Abstract
Hippocampal place cells assemblies are believed to support the cognitive map, and their reactivations during sleep are thought to be involved in spatial memory consolidation. By triggering intracranial rewarding stimulations by place cell spikes during sleep, we induced an explicit memory trace, leading to a goal-directed behavior toward the place field. This demonstrates that place cells' activity during sleep still conveys relevant spatial information and that this activity is functionally significant for navigation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
O'Keefe, J. & Nadel, L. The Hippocampus as a Cognitive Map 1–570 (Oxford University Press, 1978).
Lee, A.K. & Wilson, M.A. Neuron 36, 1183–1194 (2002).
Wilson, M.A. & McNaughton, B.L. Science 265, 676–679 (1994).
Buzsáki, G. Neuroscience 31, 551–570 (1989).
Girardeau, G., Benchenane, K., Wiener, S.I., Buzsáki, G. & Zugaro, M.B. Nat. Neurosci. 12, 1222–1223 (2009).
Ego-Stengel, V. & Wilson, M.A. Hippocampus 20, 1–10 (2010).
Lenck-Santini, P.-P., Muller, R.U., Save, E. & Poucet, B. J. Neurosci. 22, 9035–9047 (2002).
Jeffery, K.J., Gilbert, A., Burton, S. & Strudwick, A. Hippocampus 13, 175–189 (2003).
Delgado, J.M.R., Johnston, V.S., Wallace, J.D. & Bradley, R.J. Brain Res. 22, 347–362 (1970).
Shinkman, P.G., Bruce, C.J. & Pfingst, B.E. Science 184, 1194–1196 (1974).
Jackson, A., Mavoori, J. & Fetz, E.E. Nature 444, 56–60 (2006).
Fetz, E.E. Science 163, 955–958 (1969).
Eichenbaum, H. Behav. Brain Res. 103, 123–133 (1999).
Carlezon, W.A. & Chartoff, E.H. Nat. Protoc. 2, 2987–2995 (2007).
Kobayashi, T., Nishijo, H., Fukuda, M., Bures, J. & Ono, T. J. Neurophysiol. 78, 597–613 (1997).
Dickinson, A. & Balleine, B.W. Anim. Learn. Behav. 22, 1–18 (1994).
Hennevin, E. & Hars, B. Psychobiology (Austin, Tex.) 20, 166–176 (1992).
Weinberger, N.M., Gold, P.E. & Sternberg, D.B. Science 223, 605–607 (1984).
Arzi, A. et al. Nat. Neurosci. 15, 1460–1465 (2012).
Pavlides, C. & Winson, J. J. Neurosci. 9, 2907–2918 (1989).
Carlezon, W.A. & Chartoff, E.H. Nat. Protoc. 2, 2987–2995 (2007).
McNaughton, N. & Sedgwick, E.M. Neuroscience 3, 629–632 (1978).
Paxinos, G. & Bindra, D. Physiol. Behav. 5, 227–231 (1970).
O'Keefe, J. & Nadel, L. The Hippocampus as a Cognitive Map 1–570 (Oxford University Press, 1978).
Harris, K.D., Henze, D.A., Csicsvari, J., Hirase, H. & Buzsáki, G. J. Neurophysiol. 84, 401–414 (2000).
Hazan, L., Zugaro, M.B. & Buzsáki, G. J. Neurosci. Methods 155, 207–216 (2006).
Schmitzer-Torbert, N., Jackson, J., Henze, D., Harris, K. & Redish, A.D. Neuroscience 131, 1–11 (2005).
Delgado, J.M.R., Johnston, V.S., Wallace, J.D. & Bradley, R.J. Brain Res. 22, 347–362 (1970).
Shinkman, P.G., Bruce, C.J. & Pfingst, B.E. Science 184, 1194–1196 (1974).
Fetz, E.E. & Finocchio, D.V. Science 174, 431–435 (1971).
