Abstract
The human cerebral cortex depends for its normal development and size on a precisely controlled balance between self-renewal and differentiation of diverse neural progenitor cells. Specialized progenitors that are common in humans but virtually absent in rodents, called outer radial glia (ORG), have been suggested to be crucial to the evolutionary expansion of the human cortex. We combined progenitor subtype–specific sorting with transcriptome-wide RNA sequencing to identify genes enriched in human ORG, which included targets of the transcription factor neurogenin and previously uncharacterized, evolutionarily dynamic long noncoding RNAs. Activating the neurogenin pathway in ferret progenitors promoted delamination and outward migration. Finally, single-cell transcriptional profiling in human, ferret and mouse revealed more cells coexpressing proneural neurogenin targets in human than in other species, suggesting greater neuronal lineage commitment and differentiation of self-renewing progenitors. Thus, we find that the abundance of human ORG is paralleled by increased transcriptional heterogeneity of cortical progenitors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Florio, M. & Huttner, W.B. Neural progenitors, neurogenesis and the evolution of the neocortex. Development 141, 2182–2194 (2014).
Gorski, J.A. et al. Cortical excitatory neurons and glia, but not GABAergic neurons, are produced in the Emx1-expressing lineage. J. Neurosci. 22, 6309–6314 (2002).
Kowalczyk, T. et al. Intermediate neuronal progenitors (basal progenitors) produce pyramidal-projection neurons for all layers of cerebral cortex. Cereb. Cortex 19, 2439–2450 (2009).
Smart, I.H.M., Dehay, C., Giroud, P., Berland, M. & Kennedy, H. Unique morphological features of the proliferative zones and postmitotic compartments of the neural epithelium giving rise to striate and extrastriate cortex in the monkey. Cereb. Cortex 12, 37–53 (2002).
Fietz, S.A. et al. OSVZ progenitors of human and ferret neocortex are epithelial-like and expand by integrin signaling. Nat. Neurosci. 13, 690–699 (2010).
Hansen, D.V., Lui, J.H., Parker, P.R.L. & Kriegstein, A.R. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 464, 554–561 (2010).
Betizeau, M. et al. Precursor diversity and complexity of lineage relationships in the outer subventricular zone of the primate. Neuron 80, 442–457 (2013).
Gertz, C.C., Lui, J.H., LaMonica, B.E., Wang, X. & Kriegstein, A.R. Diverse behaviors of outer radial glia in developing ferret and human cortex. J. Neurosci. 34, 2559–2570 (2014).
Reillo, I., de Juan Romero, C., García-Cabezas, M.Á. & Borrell, V. A role for intermediate radial glia in the tangential expansion of the mammalian cerebral cortex. Cereb. Cortex 21, 1674–1694 (2011).
Martínez-Cerdeño, V. et al. Comparative analysis of the subventricular zone in rat, ferret and macaque: evidence for an outer subventricular zone in rodents. PLoS ONE 7, e30178 (2012).
Johnson, M.B. et al. Functional and evolutionary insights into human brain development through global transcriptome analysis. Neuron 62, 494–509 (2009).
Kang, H.J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 (2011).
Fietz, S.A. et al. Transcriptomes of germinal zones of human and mouse fetal neocortex suggest a role of extracellular matrix in progenitor self-renewal. Proc. Natl. Acad. Sci. USA 109, 11836–11841 (2012).
Miller, J.A. et al. Transcriptional landscape of the prenatal human brain. Nature 508, 199–206 (2014).
Lui, J.H. et al. Radial glia require PDGFD-PDGFRβ signalling in human but not mouse neocortex. Nature 515, 264–268 (2014).
Capela, A. & Temple, S. LeX is expressed by principle progenitor cells in the embryonic nervous system, is secreted into their environment and binds Wnt-1. Dev. Biol. 291, 300–313 (2006).
Mo, Z. et al. Human cortical neurons originate from radial glia and neuron-restricted progenitors. J. Neurosci. 27, 4132–4145 (2007).
Shibata, T. et al. Glutamate transporter GLAST is expressed in the radial glia-astrocyte lineage of developing mouse spinal cord. J. Neurosci. 17, 9212–9219 (1997).
Weigmann, A., Corbeil, D., Hellwig, A. & Huttner, W.B. Prominin, a novel microvilli-specific polytopic membrane protein of the apical surface of epithelial cells, is targeted to plasmalemmal protrusions of non-epithelial cells. Proc. Natl. Acad. Sci. USA 94, 12425–12430 (1997).
Uchida, N. et al. Direct isolation of human central nervous system stem cells. Proc. Natl. Acad. Sci. USA 97, 14720–14725 (2000).
