealing similar binding intensity for both isoforms;
e) Western blot analysis revealing decreased H4K20 methylation levels in primary cortical neurons derived from FLAG-LSD1n but not FLAG-LSD1c transgenic mice. The amount of protein is quantified using ImageJ software; the first control lane has an arbitrarily assigned value of 100;
f) Top five enriched transcription factor binding motifs by de novo motif analysis of lost LSD1-peaks on enhancers after KCl treatment, including CTCF motif;
g) Top five enriched transcription factor binding motifs by de novo motif analysis of gained LSD1-peaks on enhancers after KCl treatment, including MEF2c motif;
h) Top five enriched transcription factor binding motifs by de novo motif analysis of gained LSD1-peaks on promoters after KCl treatment, including CREB motif;
i) Immuno-precipitation of brain extracts using LSD1 antibody. Western blot analysis of LSD1-interacting proteins: CoREST, MEF2 and CREB;
j) Table showing the number of overlapping peaks in MEF2 and LSD1 ChIP-seqs;
k) Venn diagram showing combined (resting and depolarized neurons) peaks identified in LSD1 and MEF2 ChIP-seqs;
l) Heatmaps display of LSD1 and MEF2 genome occupancies by ChIP-seqs, centered on LSD1-peaks. Additional 3kb from the centers of the peaks is shown;
m) UCSC genome browser image of Arc locus revealing overlapping peaks from MEF2 and LSD1 ChIP-seqs;
n) UCSC genome browser image of Egr1 locus revealing overlapping peaks from MEF2 and LSD1 ChIP-seqs.
Supplementary Figure 5 LSD1n is required for neuronal activity-regulated gene transcription
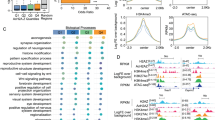
a) Neuronal activity-dependent gene expression is compromised in LSD1 KO cortical neurons assessed by RNA-seq. Box-and-whisker plots of expression values (log2 FPKM, Fragments Per Kilobase Of Exon Per Million Fragments Mapped) of regulated genes in WT and LSD1 KO cortical neurons before and after KCl treatment (6 hours). P-values denote statistic differences between treatment conditions (p=0.46 at 0hr; p=1.7E-6 at 1hr; n=473 is the number of genes up-regulated in WT neurons, unpaired t-test);
b) Upper panel shows RT-qPCR analysis of total LSD1 and LSD1n in LSD1n KO cortical neurons. Data are shown as mean ± SD; P-values denote statistic differences between WT and KO LSD1n (p=0.8027 LSD1; p=0.0115 LSD1n; n=3 technical replicates from pool of 8-12 embryos; unpaired t-test). Lower panel shows RT-PCR revealing that LSD1n KO mice express LSD1c only;
c) Western blot using an antibody against LSD1 show unchanged level of total LSD1 in LSD1n KO mice;
d) Neuronal activity-dependent gene expression is compromised in LSD1n KO cortical neurons assessed by RNA-seq. Box-and-whisker plots of expression values (log2 FPKM) of regulated genes in WT and LSD1n KO cortical neurons before and after KCl treatment (6 hours). P-values denote statistic differences between treatment conditions (p=0.30 at 0hr; p=0.018 at 1hr; n=454 is the number of genes up-regulated in WT neurons; unpaired t-test);
e) Neuronal activity-regulated Arc gene expression is compromised in LSD1n KO cortical neurons assessed by GRO-seq. UCSC genome browser image of Arc locus is shown;
f) Neuronal activity-regulated Egr1 gene expression is compromised in LSD1n KO cortical neurons assessed by GRO-seq. UCSC genome browser image of Egr1 locus is shown;
g) UCSC genome browser image of Arc enhancer revealing bi-directional eRNA expression (eRNA+ and eRNA-);
h) Neuronal activity-regulated eRNA expressions are compromised in LSD1n KO cortical neurons assessed by RT-qPCR. Data are shown as mean ± SD; N.S.= non statistically significant, ** = P-value<0.01 (p=0.3465 (Arc-eRNA KCl-); p=0.0011 (Arc eRNA- KCl+); p=0.2598 (Arc eRNA+, KCl-); p=0.0190 (Arc eRNA+ KCl+); p=0.6038 (Fos eRNA-, KCl-); p=0.0128 (Fos eRNA- KCl+); p=0.2998 (Fos eRNA+, KCl-); p=0.0100 (Fos eRNA+ KCl+); p=0.0120 (Npas4 eRNA+, KCl-); p=0.0001 (Naps4 eRNA+ KCl+); p=0.4110 (Nr4a1 eRNA-, KCl-); p=0.0084 (Nr4a1 eRNA- KCl+); n=4 technical replicates from pool of 8-12 embryos, unpaired t-test);
i) Heatmaps display of LSD1n genome occupancy in cortical neurons before and after KCl treatment (1 hour), centered on TSS sites of neuronal activity-regulated genes. Additional 2kb on each side is shown.