Jackson, A., Mavoori, J. & Fetz, E.E. Nature 444, 56–60 (2006).
Harris, K.D., Hirase, H., Leinekugel, X., Henze, D.A. & Buzsáki, G. Neuron 32, 141–149 (2001).
Muller, R.U., Kubie, J.L. & Ranck, J.B.J. J. Neurosci. 7, 1935–1950 (1987).
Maei, H.R., Zaslavsky, K., Teixeira, C.M. & Frankland, P.W. Front. Integr. Neurosci. 3, 4 (2009).
Nieuwenhuis, S., Forstmann, B.U. & Wagenmakers, E. Nat. Neurosci. 14, 1105–1107 (2011).
Acknowledgements
We thank F.P. Battaglia, M.B. Zugaro, P. Faure, L. Roux, S. Bagur and S.I. Wiener for discussions, and A. Peyrache for discussion and critical readings of an earlier version of the manuscript. Some experiments were done with software developed by G. Cournelle and A. Baelde, engineering students from the ESPCI-ParisTech. This work was supported by the Fondation pour la Recherche Médicale DEQ20120323730, France, by the National Agency for Research ANR-REG-071220-01-01, France, by the CNRS: Programmes Interdisciplinaires de Recherche (Neuro-IC), ATIP-Avenir (2014) and by the city of Paris (Grant Emergence 2014). This work also received support under the program Investissements d'Avenir launched by the French Government and implemented by the ANR, with the references: ANR-10-LABX-54 MEMO LIFE and ANR-11-IDEX-0001-02 PSL* Research University. G.d.L. and M.M.L. were funded by the Ministère de l'Enseignement Supérieur et de la Recherche, France.
Author information
Authors and Affiliations
Contributions
K.B. designed the experiment. G.d.L. and M.M.L. did the experiments. K.B., G.d.L. and M.M.L. analyzed the data. K.B., G.d.L., M.M.L. and L.R.-R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 The place preference tasks.
a. Wake pairing protocol. The paradigm consisted in four sessions: 1) free exploration and place field identification, 2) basal exploration (PRE), 3) 15 minutes exploration with pairing where online detected spikes of a given place cell triggered MFB stimulation, and 4) test exploration (POST). In PRE and POST session a minimum of eight 60s—trials with changing starting point (gray circles) is performed. Analyses were carried on the first 4 trials where animal was actively exploring. b. Sleep pairing protocol. As in panel a, the sleep protocol is composed of the same four sessions except that spike—stimulation pairing is performed during 1h of sleep. This protocol was always followed by free exploration to erase the learning (extinction exploration) and a control wake pairing. c. Classical Place preference task. Mouse position was tracked in real time with a video camera during a session lasting 15 minutes. Rewarding stimulations were then delivered when mice reached a pre—established area, covering around 5% of the total surface of exploration. This stimulation phase was preceded by a basal exploration (PRE) and followed by a probe exploration (POST) as described in panel a. d. Method for the choice of the 4 starting points used in PRE and POST sessions of all place preference paradigms.
Supplementary Figure 2 Characteristics of sleep pairing sessions with rewarding and non-rewarding MFB stimulations, showing similar properties and absence of sleep perturbation.
a—b. Power spectrum averaged on 3s—period before MFB stimulations train (black), and on 3s period (starting 1s after stimulation train onset to avoid electrical artifact) after rewarding stimulation (a, red) or non—rewarding stimulation (b, blue) during sleep pairing sessions. From left to right: row power spectrum, spectrum normalized by 1/f and log scaled power spectrum. Colored areas represent SEM. c. Percentage of time spent in REM during the total sleep period of the sleep pairing experiment, with either rewarding (grey, n=5) or non—rewarding (black, n=2) stimulations. d. Number of stimulation in SWS and REM (Wilcoxon matched pairs test, rewarding condition, n=5, Z=2.02, P=0.043). e. Frequency of rewarding and non—rewarding stimulations during the first 10min (beginning) and the last 10min (end) of the sleep pairing sessions (Wilcoxon matched pairs test, rewarding condition, n=5, Z=0.13, P=0.89). This shows that the number of place cell—triggered stimulations is not modified along the pairing protocol, excluding an operant conditioning of the neurons. f—g. Sparsity (f) and spatial information (g) of online and offline detected place cells used to trigger rewarding or non—rewarding stimulations, in sleep and wake—pairing protocols. Wilcoxon matched pairs test, rewarding condition, n=10, f: Z=1.58, P=0.11; g: Z=1.27, P=0.20.