Seo, S., Lim, J.-W., Yellajoshyula, D., Chang, L.-W. & Kroll, K.L. Neurogenin and NeuroD direct transcriptional targets and their regulatory enhancers. EMBO J. 26, 5093–5108 (2007).
Gohlke, J.M. et al. Characterization of the proneural gene regulatory network during mouse telencephalon development. BMC Biol. 6, 15 (2008).
Ochiai, W. et al. Periventricular notch activation and asymmetric Ngn2 and Tbr2 expression in pair-generated neocortical daughter cells. Mol. Cell. Neurosci. 40, 225–233 (2009).
Rousso, D.L. et al. Foxp-mediated suppression of N-cadherin regulates neuroepithelial character and progenitor maintenance in the CNS. Neuron 74, 314–330 (2012).
Kovach, C. et al. Neurog2 simultaneously activates and represses alternative gene expression programs in the developing neocortex. Cereb. Cortex 23, 1884–1900 (2013).
Iacopetti, P. et al. Expression of the antiproliferative gene TIS21 at the onset of neurogenesis identifies single neuroepithelial cells that switch from proliferative to neuron-generating division. Proc. Natl. Acad. Sci. USA 96, 4639–4644 (1999).
Kawaguchi, A. et al. Differential expression of Pax6 and Ngn2 between pair-generated cortical neurons. J. Neurosci. Res. 78, 784–795 (2004).
Reid, C.B., Tavazoie, S.F. & Walsh, C.A. Clonal dispersion and evidence for asymmetric cell division in ferret cortex. Development 124, 2441–2450 (1997).
Ware, M.L., Tavazoie, S.F., Reid, C.B. & Walsh, C.A. Coexistence of widespread clones and large radial clones in early embryonic ferret cortex. Cereb. Cortex 9, 636–645 (1999).
Jackson, C.A., Peduzzi, J.D. & Hickey, T.L. Visual cortex development in the ferret. I. Genesis and migration of visual cortical neurons. J. Neurosci. 9, 1242–1253 (1989).
Noctor, S.C., Scholnicoff, N.J. & Juliano, S.L. Histogenesis of ferret somatosensory cortex. J. Comp. Neurol. 387, 179–193 (1997).
Borrell, V. In vivo gene delivery to the postnatal ferret cerebral cortex by DNA electroporation. J. Neurosci. Methods 186, 186–195 (2010).
Treutlein, B. et al. Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375 (2014).
Jaitin, D.A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Necsulea, A. et al. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature 505, 635–640 (2014).
Ng, S.-Y., Bogu, G.K., Soh, B.S. & Stanton, L.W. The long noncoding RNA RMST interacts with SOX2 to regulate neurogenesis. Mol. Cell 51, 349–359 (2013).
Sauvageau, M. et al. Multiple knockout mouse models reveal lincRNAs are required for life and brain development. eLife 2, e01749 (2013).
Hu, P. et al. LncRNA expression signatures of twist-induced epithelial-to-mesenchymal transition in MCF10A cells. Cell. Signal. 26, 83–93 (2014).
Cabili, M.N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 25, 1915–1927 (2011).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Aprea, J. et al. Transcriptome sequencing during mouse brain development identifies long non-coding RNAs functionally involved in neurogenic commitment. EMBO J. 32, 3145–3160 (2013).
Kelava, I. et al. Abundant occurrence of basal radial glia in the subventricular zone of embryonic neocortex of a lissencephalic primate, the common marmoset Callithrix jacchus. Cereb. Cortex 22, 469–481 (2012).
Lewitus, E., Kelava, I., Kalinka, A.T., Tomancak, P. & Huttner, W.B. An adaptive threshold in mammalian neocortical evolution. PLoS Biol. 12, e1002000 (2014).
Franco, S.J. et al. Fate-restricted neural progenitors in the mammalian cerebral cortex. Science 337, 746–749 (2012).
Guo, C. et al. Fezf2 expression identifies a multipotent progenitor for neocortical projection neurons, astrocytes, and oligodendrocytes. Neuron 80, 1167–1174 (2013).
Kawaguchi, A. et al. Single-cell gene profiling defines differential progenitor subclasses in mammalian neurogenesis. Development 135, 3113–3124 (2008).
Florio, M. et al. Human-specific gene ARHGAP11B promotes basal progenitor amplification and neocortex expansion. Science doi:10.1126/science.aaa1975 (26 February 2015).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Hiller, M. et al. A 'forward genomics' approach links genotype to phenotype using independent phenotypic losses among related species. Cell Reports 2, 817–823 (2012).