Supplementary Figure 6 LSD1n removes H4K20 methylation in vivo
a) LSD1n does not remove H3K4me1 in vivo. Histogram plots of normalized ChIP-seq tag intensities of H3K4me1 on LSD1n-binding sites at promoters and enhancers of neuronal activity-regulated genes in WT and LSD1n KO cortical neurons after KCl treatment (1 hour);
b) LSD1n does not remove H3K9me2 in vivo. Histogram plots of normalized ChIP-seq tag intensities of H3K9me2 on LSD1n-binding sites at promoters and enhancers of neuronal activity-regulated genes in WT and LSD1n KO cortical after upon KCl treatment (1 hour);
c) LSD1n does not remove H3K36me3 in vivo. Histogram plots of normalized ChIP-seq tag intensities of H3K36me3 on LSD1n-binding sites at promoters and enhancers of neuronal activity-regulated genes in WT and LSD1n KO cortical neurons after KCl treatment (1 hour);
d) LSD1n removes H4K20me1 in vivo. Histogram plots of normalized ChIP-seq tag intensities of H4K20me1 on LSD1n-binding sites at promoters and enhancers of neuronal activity-regulated genes in WT and LSD1n KO cortical neurons after KCl treatment (1 hour);
e) ChIP-qPCR analysis of H3K4me2 level on Npas4 and Arc promoters in WT and LSD1n KO cortical neurons after KCl treatment (1 hour). Data are shown as mean ± SD; N.S. = non statistically significant (p=0.904 Npas4, p=0.494 Arc, n=4 technical replicates from pool of 8-12 embryos; unpaired t-test);
f) ChIP-qPCR analysis of H3K9me2 level on Npas4 and Arc promoters in WT and LSD1n KO cortical neurons after KCl treatment (1 hour). Data are shown as mean ± SD; N.S. = non statistically significant (p=0.247 Npas4, p=0.019 Arc, n=4 technical from pool of 8-12 embryos, unpaired t-test);
g) ChIP-qPCR analysis of H3K79me2 level on Npas4 and Arc promoters in WT and LSD1n KO cortical neurons after KCl treatment (1 hour). Data are shown as mean ± SD; N.S. = non statistically significant (p=0.363 Npas4, p=0.014 Arc, n=4 technical from pool of 8-12 embryos, unpaired t-test);
h) Increased H4K20 methylation in LSD1n KO cortical neurons measured by Western blot. The amount of protein is quantified using ImageJ software; the 1x control lane has an arbitrarily assigned value of 100;
i) Analysis of gene expression of LSD1, Phf8 and Svil in mouse cortical neurons by RT-qPCR (normalized by Actb) and by RNA-seqs. Data are shown as mean ± SD or FPKM value (Fragments Per Kilobase Of Exon Per Million Fragments Mapped);
j) Phf8 knockdown efficiency in cortical neurons using shRNA is assessed by RT-qPCR. Data are shown as mean ± SEM; * = P-value<0.01 (p=0.0074 on minus KCl condition; p=0.0052 on plus KCl condition; n=3 biological replicates/condition; unpaired t-test);
k) Neuronal activity-regulated gene expression is not significantly affected upon Phf8 knockdown. Gene expression of Arc, Btg2, Cyr61, Egr3, Npas4 and Pcsk1 in control or Phf8 deficient cortical neurons is measured by RT-qPCR. Values are normalized against Actb. Data are shown as mean ± SEM; N.S. = non statistically significant (p=0.4562 Arc; p=0.3708 Btg2; p=0.1894 Cyr61; p=0.3875 Egr3; p=0.2828 Npas4; p=0.3663 Pcsk1; n=3 biological replicates/condition; unpaired t-test).