* P <0.05, NS: non significant. Error bars indicate SEM.
Supplementary Figure 3 Place cells activity and theta oscillations remain stable after MFB stimulations.
a. Place fields of all online detected place cells used to trigger stimulation during sleep (left), and of all other simultaneously recorded place cells (right), during exploration of the same environment before (PRE) or after (POST) the sleep pairing protocol. Color scale indicates firing rate, maximum being indicated above the map. b. Firing map and occupancy map before and during the control wake pairing protocol. c. Exploration sessions with classical remapping of place cells when the mouse explores two different environments. d. Quantification of place field stability by correlation analyses in the three situations shown in panels a—c: Sleep pairing: correlation of firing maps obtained in exploration sessions performed before and after the sleep—pairing procedure (panel a); Wake pairing: correlation of firing maps obtained before and during the wake—pairing procedure (panel b); Control remapping session: correlation of firing maps obtained in two different environements (panel c). (Mann & Whitney, Sleep—pairing session vs. Control remapping session, n=11, n=5, Z=2.77, P=0.005; Wake Pairing session vs. Control remapping session n=10, n=5, Z=2.59, P=0.0094; Sleep Pairing session vs. Wake Pairing session n=11, n=10, Z=0.53, P=0.59). ** P <0.01, NS: non significant.
e—f. Instantaneous phase distribution of theta filtered LFP 500ms before MFB stimulations trains and 500ms after stimulation artifacts, with stimulation either triggered manually during the classical place preference (e) or by a theta modulated place cell during wake—pairing protocol (f). For phase and kappa value, black and red dots indicate a significant locking of the phase (different from random phase distribution). The p value corresponds to the statistical difference between the observed distribution of the phases of the LFP filtered in the Theta band before and after the stimulation and a uniform circular distribution.
These panels show that in our conditions, MFB stimulation did not modify theta oscillations during exploration. This is an important control since stimulations in the hypothalamic region can drive (or reset) theta oscillations22,23, Also MFB stimulation delivered either during sleep— or wake—pairing protocols, induced no modification of the place fields’ properties, as correlations were significantly higher compared to an additional session where mice are placed in two different environments causing place cells remapping. Finally, these panels must be related to the main result of the paper: rewarding stimulations triggered by spikes of a place cell could induce place preference (Fig. 2a), further supporting the absence of modification of place fields by MFB stimulations as previously observe by Kobayashi and colleagues15.
Supplementary Figure 4 MFB stimulation does not alter sleep sharp-wave ripples (SPW-Rs), nor associated neuronal activity.
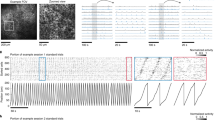
a. Sleep SPW—Rs mean amplitude, duration and frequency with co—occurrence of a single MFB stimulation (red) or without (black). Wilcoxon matched pairs test, SPW—Rs with stimulation: n=35, SPW—Rs without stimulation: n=41; Amplitude, Z=1.20, P=0.30, Duration: Z=1.20, P=0.22, Frequency: Z=0.98, P=0.31. b. Averaged hippocampal LFP at time of SPW—Rs with concomitant single MFB stimulation after removal of the stimulation artifact (red) or without (black). c. Cross—correlogram between SPW—Rs and MFB Stimulation train, averaged on 3s before or after stimulation (Wilcoxon matched pairs test, n=5, Z=1.21, P=0.22). d. Representative recording period of selected channels from two stereotrodes (red and blue) within hippocampal pyramidal layer during a train of MFB stimulation. From top to bottom: Raw data, filtered signal (> 600 Hz) with colored spikes and raster plot of offline—clustered neurons. Red stars indicate SPW—R events. The MFB stimulation train does not preclude burst of ripples classically observed. e. High magnification of the stimulation train. Ripple—related spiking activity is not affected by MFB stimulation train as assessed by the presence of the interneuron’s spikes (black raster) during the stimulation train. f. Higher magnification within the stimulation train. Note the presence of the spike crossing the threshold (dashed line) that triggered the MFB stimulation train (black arrow, spike could not be clustered offline due to stimulation artifact) and during the stimulation train (red spikes on blue traces).