Hubisz, M.J., Pollard, K.S. & Siepel, A. PHAST and RPHAST: phylogenetic analysis with space/time models. Brief. Bioinform. 12, 41–51 (2011).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet. 16, 276–277 (2000).
Acknowledgements
We thank J. Partlow for coordinating human tissue protocols, D. Gonzalez for animal protocol and technical experimental assistance, S. Lazo-Kallanian for single-cell FACS assistance and all members of the Walsh laboratory for comments and discussion. The Neurog2-VP16 construct was a generous gift from C. Schuurmans (University of Calgary). This work was supported by grants to C.A.W. from the US National Institutes of Neurological Disease and Stroke (R01 NS032457) and the Paul G. Allen Family Foundation. M.B.J. was supported by a fellowship from the Nancy Lurie Marks Family Foundation. P.P.W. was supported by the Stuart H.Q. & Victoria Quan Fellowship at Harvard Medical School. Single-cell expression profiling experiments were performed at the Molecular Genetics Core at Boston Children's Hospital (BCH IDDRC, P30 HD18655). Transcriptome analysis was performed using Harvard Medical School's Orchestra high-performance computing cluster, which is partially supported by US National Institutes of Health grant NCRR 1S10RR028832-01. C.A.W. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
M.B.J., P.P.W. and R.N.D. designed and conducted experiments and analyzed data. K.D.A. and E.A.M. performed experiments and analyzed data. J.L.H. procured and examined human tissue samples. M.B.J., P.P.W. and C.A.W. interpreted the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Isolation of human RGC subpopulations by FACS
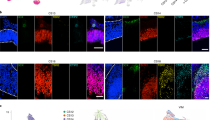
a, Both LG+Prhi and LG+Prlo subpopulations are enriched for known RGC-expressed genes (GFAP, VIM, GLAST, PAX6, SOX2, BLBP), and depleted for neuronal markers (DCX, TUJ1, NeuN, MEF2C). The LG+Prhi subpopulation was enriched relative to the LG+Prlo subpopulation for PROM1 transcript as well as three other transcripts encoding apical membrane domain-specific proteins (PARD3 [Par3], TJP1 [ZO-1], MPP5 [Pals]). Data represents four biological replicates (mean ± SEM) ranging from 16 WG to 23 WG. b, Primary neurospheres derived from LeX− and LeX+ cells sorted from dissociated human fetal cortex. Neurospheres were serially passaged at clonal density and immunolabeled for RGC marker SOX2.
Supplementary Figure 2 Gene set enrichment in human RGC subpopulations
Gene set enrichment analysis confirmed the RGC progenitor nature of both the LG+Prhi and LG+Prlo subpopulations, with enrichment of important progenitor signaling pathways (e.g. Wnt/Bmp/Tgf) and gene ontology terms (cell cycle control, neural development) in both subpopulations relative to LG−Pr− neurons and other cell types.
Supplementary Figure 3 Selected LG+Prlo-enriched candidate non-apical RGC genes validated by qRT-PCR in independent biological replicates of FACS-purified human fetal RGC.
Relative expression levels in the LG+Prlo subpopulation compared to LG+Prhi after normalization to housekeeping genes ACTB and GAPDH. Data represents six biological replicates (mean ± SEM) ranging from 16 WG to 23 WG (asterisk denotes p < 0.05, paired t-test; n=6, max p=0.045, all others were lower).
Supplementary Figure 4 Upregulation of proneural neurogenin targets in NEUROG2-VP16 electroporated ferret cells
In vivo delivery of GFP control and NEUROG2-VP16 constructs to ferret apical RGCs was performed by intraventricular injection and electroporation in neonatal ferret kits (n=2 per condition at postnatal day 1) as described in Figure 2. After 48 hours post-electroporation, electroporated cells were isolated for qRT-PCR analysis by enzymatic dissociation and FACS using their GFP fluorescence. Relative to GFP+ control electroporated cells, NEUROG2-VP16 expressing cells showed upregulation of many previously described NEUROG2 effector genes including Cbfa2t2, Foxn2, Foxp2, Hes6, Myt1, Neurod1, Neurod4, Neurog1, and Nhlh1, and down-regulation of Sox2. In addition, we also tested expression of ferret orthologs of human ORG-enriched genes and found that several including Gadd45g, Ttyh2, Sstr2, and Plcb4 were also upregulated in NEUROG2-VP16 cells compared to controls.