Supplementary Figure 7 LSD1n promotes transcriptional elongation
a) Arc gene expressions is regulated at LSD1n-dependent transcriptional elongation step assessed by RNA Pol II ChIP-seq. UCSC genome browser image of Arc locus is shown;
b) Egr1 gene expression is regulated at LSD1n-dependent transcriptional elongation step assessed by Pol II ChIP-seq. UCSC genome browser image of Egr1 locus is shown;
c) Traveling ratio plots of fraction of neuronal activity-regulated genes assessed by GRO-seq in WT and LSD1n KO cortical neurons showing that neuronal activity-dependent gene expression is regulated by LSD1n at the transcriptional elongation step; P-value denotes statistic difference between treatments (p=0.00192; two-tailed KS test);
d) Analysis of RNA Pol II recruitments by ChIP assay on Arc 5’ region (TSS). Data are shown as mean ± SD; N.S.= non statistically significant, **= P-value<0.01 (p=0.000461591; p=0.130184533; n=4 technical replicates from pool of 8-12 embryos; unpaired t-test);
e) Analysis of RNA Pol II recruitments by ChIP assay on Arc 3’ region (coding region). Data are shown as mean ± SD; ** P-value<0.01 (p=0.00016313; p=0.000237331; n=4 technical replicates from pool of 8-12 embryos; unpaired t-test);
f) RNA Pol II ChIP analysis revealing that elongation of RNA Pol II is compromised (decreased 3’/5’ ratio) in LSD1n KO cortical neurons after KCl treatment (1 hour). Data are shown as mean ± SD; ** = P-value<0.01 (p=0.0129; p=0.0012; n=4 technical replicates from pool of 8-12 embryos; unpaired t-test);
g) Analysis of RNA Pol II recruitment by ChIP assay on Egr1 5’ region (TSS). Data are shown as mean ± SD; ** = P-value<0.01 (p=1.7E-5; p=0.0003; n= 4 technical replicates from pool of 8-12 embryos; unpaired t-test);
h) Analysis of RNA Pol II recruitment by ChIP assay on Egr1 3’ region (coding region). Data are shown as mean ± SD; ** = P-value<0.01 (p=6.0E-7; p=1.3E-7; n=4 technical replicates from pool of 8-12 embryos; unpaired t-test);
i) RNA Pol II ChIP analysis revealing elongation of RNA Pol II is compromised (decreased 3’/5’ ratio) in LSD1n KO cortical neurons after KCl treatment (1 hour). Data are shown as mean ± SD; ** = P-value<0.01 (p=0.0015; p=0.0099; n=4 technical replicates from pool of 8-12 embryos; unpaired t-test);
j) Neuronal activity-regulated gene expression is modulated by elongation control. RNA Pol II traveling ratio plot assessed by RNA Pol II ChIP-seq for the fraction of neuronal activity-regulated genes before and after KCl treatment (1 hour);
k) ChIP-qPCR analysis of H4K16Ac level on Npas4 and Arc promoters in WT and LSD1n KO cortical neurons. Data are shown as mean ± SD; (p=0.040 Npas4; p=0.006 Arc; n=4 technical replicates from pool of 8-12 embryos; unpaired t-test);
l) ChIP-qPCR analysis of L3MBTL1 occupancy on Npas4 and Arc promoters in WT and LSD1n KO cortical neurons. Data are shown as mean ± SD; (p=0.002 Npas4; p=0.007 Arc; n=3 technical replicates from pool of 8-12 embryos; unpaired t-test);
m) ChIP-qPCR analysis of Brd4 occupancy on Npas4 and Arc promoters in WT and LSD1n KO cortical neurons. Data are shown as mean ± SD; (p=0.002 Npas4; p=0.0001 Arc; n=3 technical replicates from pool of 8-12 embryos; unpaired t-test);
n) ChIP-seq analysis of FLAG-LSD1n showing that LSD1n recruitment is enriched on transcribed coding regions of neuronal activity-regulated genes after KCl treatment (1 hour);
o) ChIP-seq analysis of FLAG-LSD1c showing that LSD1c recruitment is not enriched on transcribed coding regions of neuronal activity-regulated genes after KCl treatment (1 hour);
p) Heatmaps display of LSD1n and LSD1c genome occupancies on transcribed coding regions of neuronal activity-regulated genes, revealing that LSD1n but not LSD1c is recruited to coding regions after neuron KCl treatment (1 hour).