g. Top: PETH centered on the start of MFB stimulation train (red bar) and spikes raster plot of the interneuron displayed in black in the raster shown in panel d. Bottom: Same PETH superimposed with the symmetric trace of the pre— stimulation train period (grey). h. Mean +/—SEM (left) and superimposition of waveforms (right) of the same interneuron’s spikes on seven channels of an octrode, during sleep outside (left panel) or within (right panel) train of MFB stimulation. i. Left: SPW—Rs centered PETH and raster of the same interneuron spikes during sleep without MFB stimulation. Right: same PETH without (black) or with an additional jitter that smoothes oscillating firing during SPW—Rs (blue). j. Superimposition of the raw (left panel, black) or smoothed (right panel, blue) SPW—Rs centered PETH (as in panel i) to the stimulation centered PETH (as in panel g, red).
NS: non significant. Error bars indicate SEM.
Supplementary Figure 5 Offline spike sorting and online spike detection.
a. Comparison between offline clustering and online voltage threshold—based spike sorting. Each column corresponds to the respective waveforms of eight hippocampal neurons recorded from one stereotrode and isolated by offline automatic clustering. The dashed squares emphasize that spikes detected online correspond to the black waveform without ambiguity. MFB stimulation automatically triggered by spike detection causes an electrical artifact that prevents the recording of the end of the waveform. b. Cross—correlograms (grey) and autocorrelogramms (color) of the same eight neurons as in panel a. Mean firing rates (Hz) are indicated underneath each autocorrelogram. c. Specificity quantification of online spike sorting compared to offline clustering for all pairing protocols (sleep and wake). Percentage of detected events categorized into a unique cluster corresponding to the selected place cell (black), into multi—unit activity (MUA, grey), or into other (erroneous) clusters (blue). Insert: detail analysis of clusters containing misassigned spikes. For all sessions: mean percentage of detected spikes in each erroneous cluster and maximum percentage of detected spikes found over all erroneous clusters. Note that among all spikes detected online, offline clustering shows that 80% of them belonged to the cluster of the place cell of interest whereas less than 10% were clustered into multi—unit activity. Multi—unit cluster mostly corresponds to the spikes of the place cell of interest, to very few noisy signal and spikes of other units. For all recording sessions, on average, 1% of the detection events corresponded to other clustered cells. For each animal, the cluster with the maximum of misassigned spikes represented on average 5% of all detected events, meaning that a maximum of 5% of the MFB stimulations were paired to a different neuron. d. Characterization of the cells used in the pairing protocols (n=10) when detected online versus clustered offline in terms of firing rate (Wilcoxon matched pairs test, n=10, Z=2.8, P=0.005), sparsity (Wilcoxon matched pairs test, Z=1.58, P=0.11) and spatial information (Wilcoxon matched pairs test, Z=1.27, P=0.20). Note that sparsity and spatial information were similar for the place cells detected with online and offline methods, whereas there is a decrease in peak and mean firing rate obtained with the online method, consistent with the absence of detection of lower spikes during burst of complex spikes cells. Mean correlation between firing maps for online and offline detected spikes was 0.80+/—0.12.