Supplementary Figure 5 Single-cell expression profiles of human and mouse RGC
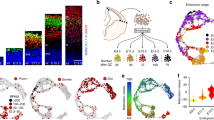
a, Violin plots of RGC marker gene expression in human and mouse single sorted RGC reveals largely similar pattern of gene expression for RGC markers including SOX2, VIM, GLAST, BLBP, PAX6, NES. Interestingly, significant numbers of human RGC express GFAP and DCX but these genes are nearly absent in mouse RGC. b, Principle component analysis of 546 human (left) and 226 mouse (right) single RGC indicates distinct distributions of transcriptional states in human compared to mouse RGC. Here, “apical” is defined by expression of at least two of the four apical complex marker transcripts, and “proneural” by expression of at least two of the four Neurogenin pathway genes. In both species, the first PC (x-axis) reflects the proneural+/− dimension, with “multipotent” (presumptively pre-Neurogenin-pathway-expressing) RGC tending towards the left (red and blue cells) and proneural RGC on the right (black and green cells). Human cortex contains a greater proportion of proneural RGC, whereas mouse has fewer proneural cells which are less distinct, as indicated by the greater overlap of black and red cells in the mouse. In addition, human cortex displays far more non-apical (blue and green) cells than mouse, which again are more distinct from the apical (red and black) cells along the second PC (y-axis). In contrast, mouse non-apical RGC (blue and green) are scarce and not transcriptionally distinct from apical cells, as indicated by the lack of separation along the y-axis.
Supplementary Figure 6 Differential expression of novel unannotated lncRNAs in human RGC subtypes
RNA-seq reads displayed in genomic context for the LG+Prhi apical RGC (red), LG+Prlo ORG (green), and LG−Pr− cells (black). Novel transcripts assembled from the RNA-seq data are shown in blue, and previously catalogued lncRNA transcripts are shown in brown39. a, Two intergenic lncRNAs on chromosome 2 with distinct expression patterns in the human fetal cortex share a bidirectional promoter and overlap at their 5′ ends. The plus-strand lncRNA is enriched in apical RGC, whereas the minus-strand lncRNA is relatively enriched in ORG and neurons. Blue boxed region highlights the overlapping transcription start sites (TSS), and is enlarged below. Black arrows indicate read peaks from each strand's TSS. Bottom part of (a) shows the promoter at higher magnification, with expression levels of the two lncRNAs (in FPKM) plotted at right. b, Example of an ORG-enriched lncRNA. Multiple alternatively spliced isoforms of this multi-exon locus are expressed in all cell types assayed, but are significantly enriched in the LG+Prlo non-apical subpopulation. A partial transcript overlapping the 5′ end of the locus was previously detected by ultra-high depth RNA sequencing39; our data demonstrate that even low-abundance transcripts can be captured and fully reconstructed from an order of magnitude fewer reads when RNA is sequenced from the specific cell types that express the gene, rather than from heterogeneous bulk tissue. c, Example of a novel apical RGC-specific intergenic transcript not detected by previous deep-sequencing experiments.
Supplementary Figure 7 Differential enrichment of lncRNAs in human and mouse RGC populations
We performed qRT-PCR of several conserved lncRNAs in FACS-purified human (n=4 biological replicates ranging from 16 WG to 23 WG) and mouse RGC populations (n=3 from E15.5) comparing human ORG (LG+Prlo) and apical RGC (LG+Prhi) with neurons (LG−Pr−) and mouse RGC (L+Pr+) with neurons (L−Pr−) (mean ± SEM). We find that several conserved lncRNAs including LINC-PINT, TUNAR, CRNDE, MIR22HG are enriched in human RGC progenitor populations but depleted in mouse RGC suggesting potentially divergent roles in human radial progenitor evolution and function.
Supplementary information
Supplementary Figures and Tables
Supplementary Figures 1–7 and Supplementary Tables 1–3 (PDF 5788 kb)
Supplementary Methods Checklist
Reporting Checklist for Nature Neuroscience (PDF 116 kb)
Rights and permissions
About this article
Cite this article
Johnson, M., Wang, P., Atabay, K. et al. Single-cell analysis reveals transcriptional heterogeneity of neural progenitors in human cortex. Nat Neurosci 18, 637–646 (2015). https://doi.org/10.1038/nn.3980
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3980
This article is cited by
-
The interaction between ageing and Alzheimer's disease: insights from the hallmarks of ageing
Translational Neurodegeneration (2024)
-
Functional synergy of a human-specific and an ape-specific metabolic regulator in human neocortex development
Nature Communications (2024)
-
Sorting Technology for Mesenchymal Stem Cells from a Single Tissue Source
Stem Cell Reviews and Reports (2024)
-
Astroblastomas exhibit radial glia stem cell lineages and differential expression of imprinted and X-inactivation escape genes
Nature Communications (2022)
-
Brain organoid: a 3D technology for investigating cellular composition and interactions in human neurological development and disease models in vitro
Stem Cell Research & Therapy (2021)