Supplementary Figure 8 LSD1n is required for learning and memory
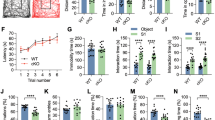
a) Analysis of brain anatomy of WT and LSD1n KO mice using Nissl stain, revealing normal brain morphology of LSD1n KO mice;
b) Optomotor behavioral test of male or female WT and LSD1n KO mice revealing normal vision in LSD1n KO mice. Data are shown as mean ± SD; N.S. = non statistically significant (p=0.111), ANOVA test (n=10);
c) Percentage of spontaneous alternation (Y maze) showing normal working memory in LSD1n KO mice. Data are shown as mean ± SD; N.S. = non statistically significant (p=0.7449); ANOVA test (n=10);
d) Number of total arm entries (Y maze) showing normal working memory in LSD1n KO mice. Data are shown as mean ± SD; N.S. = non statistically significant (p=0.0550); ANOVA test (n=10);
e) Normal locomotor activity in LSD1n KO mice assessed by ambulation counts in dark and light cycles (n=10);
f) Western blot analysis showing increased H4K20 methylation in cortex of LSD1n knockout mice. The amount of protein is quantified using ImageJ software, the first control lane has an arbitrarily assigned value of 100;
g) RT-PCR analysis showing decreased expression of LSD1n in 28-month-old mice (Old) compared to 4-month-old mice (Young);
h) RT-qPCR showing decreased expression of LSD1n in 28-month-old mice (Old) compared to 4-month-old mice (Young). Data are shown as mean ± SD; N.S. = non statistically significant, ** = P-value<0.01 (p=0.1896 LSD1; p=0.0001 LSD1c; p=3.7E-5 LSD1n; n=3 technical replicates from young or old mice; unpaired t-test).
Supplementary Figure 9 Model: LSD1n controls neuronal activity-dependent gene transcription essential for learning and memory
Neuronal activity-induced recruitment of LSD1n to MEF2- and CREB-bound enhancers and promoters of neuronal activity-regulated genes, which leads to removal of H4K20 methylations on enhancers, promoters and transcribed coding regions to promote gene transcription initiation and elongation.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 (PDF 2688 kb)
Supplementary Methods Checklist
(PDF 488 kb)
Rights and permissions
About this article
Cite this article
Wang, J., Telese, F., Tan, Y. et al. LSD1n is an H4K20 demethylase regulating memory formation via transcriptional elongation control. Nat Neurosci 18, 1256–1264 (2015). https://doi.org/10.1038/nn.4069
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4069
This article is cited by
-
LSD1/PRMT6-targeting gene therapy to attenuate androgen receptor toxic gain-of-function ameliorates spinobulbar muscular atrophy phenotypes in flies and mice
Nature Communications (2023)
-
Histone demethylase Lsd1 is required for the differentiation of neural cells in Nematostella vectensis
Nature Communications (2022)
-
Roles of lysine-specific demethylase 1 (LSD1) in homeostasis and diseases
Journal of Biomedical Science (2021)
-
LSD1 inhibition sustains T cell invigoration with a durable response to PD-1 blockade
Nature Communications (2021)
-
LSD1: more than demethylation of histone lysine residues
Experimental & Molecular Medicine (2020)