** P <0.01, NS: non significant.
Supplementary Figure 6 Additional statistical quantification of the place preference induced by the sleep-pairing protocol.
a. Left. Correlation between PRE and POST sessions of sleep—pairing (n=5, red dots) or wake—pairing (n=5, black dots) protocols, in term of percentages of time spent in PF. Note that POST values are always bigger than PRE. Right. Correlation between PRE and POST sessions of those same sleep— and wake—pairing experiments, color scale corresponds to the percentage of the exploring area occupied by the PF. b. Table with exhaustive Khi2 proportion comparison between proportion of time spent inside and outside the place fields (PF) in PRE and POST session of sleep—pairing protocol (2nd column) or between PRE and stimulation sessions of the control wake—pairing protocol (3rd column), see Figure 3c for bar representation. First column corresponds to the comparison between proportion of time spent in the PF in PRE session of sleep—pairing protocol and the theoretical proportion of time spent in the PF according to PF size in case of random exploration (see Online Methods for details). On the basis of the significance in 3rd column, animals were either included (n=5) or excluded (n=2), see exclusion criterion 3. c. Distribution of percentage of time spent in PF (normalized by PF size) for random combination of 4 trials (out of 8) obtained from PRE sessions of all protocols (sleep—pairing protocol, wake—pairing protocol and classical place preference). Grey area represents the generated distribution, with the values of PRE sessions (white dots) and POST sessions (black dots) of the sleep—pairing protocol displayed on top of it. Dotted line represented the 95th percentile. Four of the 5 mice of the sleep—pairing protocol stood in the 5% extreme part of the distribution (the last one was at the 93th percentile). Left. Distribution computed by using time spent in PF, not significantly different from a homogenous exploration of the environment (see Online Methods), for which the normalized value is 1 (student t test, n=5914, P=0.47). Right. Same result is obtained by using time spent in the enlarged PF (student t test, n=5914, P=0.96).
Supplementary Figure 7 Partial correlations of occupancy and firing map before, during and after wake-pairing protocol.
Correlation of occupancy maps in PRE versus POST wake pairing session (grey), or correlation between detected place cell’s firing map (FM) and occupancy map in PRE and POST wake pairing session, or during wake pairing (self—stim) sessions (Friedman test, khi2=16.16 (n=5,df=4) *P=0.0023). Correlation with FM is increased during POST (n=5, correlation PRE/FM vs. POST/FM, Wilcoxon matched pairs test, Z=2.02, *P=0.043, correlation PRE/POST vs. POST/FM, Wilcoxon matched pairs test, Z=2.02, *P=0.043, correlation PRE/FM vs. PRE/POST, Wilcoxon matched pairs test, Z=2.02, *P=0.043). Same results were observed with partial correlation (#) between POST occupancy map and FM when corrected by PRE occupancy map (n=5, correlation PRE/FM vs. POST/FM corrected from PRE Wilcoxon matched pairs test, Z=2.02, *P=0.043, correlation POST/FM vs. POST/FM corrected from PRE, Wilcoxon matched pairs test, Z=1.21, P=0.22). Interestingly, this positive correlation is comparable to the one seen during the control wake—pairing session, where the mouse is actually self—stimulating (yellow, as in fig.2e) (PRE/FM vs. self—stim/FM: Wilcoxon matched pairs test, Z=2.02, *P=0.043, POST/FM vs. self—stim/FM, Wilcoxon matched pairs test, Z=1.21, P=0.22, self—stim/FM vs. POST/FM corrected from PRE Wilcoxon matched pairs test, Z=0.13, P=0.89). Note the absence of correlation in PRE sessions excluding a bias of behavior induced by arena properties. NS: not significant.
Supplementary Figure 8 Comparison between place preference tasks in all configurations and effect of the circadian rhythm.
a—b. Percentage of time spent within normal place field (left) or enlarged place field (right) in PRE and POST sessions of the sleep–pairing protocol (a, 12:00—15:00, n=5) or the wake—pairing protocol (b, 12:00—15:00, n=5). c—d. Percentage of time spent within rewarded area (R—area, left) or enlarged R—area (right) in PRE and POST sessions of the classical place preference using manual stimulation, induced either during the light phase (c, 12:00—15:00, n=7) or the dark phase (d, 0:00—3:00, n=5) of the day. Panels a to d, Wilcoxon matched pairs test, all comparison PRE vs. POST, a—b,d: n=5, Z=2.02, P=0.043; c: n=7, Z=2.37, P=0.018. *P <0.05.
e—f. Comparison of mouse behavior between place cell—stimulation pairing during sleep or wakefulness, with classical place preference (manual place—stimulation pairing) during light or dark phase for different parameters: e. Percentage of time spent in place field or rewarded area (R—area) in the POST session normalized by the PRE session, expressed either as a ratio POST/PRE (left, Kruskall Wallis, H(3,22)=3.27, P=0.35) or as a difference POST—PRE (right, Kruskall Wallis, H(3,22)=1.40, P=0.70). f. Mean running speed averaged in the different protocols (Kruskall Wallis, H(3,22)=4.279, P=0.23).
Supplementary Figure 9 Quantification of behavioral measures expressed in the sleep-pairing protocol (n = 5) and normalization methods.
a. Effect of the definition of the place field (PF). Percentage of time spent in PF (i) and density of occupation inside PF (ii) where “enlarged PF” corresponds to the PF artificially enlarged by the Matlab function imdilate. b. Time spent in PF during PRE and POST session, either not normalized (i, as in panel a), normalized by the time spent in PF during PRE (ii), or normalized by the PF size (iii). c. Time of first entry into PF in real place fields and in area of standardized size surrounding the center of PF. d. Cumulative distance to PF. a—d, Wilcoxon matched pairs test, all comparison PRE vs. POST, n=5, Z=2.02, P=0.043. * P <0.05.
Supplementary Figure 10 Fine characterization of mouse behavior during experiments
Left. Raw trajectories before and after sleep learning
a. Left Column. Firing map with delimited place fields (white line) and maximum firing rate (Hz, red) of the detected place cells. Middle and Right columns. Trajectories during the 4 trials from PRE and POST sessions of the sleep pairing protocol for the five included mice. Blue dots correspond to starting points of each trial. Red areas correspond to place fields used for analysis.
Right. Homogenous exploratory behavior with cable—restrained method
b.Trajectories (up) and mean occupancy maps (bottom) exploration (30 minutes) in an octagonal classical open field (left) and in the same open field (identical shape and external clues) where exploration is restricted by the recording cable length (right). White lines represent central area, corresponding to 50% of the total reachable surface of the environment. c. Number of transitions per minute between central and external area (Wilcoxon matched pairs test, n=6, Z=2.20, P=0.027), and percentage of time spent within the central area (Wilcoxon matched pairs test, n=6, Z=2.20, P=0.027) over 30min—exploration in classical open field (white) versus cable—restrained exploration (black). Error bars indicate SEM.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 (PDF 2216 kb)
Rights and permissions
About this article
Cite this article
de Lavilléon, G., Lacroix, M., Rondi-Reig, L. et al. Explicit memory creation during sleep demonstrates a causal role of place cells in navigation. Nat Neurosci 18, 493–495 (2015). https://doi.org/10.1038/nn.3970
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3970
This article is cited by
-
Dynamic synchronization between hippocampal representations and stepping
Nature (2023)
-
Closed-loop brain stimulation augments fear extinction in male rats
Nature Communications (2023)
-
Spatial maps and oscillations in the healthy hippocampus of Octodon degus, a natural model of sporadic Alzheimer’s disease
Scientific Reports (2022)
-
Neurons as will and representation
Nature Reviews Neuroscience (2022)
-
Replay, the default mode network and the cascaded memory systems model
Nature Reviews Neuroscience (2022)